GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0001508 | Action potential |
| Process | GO:0001666 | Response to hypoxia |
| Process | GO:0003008 | System process |
| Process | GO:0006810 | Transport |
| Process | GO:0006811 | Ion transport |
| Process | GO:0006812 | Cation transport |
| Process | GO:0006814 | Sodium ion transport |
| Process | GO:0006915 | Apoptotic process |
| Process | GO:0006950 | Response to stress |
| Process | GO:0006970 | Response to osmotic stress |
| Process | GO:0007154 | Cell communication |
| Process | GO:0007165 | Signal transduction |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0007399 | Nervous system development |
| Process | GO:0007610 | Behavior |
| Process | GO:0007611 | Learning or memory |
| Process | GO:0007613 | Memory |
| Process | GO:0008219 | Cell death |
| Process | GO:0008627 | Intrinsic apoptotic signaling pathway in response to osmotic stress |
| Process | GO:0009628 | Response to abiotic stimulus |
| Process | GO:0009987 | Cellular process |
| Process | GO:0012501 | Programmed cell death |
| Process | GO:0019226 | Transmission of nerve impulse |
| Process | GO:0019228 | Neuronal action potential |
| Process | GO:0023052 | Signaling |
| Process | GO:0030001 | Metal ion transport |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032879 | Regulation of localization |
| Process | GO:0033554 | Cellular response to stress |
| Process | GO:0034220 | Ion transmembrane transport |
| Process | GO:0034762 | Regulation of transmembrane transport |
| Process | GO:0034765 | Regulation of ion transmembrane transport |
| Process | GO:0035556 | Intracellular signal transduction |
| Process | GO:0035637 | Multicellular organismal signaling |
| Process | GO:0035725 | Sodium ion transmembrane transport |
| Process | GO:0036293 | Response to decreased oxygen levels |
| Process | GO:0036294 | Cellular response to decreased oxygen levels |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0042391 | Regulation of membrane potential |
| Process | GO:0043269 | Regulation of ion transport |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050877 | Nervous system process |
| Process | GO:0050890 | Cognition |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0051049 | Regulation of transport |
| Process | GO:0051179 | Localization |
| Process | GO:0051234 | Establishment of localization |
| Process | GO:0051402 | Neuron apoptotic process |
| Process | GO:0051716 | Cellular response to stimulus |
| Process | GO:0051899 | Membrane depolarization |
| Process | GO:0055085 | Transmembrane transport |
| Process | GO:0062197 | Cellular response to chemical stress |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065008 | Regulation of biological quality |
| Process | GO:0070482 | Response to oxygen levels |
| Process | GO:0070887 | Cellular response to chemical stimulus |
| Process | GO:0070997 | Neuron death |
| Process | GO:0071214 | Cellular response to abiotic stimulus |
| Process | GO:0071453 | Cellular response to oxygen levels |
| Process | GO:0071456 | Cellular response to hypoxia |
| Process | GO:0071470 | Cellular response to osmotic stress |
| Process | GO:0086010 | Membrane depolarization during action potential |
| Process | GO:0097190 | Apoptotic signaling pathway |
| Process | GO:0097193 | Intrinsic apoptotic signaling pathway |
| Process | GO:0098655 | Cation transmembrane transport |
| Process | GO:0098660 | Inorganic ion transmembrane transport |
| Process | GO:0098662 | Inorganic cation transmembrane transport |
| Process | GO:0099505 | Regulation of presynaptic membrane potential |
| Process | GO:0104004 | Cellular response to environmental stimulus |
| Component | GO:0001518 | Voltage-gated sodium channel complex |
| Component | GO:0005886 | Plasma membrane |
| Component | GO:0005887 | Integral component of plasma membrane |
| Component | GO:0005911 | Cell-cell junction |
| Component | GO:0014704 | Intercalated disc |
| Component | GO:0016020 | Membrane |
| Component | GO:0016021 | Integral component of membrane |
| Component | GO:0030054 | Cell junction |
| Component | GO:0030315 | T-tubule |
| Component | GO:0030424 | Axon |
| Component | GO:0031224 | Intrinsic component of membrane |
| Component | GO:0031226 | Intrinsic component of plasma membrane |
| Component | GO:0032991 | Protein-containing complex |
| Component | GO:0033268 | Node of Ranvier |
| Component | GO:0033270 | Paranode region of axon |
| Component | GO:0034702 | Ion channel complex |
| Component | GO:0034703 | Cation channel complex |
| Component | GO:0034706 | Sodium channel complex |
| Component | GO:0042383 | Sarcolemma |
| Component | GO:0042734 | Presynaptic membrane |
| Component | GO:0042995 | Cell projection |
| Component | GO:0043005 | Neuron projection |
| Component | GO:0043194 | Axon initial segment |
| Component | GO:0044291 | Cell-cell contact zone |
| Component | GO:0044304 | Main axon |
| Component | GO:0045202 | Synapse |
| Component | GO:0070161 | Anchoring junction |
| Component | GO:0071944 | Cell periphery |
| Component | GO:0097060 | Synaptic membrane |
| Component | GO:0098590 | Plasma membrane region |
| Component | GO:0098793 | Presynapse |
| Component | GO:0098796 | Membrane protein complex |
| Component | GO:0098797 | Plasma membrane protein complex |
| Component | GO:0098889 | Intrinsic component of presynaptic membrane |
| Component | GO:0098978 | Glutamatergic synapse |
| Component | GO:0099056 | Integral component of presynaptic membrane |
| Component | GO:0099240 | Intrinsic component of synaptic membrane |
| Component | GO:0099699 | Integral component of synaptic membrane |
| Component | GO:0110165 | Cellular anatomical entity |
| Component | GO:0120025 | Plasma membrane bounded cell projection |
| Component | GO:1902495 | Transmembrane transporter complex |
| Component | GO:1990351 | Transporter complex |
| Function | GO:0005215 | Transporter activity |
| Function | GO:0005216 | Ion channel activity |
| Function | GO:0005244 | Voltage-gated ion channel activity |
| Function | GO:0005248 | Voltage-gated sodium channel activity |
| Function | GO:0005261 | Cation channel activity |
| Function | GO:0005272 | Sodium channel activity |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0008324 | Cation transmembrane transporter activity |
| Function | GO:0015075 | Ion transmembrane transporter activity |
| Function | GO:0015081 | Sodium ion transmembrane transporter activity |
| Function | GO:0015267 | Channel activity |
| Function | GO:0015318 | Inorganic molecular entity transmembrane transporter activity |
| Function | GO:0019904 | Protein domain specific binding |
| Function | GO:0022803 | Passive transmembrane transporter activity |
| Function | GO:0022832 | Voltage-gated channel activity |
| Function | GO:0022836 | Gated channel activity |
| Function | GO:0022857 | Transmembrane transporter activity |
| Function | GO:0022890 | Inorganic cation transmembrane transporter activity |
| Function | GO:0030275 | LRR domain binding |
| Function | GO:0031402 | Sodium ion binding |
| Function | GO:0031420 | Alkali metal ion binding |
| Function | GO:0043167 | Ion binding |
| Function | GO:0043169 | Cation binding |
| Function | GO:0043522 | Leucine zipper domain binding |
| Function | GO:0046872 | Metal ion binding |
| Function | GO:0046873 | Metal ion transmembrane transporter activity |
| Function | GO:0099508 | Voltage-gated ion channel activity involved in regulation of presynaptic membrane potential |
Expression profile of Scn2a in omics data
| ID | GSE109165 |
| Title | Suppression of RGSz1 function optimizes the actions of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway |
| Organism | Mus musculus |
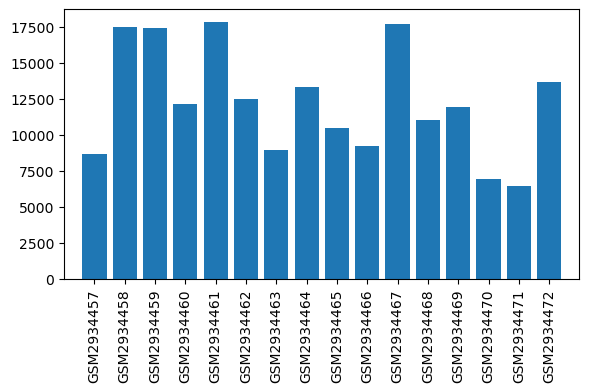
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2934457 | WT_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 8692.0 |
| GSM2934458 | WT_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 17466.0 |
| GSM2934459 | WT_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 17417.0 |
| GSM2934460 | WT_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 12157.0 |
| GSM2934461 | KO_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 17842.0 |
| GSM2934462 | KO_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 12509.0 |
| GSM2934463 | KO_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 8953.0 |
| GSM2934464 | KO_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 13341.0 |
| GSM2934465 | WT_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 10475.0 |
| GSM2934466 | WT_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 9187.0 |
| GSM2934467 | WT_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 17712.0 |
| GSM2934468 | WT_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 11004.0 |
| GSM2934469 | KO_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 11953.0 |
| GSM2934470 | KO_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 6935.0 |
| GSM2934471 | KO_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 6424.0 |
| GSM2934472 | KO_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 13644.0 |
| ID | GSE126662 |
| Title | Comparison between opioid (morphine) and vehicle treatments in mouse trigeminal ganglia and nucleus accumbens |
| Organism | Mus musculus |
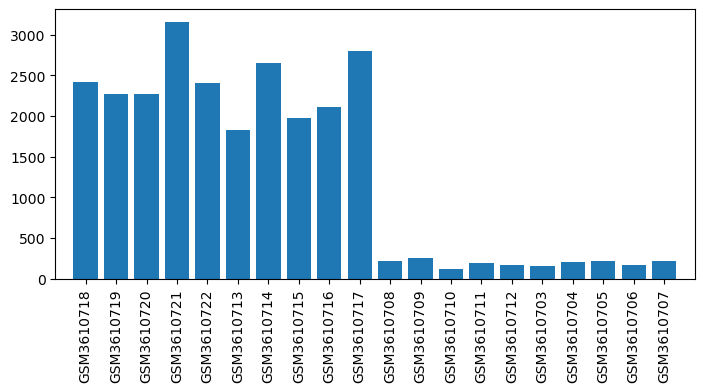
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM3610703 | TG_M1: OIH_M1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 155.2324 |
| GSM3610704 | TG_M2: OIH_M2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 207.84799999999998 |
| GSM3610705 | TG_M3: OIH_M3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 218.22923333333333 |
| GSM3610706 | TG_M4: OIH_M4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 167.37 |
| GSM3610707 | TG_M5: OIH_M5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 215.23927999999998 |
| GSM3610708 | TG_V1: VehicleOIH_V1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 221.6955 |
| GSM3610709 | TG_V2: VehicleOIH_V2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 255.4025333333333 |
| GSM3610710 | TG_V3: VehicleOIH_V3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 117.75306666666667 |
| GSM3610711 | TG_V4: VehicleOIH_V4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 198.48166666666665 |
| GSM3610712 | TG_V5: VehicleOIH_V5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 161.71166666666667 |
| GSM3610713 | NAc_M1: OIH_M1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 1824.4250000000002 |
| GSM3610714 | NAc_M2: OIH_M2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 2645.6916666666666 |
| GSM3610715 | NAc_M3: OIH_M3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 1979.3716666666667 |
| GSM3610716 | NAc_M4: OIH_M4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 2105.623 |
| GSM3610717 | NAc_M5: OIH_M5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 2803.1963333333333 |
| GSM3610718 | NAc_V1: VehicleOIH_V1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 2416.0156666666667 |
| GSM3610719 | NAc_V2: VehicleOIH_V2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 2264.527333333333 |
| GSM3610720 | NAc_V3: VehicleOIH_V3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 2264.7190000000005 |
| GSM3610721 | NAc_V4: VehicleOIH_V4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 3154.3143333333337 |
| GSM3610722 | NAc_V5: VehicleOIH_V5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 2400.072333333333 |
| ID | GSE205201 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [mouse] |
| Organism | Mus musculus |
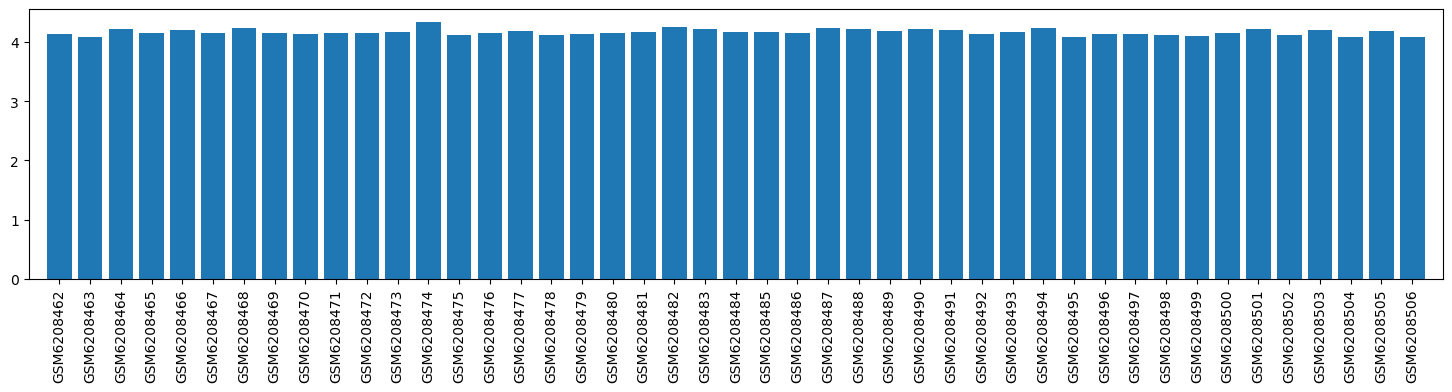
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208462 | Mouse_Baseline_0h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 4.131543 |
| GSM6208463 | Mouse_Baseline_0h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 4.083852 |
| GSM6208464 | Mouse_Baseline_0h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 4.222618 |
| GSM6208465 | Mouse_Baseline_0h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 4.148036 |
| GSM6208466 | Mouse_Baseline_0h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 4.203738 |
| GSM6208467 | Mouse_Vehicle_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 4.159455 |
| GSM6208468 | Mouse_Vehicle_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 4.243804 |
| GSM6208469 | Mouse_Vehicle_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 4.147151 |
| GSM6208470 | Mouse_Vehicle_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 4.145084 |
| GSM6208471 | Mouse_Vehicle_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 4.146239 |
| GSM6208472 | Mouse_Vehicle_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 4.155613 |
| GSM6208473 | Mouse_Vehicle_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 4.164042 |
| GSM6208474 | Mouse_Vehicle_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 4.340091 |
| GSM6208475 | Mouse_Vehicle_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 4.123433 |
| GSM6208476 | Mouse_Vehicle_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 4.162226 |
| GSM6208477 | Mouse_Vehicle_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 4.193707 |
| GSM6208478 | Mouse_Vehicle_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 4.127584 |
| GSM6208479 | Mouse_Vehicle_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 4.138233 |
| GSM6208480 | Mouse_Vehicle_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 4.152025 |
| GSM6208481 | Mouse_Vehicle_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 4.168245 |
| GSM6208482 | Mouse_Vehicle_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 4.249787 |
| GSM6208483 | Mouse_Vehicle_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 4.218005 |
| GSM6208484 | Mouse_Vehicle_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 4.177789 |
| GSM6208485 | Mouse_Vehicle_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 4.164042 |
| GSM6208486 | Mouse_Vehicle_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 4.157894 |
| GSM6208487 | Mouse_APAP_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 4.244054 |
| GSM6208488 | Mouse_APAP_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 4.219739 |
| GSM6208489 | Mouse_APAP_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 4.190683 |
| GSM6208490 | Mouse_APAP_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 4.21786 |
| GSM6208491 | Mouse_APAP_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 4.211079 |
| GSM6208492 | Mouse_APAP_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 4.141416 |
| GSM6208493 | Mouse_APAP_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 4.170709 |
| GSM6208494 | Mouse_APAP_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 4.240588 |
| GSM6208495 | Mouse_APAP_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 4.087545 |
| GSM6208496 | Mouse_APAP_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 4.129913 |
| GSM6208497 | Mouse_APAP_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 4.144052 |
| GSM6208498 | Mouse_APAP_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 4.121893 |
| GSM6208499 | Mouse_APAP_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 4.099428 |
| GSM6208500 | Mouse_APAP_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 4.149867 |
| GSM6208501 | Mouse_APAP_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 4.221038 |
| GSM6208502 | Mouse_APAP_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 4.120994 |
| GSM6208503 | Mouse_APAP_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 4.197545 |
| GSM6208504 | Mouse_APAP_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 4.086139 |
| GSM6208505 | Mouse_APAP_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 4.195968 |
| GSM6208506 | Mouse_APAP_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 4.079228 |
| ID | GSE223541 |
| Title | Oxycodone withdrawal induces HDAC1/2-dependent transcriptional maladaptations in the reward pathway in a murine model of peripheral nerve injury |
| Organism | Mus musculus |
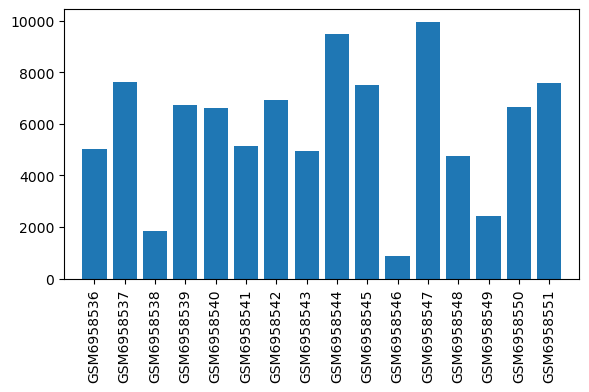
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6958536 | B1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sa | 5033.0 |
| GSM6958537 | C1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 7627.0 |
| GSM6958538 | F1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 1863.0 |
| GSM6958539 | H1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 6717.0 |
| GSM6958540 | B2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 6612.0 |
| GSM6958541 | C2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 5147.0 |
| GSM6958542 | F2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 6938.0 |
| GSM6958543 | H2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 4946.0 |
| GSM6958544 | B3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 9481.0 |
| GSM6958545 | C3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 7506.0 |
| GSM6958546 | F3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 868.0 |
| GSM6958547 | H3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 9944.0 |
| GSM6958548 | B4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 4757.0 |
| GSM6958549 | C4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 2434.0 |
| GSM6958550 | F4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 6639.0 |
| GSM6958551 | H4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 7563.0 |

