GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0001655 | Urogenital system development |
| Process | GO:0001676 | Long-chain fatty acid metabolic process |
| Process | GO:0001822 | Kidney development |
| Process | GO:0006082 | Organic acid metabolic process |
| Process | GO:0006629 | Lipid metabolic process |
| Process | GO:0006631 | Fatty acid metabolic process |
| Process | GO:0006690 | Icosanoid metabolic process |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0009058 | Biosynthetic process |
| Process | GO:0009410 | Response to xenobiotic stimulus |
| Process | GO:0009719 | Response to endogenous stimulus |
| Process | GO:0009725 | Response to hormone |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010033 | Response to organic substance |
| Process | GO:0016053 | Organic acid biosynthetic process |
| Process | GO:0019369 | Arachidonic acid metabolic process |
| Process | GO:0019752 | Carboxylic acid metabolic process |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032787 | Monocarboxylic acid metabolic process |
| Process | GO:0033559 | Unsaturated fatty acid metabolic process |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0043436 | Oxoacid metabolic process |
| Process | GO:0043651 | Linoleic acid metabolic process |
| Process | GO:0044237 | Cellular metabolic process |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0044249 | Cellular biosynthetic process |
| Process | GO:0044255 | Cellular lipid metabolic process |
| Process | GO:0044281 | Small molecule metabolic process |
| Process | GO:0044283 | Small molecule biosynthetic process |
| Process | GO:0046394 | Carboxylic acid biosynthetic process |
| Process | GO:0046456 | Icosanoid biosynthetic process |
| Process | GO:0048252 | Lauric acid metabolic process |
| Process | GO:0048513 | Animal organ development |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0051791 | Medium-chain fatty acid metabolic process |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0072001 | Renal system development |
| Process | GO:0120254 | Olefinic compound metabolic process |
| Process | GO:1901576 | Organic substance biosynthetic process |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005737 | Cytoplasm |
| Component | GO:0005783 | Endoplasmic reticulum |
| Component | GO:0005789 | Endoplasmic reticulum membrane |
| Component | GO:0012505 | Endomembrane system |
| Component | GO:0016020 | Membrane |
| Component | GO:0031090 | Organelle membrane |
| Component | GO:0031984 | Organelle subcompartment |
| Component | GO:0042175 | Nuclear outer membrane-endoplasmic reticulum membrane network |
| Component | GO:0043226 | Organelle |
| Component | GO:0043227 | Membrane-bounded organelle |
| Component | GO:0043229 | Intracellular organelle |
| Component | GO:0043231 | Intracellular membrane-bounded organelle |
| Component | GO:0098827 | Endoplasmic reticulum subcompartment |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0003824 | Catalytic activity |
| Function | GO:0004497 | Monooxygenase activity |
| Function | GO:0005488 | Binding |
| Function | GO:0005504 | Fatty acid binding |
| Function | GO:0005506 | Iron ion binding |
| Function | GO:0008289 | Lipid binding |
| Function | GO:0008391 | Arachidonic acid monooxygenase activity |
| Function | GO:0008392 | Arachidonic acid epoxygenase activity |
| Function | GO:0008405 | Arachidonic acid 11,12-epoxygenase activity |
| Function | GO:0016491 | Oxidoreductase activity |
| Function | GO:0016705 | Oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen |
| Function | GO:0016713 | Oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced iron-sulfur protein as one donor, and incorporation of one atom of oxygen |
| Function | GO:0018685 | Alkane 1-monooxygenase activity |
| Function | GO:0020037 | Heme binding |
| Function | GO:0031406 | Carboxylic acid binding |
| Function | GO:0033293 | Monocarboxylic acid binding |
| Function | GO:0036041 | Long-chain fatty acid binding |
| Function | GO:0036094 | Small molecule binding |
| Function | GO:0043167 | Ion binding |
| Function | GO:0043168 | Anion binding |
| Function | GO:0043169 | Cation binding |
| Function | GO:0043177 | Organic acid binding |
| Function | GO:0046872 | Metal ion binding |
| Function | GO:0046906 | Tetrapyrrole binding |
| Function | GO:0046914 | Transition metal ion binding |
| Function | GO:0050542 | Icosanoid binding |
| Function | GO:0050543 | Icosatetraenoic acid binding |
| Function | GO:0050544 | Arachidonic acid binding |
| Function | GO:0097159 | Organic cyclic compound binding |
| Function | GO:0102033 | Long-chain fatty acid omega-hydroxylase activity |
| Function | GO:0120250 | Fatty acid omega-hydroxylase activity |
| Function | GO:1901363 | Heterocyclic compound binding |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| WikiPathways | WP133 | Fatty acid omega-oxidation |
| KEGG | rno00071 | Fatty acid degradation |
| KEGG | rno00590 | Arachidonic acid metabolism |
| KEGG | rno00830 | Retinol metabolism |
| KEGG | rno01100 | Metabolic pathways |
| KEGG | rno03320 | PPAR signaling pathway |
| KEGG | rno04270 | Vascular smooth muscle contraction |
| KEGG | rno04750 | Inflammatory mediator regulation of TRP channels |
Expression profile of Cyp4a2 in omics data
| ID | GSE100499 |
| Title | Exploring acupuncture mechanism in preconditioning of IRI |
| Organism | Rattus norvegicus |
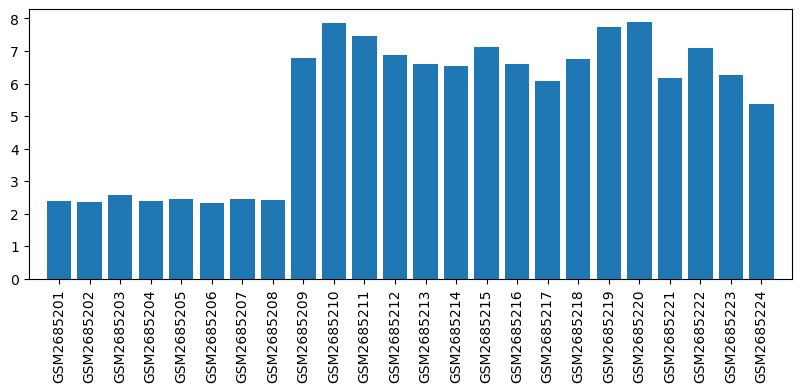
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2685201 | MX1 | tissue: Myocardium gender: male | 2.3894625 |
| GSM2685202 | MX2 | tissue: Myocardium gender: female | 2.3464642 |
| GSM2685203 | MX3 | tissue: Myocardium gender: female | 2.5737011 |
| GSM2685204 | MX4 | tissue: Myocardium gender: male | 2.3913832 |
| GSM2685205 | DZ1 | tissue: Myocardium gender: male | 2.452026 |
| GSM2685206 | DZ2 | tissue: Myocardium gender: male | 2.3219855 |
| GSM2685207 | DZ3 | tissue: Myocardium gender: female | 2.457881 |
| GSM2685208 | DZ4 | tissue: Myocardium gender: female | 2.424425 |
| GSM2685209 | JJ1 | tissue: Myocardium gender: male | 6.780395 |
| GSM2685210 | JJ2 | tissue: Myocardium gender: female | 7.8689604 |
| GSM2685211 | JJ3 | tissue: Myocardium gender: female | 7.4710317 |
| GSM2685212 | JJ4 | tissue: Myocardium gender: male | 6.8769064 |
| GSM2685213 | NG1 | tissue: Myocardium gender: male | 6.6138105 |
| GSM2685214 | NG2 | tissue: Myocardium gender: female | 6.5330486 |
| GSM2685215 | NG3 | tissue: Myocardium gender: male | 7.131818 |
| GSM2685216 | NG4 | tissue: Myocardium gender: female | 6.59519 |
| GSM2685217 | YL1 | tissue: Myocardium gender: male | 6.0828485 |
| GSM2685218 | YL2 | tissue: Myocardium gender: male | 6.7483344 |
| GSM2685219 | YL3 | tissue: Myocardium gender: female | 7.75232 |
| GSM2685220 | YL4 | tissue: Myocardium gender: female | 7.892209 |
| GSM2685221 | QC1 | tissue: Myocardium gender: male | 6.1719384 |
| GSM2685222 | QC2 | tissue: Myocardium gender: male | 7.0858316 |
| GSM2685223 | QC3 | tissue: Myocardium gender: female | 6.277372 |
| GSM2685224 | QC4 | tissue: Myocardium gender: female | 5.370904 |
| ID | GSE136833 |
| Title | Effects of lidocaine on the expression of rat spinal cord |
| Organism | Rattus norvegicus |
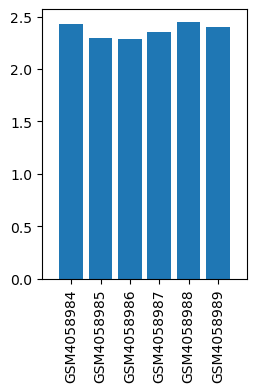
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4058984 | Control_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 2.429382619 |
| GSM4058985 | Control_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 2.295745935 |
| GSM4058986 | Control_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 2.286461304 |
| GSM4058987 | Lidocaine_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 2.358342771 |
| GSM4058988 | Lidocaine_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 2.449313333 |
| GSM4058989 | Lidocaine_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 2.401637681 |
| ID | GSE138454 |
| Title | Impact of inflammation on hepatic subcellular energetics in anesthetized rats |
| Organism | Rattus norvegicus |
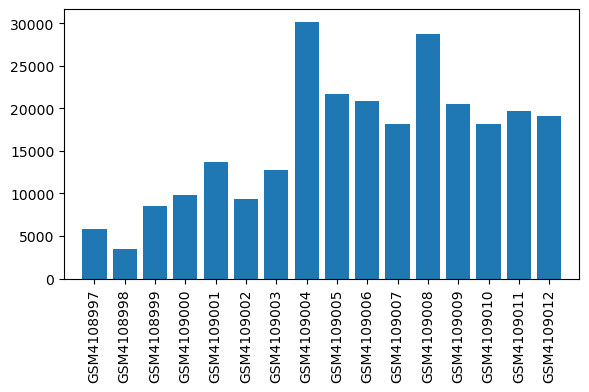
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4108997 | Sample 1 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 5845.0 |
| GSM4108998 | Sample 2 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 3499.0 |
| GSM4108999 | Sample 3 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 8543.0 |
| GSM4109000 | Sample 4 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 9799.0 |
| GSM4109001 | Sample 5 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 13717.0 |
| GSM4109002 | Sample 6 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 9324.0 |
| GSM4109003 | Sample 7 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 12738.0 |
| GSM4109004 | Sample 8 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 30140.0 |
| GSM4109005 | Sample9 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 21686.0 |
| GSM4109006 | Sample10 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 20885.0 |
| GSM4109007 | Sample11 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 18115.0 |
| GSM4109008 | Sample12 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 28680.0 |
| GSM4109009 | Sample13 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 20518.0 |
| GSM4109010 | Sample14 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 18174.0 |
| GSM4109011 | Sample 15 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 19685.0 |
| GSM4109012 | Sample16 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 19152.0 |
| ID | GSE139220 |
| Title | Expression data from sevoflurane anesthesia treated rat |
| Organism | Rattus norvegicus |
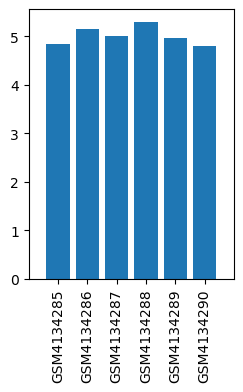
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4134285 | Brain_Control_rep1 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 4.840479 |
| GSM4134286 | Brain_Control_rep2 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 5.139357 |
| GSM4134287 | Brain_Control_rep3 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 5.010787 |
| GSM4134288 | Brain_Sevoflurane_rep1 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 5.294138 |
| GSM4134289 | Brain_Sevoflurane_rep2 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 4.961997 |
| GSM4134290 | Brain_Sevoflurane_rep3 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 4.808089 |
| ID | GSE143895 |
| Title | GPR160 de-orphanization reveals critical roles in neuropathic pain in rodents |
| Organism | Rattus norvegicus |
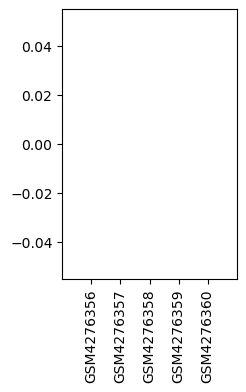
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4276356 | dhsc01 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.0 |
| GSM4276357 | dhsc03 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.0 |
| GSM4276358 | dhsc05 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.0 |
| GSM4276359 | dhsc13 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 0.0 |
| GSM4276360 | dhsc15 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 0.0 |
| ID | GSE157924 |
| Title | Regulation of acupuncture on genes in brain stem |
| Organism | Rattus norvegicus |
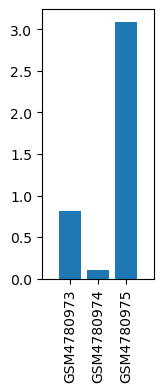
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4780973 | Rattus_JSSZ(Sham) | tissue: Rattus brainstem of modeling treatment: Sham | 0.813715461 |
| GSM4780974 | Rattus_JXZ(Hand Acu) | tissue: Rattus brainstem of modeling treatment: Hand Acu | 0.099622869 |
| GSM4780975 | Rattus_MXZ(TBI) | tissue: Rattus brainstem of modeling treatment: TBI | 3.089129098 |
| ID | GSE190472 |
| Title | Study on VEGF regulation mechanism of acupuncture improving endometrial receptivity in IVF-ET |
| Organism | Rattus norvegicus |
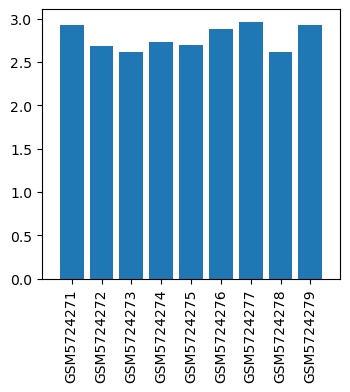
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5724271 | model 21 | Sex: female tissue: Uterine tissue condition: model | 2.9282 |
| GSM5724272 | model 22 | Sex: female tissue: Uterine tissue condition: model | 2.6847 |
| GSM5724273 | model 23 | Sex: female tissue: Uterine tissue condition: model | 2.6155 |
| GSM5724274 | treatment31 | Sex: female tissue: Uterine tissue condition: treatment | 2.7305 |
| GSM5724275 | treatment32 | Sex: female tissue: Uterine tissue condition: treatment | 2.6954 |
| GSM5724276 | treatment33 | Sex: female tissue: Uterine tissue condition: treatment | 2.8812 |
| GSM5724277 | control 41 | Sex: female tissue: Uterine tissue condition: control | 2.9603 |
| GSM5724278 | control 42 | Sex: female tissue: Uterine tissue condition: control | 2.6113 |
| GSM5724279 | control 43 | Sex: female tissue: Uterine tissue condition: control | 2.93 |
| ID | GSE192399 |
| Title | Gene expression of brain regions activated by sevoflurane and propofol |
| Organism | Rattus norvegicus |
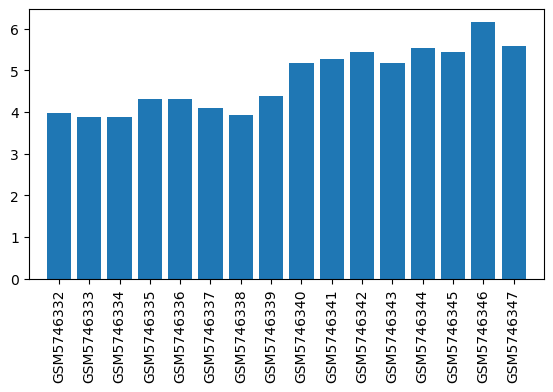
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5746332 | LHb, Sevoflurane, 60mins | brain region: the lateral habenula (LHb) treatment: Sevoflurane | 3.97693 |
| GSM5746333 | LHb, O2, 60mins | brain region: the lateral habenula (LHb) treatment: O2 | 3.87258 |
| GSM5746334 | MHb, Sevoflurane, 60mins | brain region: the medial habenula (MHb) treatment: Sevoflurane | 3.86996 |
| GSM5746335 | MHb, O2, 60mins | brain region: the medial habenula (MHb) treatment: O2 | 4.29979 |
| GSM5746336 | Sol, Sevoflurane, 60mins | brain region: the solitary nucleus (Sol) treatment: Sevoflurane | 4.31574 |
| GSM5746337 | Sol, O2, 60mins | brain region: the solitary nucleus (Sol) treatment: O2 | 4.08831 |
| GSM5746338 | Mve, Sevoflurane, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Sevoflurane | 3.93379 |
| GSM5746339 | Mve, O2, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: O2 | 4.38509 |
| GSM5746340 | LHb, Propofol, 60mins | brain region: the lateral habenula (LHb) treatment: Propofol | 5.1793 |
| GSM5746341 | LHb, Intralipos, 60mins | brain region: the lateral habenula (LHb) treatment: Intralipos | 5.27507 |
| GSM5746342 | MHb, Propofol, 60mins | brain region: the medial habenula (MHb) treatment: Propofol | 5.43882 |
| GSM5746343 | MHb, Intralipos, 60mins | brain region: the medial habenula (MHb) treatment: Intralipos | 5.17531 |
| GSM5746344 | Sol, Propofol, 60mins | brain region: the solitary nucleus (Sol) treatment: Propofol | 5.52274 |
| GSM5746345 | Sol, Intralipos, 60mins | brain region: the solitary nucleus (Sol) treatment: Intralipos | 5.42494 |
| GSM5746346 | MVe, Propofol, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Propofol | 6.15649 |
| GSM5746347 | MVe, Intralipos, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Intralipos | 5.59145 |
| ID | GSE244436 |
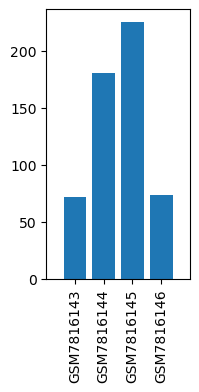
| Title | Similarity and dissimilarity in alteration of gene expression profile associated with inhalational anesthesia between sevoflurane and desflurane |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7816143 | Liver, Control for SevoAnes | tissue: liver genotype: wild type treatment: Control for SevoAnes | 71.355934 |
| GSM7816144 | Liver, Anesthetized with sevoflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with sevoflurane for 6 h | 180.586487 |
| GSM7816145 | Liver, Control for DesAnes | tissue: liver genotype: wild type treatment: Control for DesAnes | 225.195114 |
| GSM7816146 | Liver, Anesthetized with desflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with desflurane for 6 h | 73.645905 |
| ID | GSE48025 |
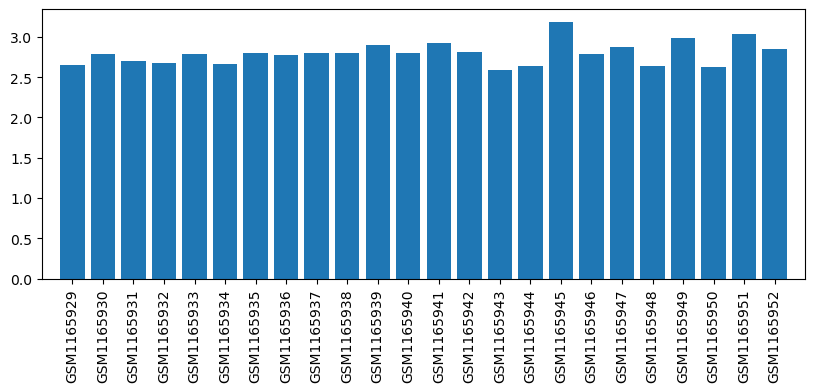
| Title | Affymetrix Chip Data of the Transcriptome of the Rheumatoid Arthritis Rat Model Treated with Acupuncture (Affymetrix, mRNA, batch1) |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1165929 | RAT_NORA_T0_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats before therapy | 2.646162889 |
| GSM1165930 | RAT_NORA_T0_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats before therapy | 2.783681069 |
| GSM1165931 | RAT_RA_T0_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats before therapy | 2.703737485 |
| GSM1165932 | RAT_RA_T0_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats before therapy | 2.678175535 |
| GSM1165933 | RAT_NORA_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 1 hour after therapy | 2.791519972 |
| GSM1165934 | RAT_NORA_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 1 hour after therapy | 2.668383904 |
| GSM1165935 | RAT_AP_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour acupuncture therapy | 2.801560526 |
| GSM1165936 | RAT_AP_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour acupuncture therapy | 2.776715486 |
| GSM1165937 | RAT_MTX_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour MTX therapy | 2.796261041 |
| GSM1165938 | RAT_MTX_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour MTX therapy | 2.796261041 |
| GSM1165939 | RAT_ANE_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour ANE therapy | 2.89785453 |
| GSM1165940 | RAT_ANE_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour ANE therapy | 2.795173427 |
| GSM1165941 | RAT_PLA_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour placebo therapy | 2.928028324 |
| GSM1165942 | RAT_PLA_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour placebo therapy | 2.809350259 |
| GSM1165943 | RAT_NORA_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 34 days after therapy | 2.585641897 |
| GSM1165944 | RAT_NORA_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 34 days after therapy | 2.641786104 |
| GSM1165945 | RAT_AP_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days acupuncture therapy | 3.184757267 |
| GSM1165946 | RAT_AP_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days acupuncture therapy | 2.7871177 |
| GSM1165947 | RAT_MTX_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days MTX therapy | 2.875258729 |
| GSM1165948 | RAT_MTX_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days MTX therapy | 2.639507931 |
| GSM1165949 | RAT_ANE_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days ANE therapy | 2.986008612 |
| GSM1165950 | RAT_ANE_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days ANE therapy | 2.623638365 |
| GSM1165951 | RAT_PLA_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days placebo therapy | 3.039391539 |
| GSM1165952 | RAT_PLA_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days placebo therapy | 2.848097879 |
| ID | GSE53897 |
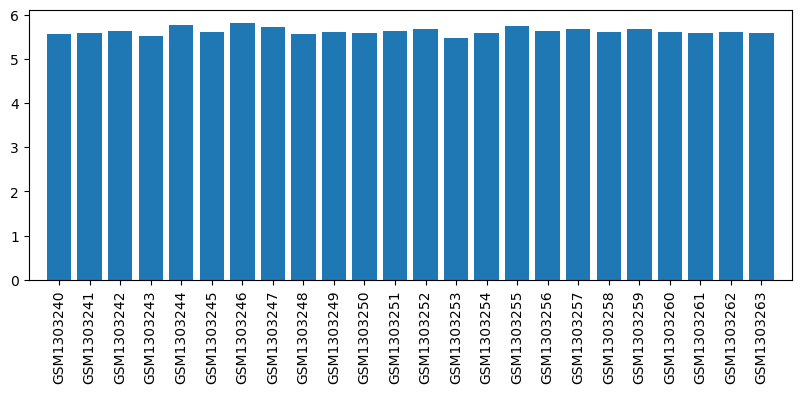
| Title | Genome-wide analysis of dorsal root ganglion (DRG) gene expression after morphine or oxycodone treatment |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1303240 | DRG RVM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.57033 |
| GSM1303241 | DRG RVM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.589932 |
| GSM1303242 | DRG RVM3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.637362 |
| GSM1303243 | DRG RVM7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.520891 |
| GSM1303244 | DRG RVOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.776291 |
| GSM1303245 | DRG RVOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.619682 |
| GSM1303246 | DRG RVOx5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.816014 |
| GSM1303247 | DRG RVOx7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.715746 |
| GSM1303248 | DRG RVS4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.567611 |
| GSM1303249 | DRG RVS5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.601662 |
| GSM1303250 | DRG RVS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.595378 |
| GSM1303251 | DRG RVS7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.623228 |
| GSM1303252 | DRG RSM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.689202 |
| GSM1303253 | DRG RSM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.485823 |
| GSM1303254 | DRG RSM4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.597361 |
| GSM1303255 | DRG RSM5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.752819 |
| GSM1303256 | DRG RSOx1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.634487 |
| GSM1303257 | DRG RSOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.678274 |
| GSM1303258 | DRG RSOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.601464 |
| GSM1303259 | DRG RSOx4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.673919 |
| GSM1303260 | DRG RSS1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.605037 |
| GSM1303261 | DRG RSS2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.579099 |
| GSM1303262 | DRG RSS3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.622863 |
| GSM1303263 | DRG RSS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.582916 |
| ID | GSE58456 |
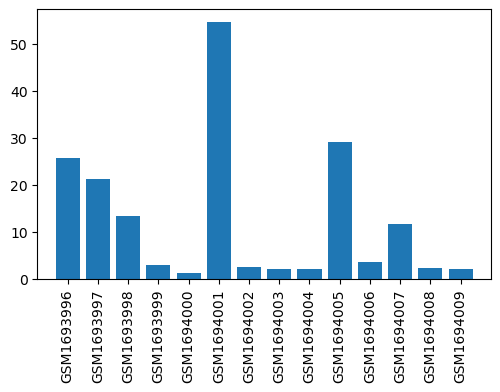
| Title | Affymetrix Chip Data of the Transcriptome of the Rheumatoid Arthritis Rat Model Treated with Acupuncture (Affymetrix, mRNA, batch 2) |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1693996 | Peripheral blood mononuclear cells from NORA before therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 25.67814749 |
| GSM1693997 | Peripheral blood mononuclear cells from NORA before therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 21.21338376 |
| GSM1693998 | Peripheral blood mononuclear cells from NORA before therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 13.3662016 |
| GSM1693999 | Peripheral blood mononuclear cells from RA before therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats before therapy | 2.863078789 |
| GSM1694000 | Peripheral blood mononuclear cells from RA before therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats before therapy | 1.203317907 |
| GSM1694001 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 54.58892413 |
| GSM1694002 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 2.518109881 |
| GSM1694003 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 2.083546871 |
| GSM1694004 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 1.975555388 |
| GSM1694005 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 29.05271316 |
| GSM1694006 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 3.573819003 |
| GSM1694007 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 11.7038656 |
| GSM1694008 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 2.350917917 |
| GSM1694009 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 2.156991597 |

