GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0000122 | Negative regulation of transcription by RNA polymerase II |
| Process | GO:0001817 | Regulation of cytokine production |
| Process | GO:0001818 | Negative regulation of cytokine production |
| Process | GO:0001819 | Positive regulation of cytokine production |
| Process | GO:0001878 | Response to yeast |
| Process | GO:0001906 | Cell killing |
| Process | GO:0001909 | Leukocyte mediated cytotoxicity |
| Process | GO:0002237 | Response to molecule of bacterial origin |
| Process | GO:0002252 | Immune effector process |
| Process | GO:0002376 | Immune system process |
| Process | GO:0002437 | Inflammatory response to antigenic stimulus |
| Process | GO:0002438 | Acute inflammatory response to antigenic stimulus |
| Process | GO:0002440 | Production of molecular mediator of immune response |
| Process | GO:0002443 | Leukocyte mediated immunity |
| Process | GO:0002444 | Myeloid leukocyte mediated immunity |
| Process | GO:0002446 | Neutrophil mediated immunity |
| Process | GO:0002523 | Leukocyte migration involved in inflammatory response |
| Process | GO:0002526 | Acute inflammatory response |
| Process | GO:0002682 | Regulation of immune system process |
| Process | GO:0002684 | Positive regulation of immune system process |
| Process | GO:0002685 | Regulation of leukocyte migration |
| Process | GO:0002691 | Regulation of cellular extravasation |
| Process | GO:0002775 | Antimicrobial peptide production |
| Process | GO:0002777 | Antimicrobial peptide biosynthetic process |
| Process | GO:0002778 | Antibacterial peptide production |
| Process | GO:0002780 | Antibacterial peptide biosynthetic process |
| Process | GO:0006355 | Regulation of transcription, DNA-templated |
| Process | GO:0006357 | Regulation of transcription by RNA polymerase II |
| Process | GO:0006508 | Proteolysis |
| Process | GO:0006518 | Peptide metabolic process |
| Process | GO:0006807 | Nitrogen compound metabolic process |
| Process | GO:0006810 | Transport |
| Process | GO:0006909 | Phagocytosis |
| Process | GO:0006950 | Response to stress |
| Process | GO:0006952 | Defense response |
| Process | GO:0006954 | Inflammatory response |
| Process | GO:0006955 | Immune response |
| Process | GO:0006959 | Humoral immune response |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0008284 | Positive regulation of cell population proliferation |
| Process | GO:0009058 | Biosynthetic process |
| Process | GO:0009314 | Response to radiation |
| Process | GO:0009411 | Response to UV |
| Process | GO:0009416 | Response to light stimulus |
| Process | GO:0009605 | Response to external stimulus |
| Process | GO:0009607 | Response to biotic stimulus |
| Process | GO:0009617 | Response to bacterium |
| Process | GO:0009620 | Response to fungus |
| Process | GO:0009628 | Response to abiotic stimulus |
| Process | GO:0009889 | Regulation of biosynthetic process |
| Process | GO:0009890 | Negative regulation of biosynthetic process |
| Process | GO:0009892 | Negative regulation of metabolic process |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010033 | Response to organic substance |
| Process | GO:0010467 | Gene expression |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010556 | Regulation of macromolecule biosynthetic process |
| Process | GO:0010558 | Negative regulation of macromolecule biosynthetic process |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0010605 | Negative regulation of macromolecule metabolic process |
| Process | GO:0010628 | Positive regulation of gene expression |
| Process | GO:0010629 | Negative regulation of gene expression |
| Process | GO:0016192 | Vesicle-mediated transport |
| Process | GO:0016477 | Cell migration |
| Process | GO:0019219 | Regulation of nucleobase-containing compound metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0019538 | Protein metabolic process |
| Process | GO:0019730 | Antimicrobial humoral response |
| Process | GO:0019731 | Antibacterial humoral response |
| Process | GO:0022407 | Regulation of cell-cell adhesion |
| Process | GO:0022409 | Positive regulation of cell-cell adhesion |
| Process | GO:0030155 | Regulation of cell adhesion |
| Process | GO:0030334 | Regulation of cell migration |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0031324 | Negative regulation of cellular metabolic process |
| Process | GO:0031326 | Regulation of cellular biosynthetic process |
| Process | GO:0031327 | Negative regulation of cellular biosynthetic process |
| Process | GO:0032496 | Response to lipopolysaccharide |
| Process | GO:0032642 | Regulation of chemokine production |
| Process | GO:0032677 | Regulation of interleukin-8 production |
| Process | GO:0032682 | Negative regulation of chemokine production |
| Process | GO:0032717 | Negative regulation of interleukin-8 production |
| Process | GO:0032757 | Positive regulation of interleukin-8 production |
| Process | GO:0033993 | Response to lipid |
| Process | GO:0034641 | Cellular nitrogen compound metabolic process |
| Process | GO:0040012 | Regulation of locomotion |
| Process | GO:0042127 | Regulation of cell population proliferation |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0042742 | Defense response to bacterium |
| Process | GO:0043043 | Peptide biosynthetic process |
| Process | GO:0043170 | Macromolecule metabolic process |
| Process | GO:0043207 | Response to external biotic stimulus |
| Process | GO:0043603 | Cellular amide metabolic process |
| Process | GO:0043604 | Amide biosynthetic process |
| Process | GO:0044237 | Cellular metabolic process |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0044249 | Cellular biosynthetic process |
| Process | GO:0044271 | Cellular nitrogen compound biosynthetic process |
| Process | GO:0044403 | Biological process involved in symbiotic interaction |
| Process | GO:0044419 | Biological process involved in interspecies interaction between organisms |
| Process | GO:0045785 | Positive regulation of cell adhesion |
| Process | GO:0045892 | Negative regulation of transcription, DNA-templated |
| Process | GO:0045934 | Negative regulation of nucleobase-containing compound metabolic process |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048519 | Negative regulation of biological process |
| Process | GO:0048522 | Positive regulation of cellular process |
| Process | GO:0048523 | Negative regulation of cellular process |
| Process | GO:0048583 | Regulation of response to stimulus |
| Process | GO:0048584 | Positive regulation of response to stimulus |
| Process | GO:0048660 | Regulation of smooth muscle cell proliferation |
| Process | GO:0048661 | Positive regulation of smooth muscle cell proliferation |
| Process | GO:0048870 | Cell motility |
| Process | GO:0050776 | Regulation of immune response |
| Process | GO:0050778 | Positive regulation of immune response |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050829 | Defense response to Gram-negative bacterium |
| Process | GO:0050832 | Defense response to fungus |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0050900 | Leukocyte migration |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051172 | Negative regulation of nitrogen compound metabolic process |
| Process | GO:0051179 | Localization |
| Process | GO:0051234 | Establishment of localization |
| Process | GO:0051239 | Regulation of multicellular organismal process |
| Process | GO:0051240 | Positive regulation of multicellular organismal process |
| Process | GO:0051241 | Negative regulation of multicellular organismal process |
| Process | GO:0051252 | Regulation of RNA metabolic process |
| Process | GO:0051253 | Negative regulation of RNA metabolic process |
| Process | GO:0051702 | Biological process involved in interaction with symbiont |
| Process | GO:0051707 | Response to other organism |
| Process | GO:0051873 | Killing by host of symbiont cells |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0061844 | Antimicrobial humoral immune response mediated by antimicrobial peptide |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0070942 | Neutrophil mediated cytotoxicity |
| Process | GO:0070943 | Neutrophil-mediated killing of symbiont cell |
| Process | GO:0070944 | Neutrophil-mediated killing of bacterium |
| Process | GO:0070945 | Neutrophil-mediated killing of gram-negative bacterium |
| Process | GO:0070947 | Neutrophil-mediated killing of fungus |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0098542 | Defense response to other organism |
| Process | GO:1901564 | Organonitrogen compound metabolic process |
| Process | GO:1901566 | Organonitrogen compound biosynthetic process |
| Process | GO:1901576 | Organic substance biosynthetic process |
| Process | GO:1901700 | Response to oxygen-containing compound |
| Process | GO:1902679 | Negative regulation of RNA biosynthetic process |
| Process | GO:1903037 | Regulation of leukocyte cell-cell adhesion |
| Process | GO:1903039 | Positive regulation of leukocyte cell-cell adhesion |
| Process | GO:1903236 | Regulation of leukocyte tethering or rolling |
| Process | GO:1903238 | Positive regulation of leukocyte tethering or rolling |
| Process | GO:1903506 | Regulation of nucleic acid-templated transcription |
| Process | GO:1903507 | Negative regulation of nucleic acid-templated transcription |
| Process | GO:1904994 | Regulation of leukocyte adhesion to vascular endothelial cell |
| Process | GO:1904996 | Positive regulation of leukocyte adhesion to vascular endothelial cell |
| Process | GO:2000145 | Regulation of cell motility |
| Process | GO:2001141 | Regulation of RNA biosynthetic process |
| Process | GO:0002812 | Biosynthetic process of antibacterial peptides active against Gram-negative bacteria |
| Component | GO:0005576 | Extracellular region |
| Component | GO:0005615 | Extracellular space |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005667 | Transcription regulator complex |
| Component | GO:0005737 | Cytoplasm |
| Component | GO:0009986 | Cell surface |
| Component | GO:0012505 | Endomembrane system |
| Component | GO:0017053 | Transcription repressor complex |
| Component | GO:0030141 | Secretory granule |
| Component | GO:0031410 | Cytoplasmic vesicle |
| Component | GO:0031982 | Vesicle |
| Component | GO:0032991 | Protein-containing complex |
| Component | GO:0043226 | Organelle |
| Component | GO:0043227 | Membrane-bounded organelle |
| Component | GO:0043229 | Intracellular organelle |
| Component | GO:0043231 | Intracellular membrane-bounded organelle |
| Component | GO:0097708 | Intracellular vesicle |
| Component | GO:0099503 | Secretory vesicle |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0002020 | Protease binding |
| Function | GO:0003712 | Transcription coregulator activity |
| Function | GO:0003714 | Transcription corepressor activity |
| Function | GO:0003824 | Catalytic activity |
| Function | GO:0004175 | Endopeptidase activity |
| Function | GO:0004252 | Serine-type endopeptidase activity |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0005539 | Glycosaminoglycan binding |
| Function | GO:0008201 | Heparin binding |
| Function | GO:0008233 | Peptidase activity |
| Function | GO:0008236 | Serine-type peptidase activity |
| Function | GO:0016787 | Hydrolase activity |
| Function | GO:0017171 | Serine hydrolase activity |
| Function | GO:0019899 | Enzyme binding |
| Function | GO:0019955 | Cytokine binding |
| Function | GO:0097367 | Carbohydrate derivative binding |
| Function | GO:0140096 | Catalytic activity, acting on a protein |
| Function | GO:0140110 | Transcription regulator activity |
| Function | GO:1901681 | Sulfur compound binding |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| RCTM | RNO-1474228 | Degradation of the extracellular matrix |
| RCTM | RNO-1474244 | Extracellular matrix organization |
| RCTM | RNO-1592389 | Activation of Matrix Metalloproteinases |
| RCTM | RNO-168249 | Innate Immune System |
| RCTM | RNO-168256 | Immune System |
| RCTM | RNO-5218859 | Regulated Necrosis |
| RCTM | RNO-5357801 | Programmed Cell Death |
| RCTM | RNO-5620971 | Pyroptosis |
| RCTM | RNO-6798695 | Neutrophil degranulation |
| RCTM | RNO-6803157 | Antimicrobial peptides |
| KEGG | rno04613 | Neutrophil extracellular trap formation |
| KEGG | rno05202 | Transcriptional misregulation in cancer |
| KEGG | rno05322 | Systemic lupus erythematosus |
Expression profile of Elane in omics data
| ID | GSE100499 |
| Title | Exploring acupuncture mechanism in preconditioning of IRI |
| Organism | Rattus norvegicus |
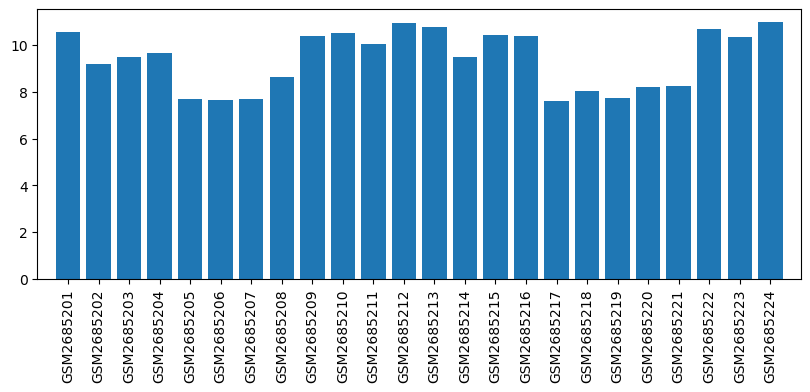
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2685201 | MX1 | tissue: Myocardium gender: male | 10.553924 |
| GSM2685202 | MX2 | tissue: Myocardium gender: female | 9.17723 |
| GSM2685203 | MX3 | tissue: Myocardium gender: female | 9.489714 |
| GSM2685204 | MX4 | tissue: Myocardium gender: male | 9.660267 |
| GSM2685205 | DZ1 | tissue: Myocardium gender: male | 7.7083898 |
| GSM2685206 | DZ2 | tissue: Myocardium gender: male | 7.665864 |
| GSM2685207 | DZ3 | tissue: Myocardium gender: female | 7.6782575 |
| GSM2685208 | DZ4 | tissue: Myocardium gender: female | 8.640213 |
| GSM2685209 | JJ1 | tissue: Myocardium gender: male | 10.39276 |
| GSM2685210 | JJ2 | tissue: Myocardium gender: female | 10.51917 |
| GSM2685211 | JJ3 | tissue: Myocardium gender: female | 10.0633545 |
| GSM2685212 | JJ4 | tissue: Myocardium gender: male | 10.935918 |
| GSM2685213 | NG1 | tissue: Myocardium gender: male | 10.780749 |
| GSM2685214 | NG2 | tissue: Myocardium gender: female | 9.511051 |
| GSM2685215 | NG3 | tissue: Myocardium gender: male | 10.413771 |
| GSM2685216 | NG4 | tissue: Myocardium gender: female | 10.398956 |
| GSM2685217 | YL1 | tissue: Myocardium gender: male | 7.59737 |
| GSM2685218 | YL2 | tissue: Myocardium gender: male | 8.053409 |
| GSM2685219 | YL3 | tissue: Myocardium gender: female | 7.7251053 |
| GSM2685220 | YL4 | tissue: Myocardium gender: female | 8.185035 |
| GSM2685221 | QC1 | tissue: Myocardium gender: male | 8.263228 |
| GSM2685222 | QC2 | tissue: Myocardium gender: male | 10.67293 |
| GSM2685223 | QC3 | tissue: Myocardium gender: female | 10.356289 |
| GSM2685224 | QC4 | tissue: Myocardium gender: female | 10.989764 |
| ID | GSE136833 |
| Title | Effects of lidocaine on the expression of rat spinal cord |
| Organism | Rattus norvegicus |
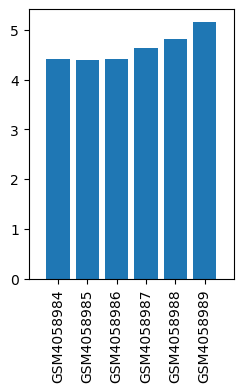
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4058984 | Control_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 4.405228545 |
| GSM4058985 | Control_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 4.394505957 |
| GSM4058986 | Control_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 4.408442525 |
| GSM4058987 | Lidocaine_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 4.633199484 |
| GSM4058988 | Lidocaine_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 4.8073751 |
| GSM4058989 | Lidocaine_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 5.157012271 |
| ID | GSE138454 |
| Title | Impact of inflammation on hepatic subcellular energetics in anesthetized rats |
| Organism | Rattus norvegicus |
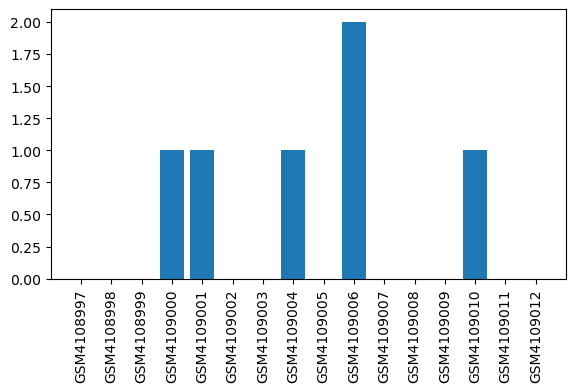
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4108997 | Sample 1 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4108998 | Sample 2 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4108999 | Sample 3 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4109000 | Sample 4 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 1.0 |
| GSM4109001 | Sample 5 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 1.0 |
| GSM4109002 | Sample 6 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 0.0 |
| GSM4109003 | Sample 7 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 0.0 |
| GSM4109004 | Sample 8 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 1.0 |
| GSM4109005 | Sample9 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4109006 | Sample10 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 2.0 |
| GSM4109007 | Sample11 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4109008 | Sample12 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4109009 | Sample13 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 0.0 |
| GSM4109010 | Sample14 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 1.0 |
| GSM4109011 | Sample 15 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 0.0 |
| GSM4109012 | Sample16 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 0.0 |
| ID | GSE139220 |
| Title | Expression data from sevoflurane anesthesia treated rat |
| Organism | Rattus norvegicus |
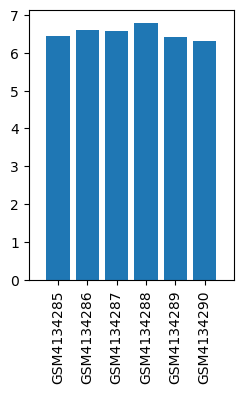
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4134285 | Brain_Control_rep1 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 6.457361 |
| GSM4134286 | Brain_Control_rep2 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 6.609035 |
| GSM4134287 | Brain_Control_rep3 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 6.583948 |
| GSM4134288 | Brain_Sevoflurane_rep1 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 6.788565 |
| GSM4134289 | Brain_Sevoflurane_rep2 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 6.43109 |
| GSM4134290 | Brain_Sevoflurane_rep3 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 6.310688 |
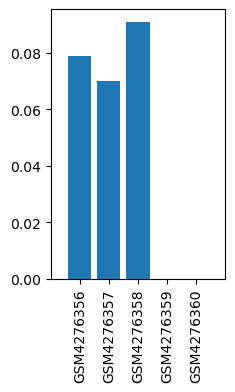
| ID | GSE143895 |
| Title | GPR160 de-orphanization reveals critical roles in neuropathic pain in rodents |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4276356 | dhsc01 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.079 |
| GSM4276357 | dhsc03 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.07 |
| GSM4276358 | dhsc05 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.091 |
| GSM4276359 | dhsc13 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 0.0 |
| GSM4276360 | dhsc15 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 0.0 |
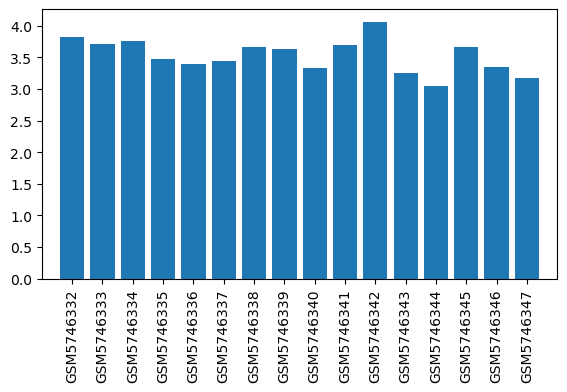
| ID | GSE192399 |
| Title | Gene expression of brain regions activated by sevoflurane and propofol |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5746332 | LHb, Sevoflurane, 60mins | brain region: the lateral habenula (LHb) treatment: Sevoflurane | 3.82072 |
| GSM5746333 | LHb, O2, 60mins | brain region: the lateral habenula (LHb) treatment: O2 | 3.70555 |
| GSM5746334 | MHb, Sevoflurane, 60mins | brain region: the medial habenula (MHb) treatment: Sevoflurane | 3.7617 |
| GSM5746335 | MHb, O2, 60mins | brain region: the medial habenula (MHb) treatment: O2 | 3.47794 |
| GSM5746336 | Sol, Sevoflurane, 60mins | brain region: the solitary nucleus (Sol) treatment: Sevoflurane | 3.40098 |
| GSM5746337 | Sol, O2, 60mins | brain region: the solitary nucleus (Sol) treatment: O2 | 3.43456 |
| GSM5746338 | Mve, Sevoflurane, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Sevoflurane | 3.65543 |
| GSM5746339 | Mve, O2, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: O2 | 3.63667 |
| GSM5746340 | LHb, Propofol, 60mins | brain region: the lateral habenula (LHb) treatment: Propofol | 3.33802 |
| GSM5746341 | LHb, Intralipos, 60mins | brain region: the lateral habenula (LHb) treatment: Intralipos | 3.68802 |
| GSM5746342 | MHb, Propofol, 60mins | brain region: the medial habenula (MHb) treatment: Propofol | 4.05883 |
| GSM5746343 | MHb, Intralipos, 60mins | brain region: the medial habenula (MHb) treatment: Intralipos | 3.24742 |
| GSM5746344 | Sol, Propofol, 60mins | brain region: the solitary nucleus (Sol) treatment: Propofol | 3.04835 |
| GSM5746345 | Sol, Intralipos, 60mins | brain region: the solitary nucleus (Sol) treatment: Intralipos | 3.65916 |
| GSM5746346 | MVe, Propofol, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Propofol | 3.34876 |
| GSM5746347 | MVe, Intralipos, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Intralipos | 3.17367 |
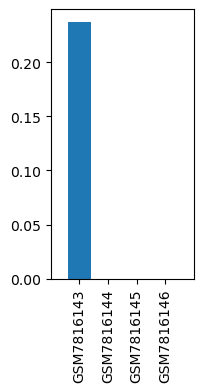
| ID | GSE244436 |
| Title | Similarity and dissimilarity in alteration of gene expression profile associated with inhalational anesthesia between sevoflurane and desflurane |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7816143 | Liver, Control for SevoAnes | tissue: liver genotype: wild type treatment: Control for SevoAnes | 0.23731 |
| GSM7816144 | Liver, Anesthetized with sevoflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with sevoflurane for 6 h | 0.0 |
| GSM7816145 | Liver, Control for DesAnes | tissue: liver genotype: wild type treatment: Control for DesAnes | 0.0 |
| GSM7816146 | Liver, Anesthetized with desflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with desflurane for 6 h | 0.0 |
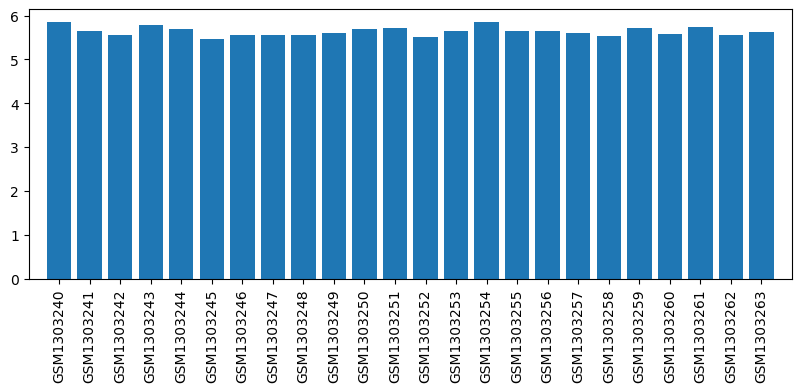
| ID | GSE53897 |
| Title | Genome-wide analysis of dorsal root ganglion (DRG) gene expression after morphine or oxycodone treatment |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1303240 | DRG RVM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.848238 |
| GSM1303241 | DRG RVM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.651389 |
| GSM1303242 | DRG RVM3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.565604 |
| GSM1303243 | DRG RVM7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.788752 |
| GSM1303244 | DRG RVOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.685966 |
| GSM1303245 | DRG RVOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.457109 |
| GSM1303246 | DRG RVOx5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.562052 |
| GSM1303247 | DRG RVOx7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.545462 |
| GSM1303248 | DRG RVS4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.556996 |
| GSM1303249 | DRG RVS5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.598609 |
| GSM1303250 | DRG RVS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.696405 |
| GSM1303251 | DRG RVS7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.724449 |
| GSM1303252 | DRG RSM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.519311 |
| GSM1303253 | DRG RSM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.652723 |
| GSM1303254 | DRG RSM4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.853512 |
| GSM1303255 | DRG RSM5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.644066 |
| GSM1303256 | DRG RSOx1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.636771 |
| GSM1303257 | DRG RSOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.611252 |
| GSM1303258 | DRG RSOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.532203 |
| GSM1303259 | DRG RSOx4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.717812 |
| GSM1303260 | DRG RSS1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.578479 |
| GSM1303261 | DRG RSS2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.735977 |
| GSM1303262 | DRG RSS3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.559193 |
| GSM1303263 | DRG RSS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.633068 |

