GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0001655 | Urogenital system development |
| Process | GO:0001701 | In utero embryonic development |
| Process | GO:0001776 | Leukocyte homeostasis |
| Process | GO:0001780 | Neutrophil homeostasis |
| Process | GO:0001817 | Regulation of cytokine production |
| Process | GO:0001819 | Positive regulation of cytokine production |
| Process | GO:0001822 | Kidney development |
| Process | GO:0002250 | Adaptive immune response |
| Process | GO:0002252 | Immune effector process |
| Process | GO:0002253 | Activation of immune response |
| Process | GO:0002262 | Myeloid cell homeostasis |
| Process | GO:0002376 | Immune system process |
| Process | GO:0002443 | Leukocyte mediated immunity |
| Process | GO:0002449 | Lymphocyte mediated immunity |
| Process | GO:0002455 | Humoral immune response mediated by circulating immunoglobulin |
| Process | GO:0002460 | Adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains |
| Process | GO:0002523 | Leukocyte migration involved in inflammatory response |
| Process | GO:0002682 | Regulation of immune system process |
| Process | GO:0002683 | Negative regulation of immune system process |
| Process | GO:0002684 | Positive regulation of immune system process |
| Process | GO:0002685 | Regulation of leukocyte migration |
| Process | GO:0002686 | Negative regulation of leukocyte migration |
| Process | GO:0002688 | Regulation of leukocyte chemotaxis |
| Process | GO:0002689 | Negative regulation of leukocyte chemotaxis |
| Process | GO:0006873 | Cellular ion homeostasis |
| Process | GO:0006874 | Cellular calcium ion homeostasis |
| Process | GO:0006875 | Cellular metal ion homeostasis |
| Process | GO:0006935 | Chemotaxis |
| Process | GO:0006950 | Response to stress |
| Process | GO:0006952 | Defense response |
| Process | GO:0006954 | Inflammatory response |
| Process | GO:0006955 | Immune response |
| Process | GO:0006956 | Complement activation |
| Process | GO:0006957 | Complement activation, alternative pathway |
| Process | GO:0006958 | Complement activation, classical pathway |
| Process | GO:0006959 | Humoral immune response |
| Process | GO:0007204 | Positive regulation of cytosolic calcium ion concentration |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0009605 | Response to external stimulus |
| Process | GO:0009607 | Response to biotic stimulus |
| Process | GO:0009611 | Response to wounding |
| Process | GO:0009790 | Embryo development |
| Process | GO:0009792 | Embryo development ending in birth or egg hatching |
| Process | GO:0009892 | Negative regulation of metabolic process |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010466 | Negative regulation of peptidase activity |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010574 | Regulation of vascular endothelial growth factor production |
| Process | GO:0010575 | Positive regulation of vascular endothelial growth factor production |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0010605 | Negative regulation of macromolecule metabolic process |
| Process | GO:0010628 | Positive regulation of gene expression |
| Process | GO:0010700 | Negative regulation of norepinephrine secretion |
| Process | GO:0010758 | Regulation of macrophage chemotaxis |
| Process | GO:0010760 | Negative regulation of macrophage chemotaxis |
| Process | GO:0010951 | Negative regulation of endopeptidase activity |
| Process | GO:0014059 | Regulation of dopamine secretion |
| Process | GO:0014061 | Regulation of norepinephrine secretion |
| Process | GO:0016064 | Immunoglobulin mediated immune response |
| Process | GO:0016477 | Cell migration |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0019724 | B cell mediated immunity |
| Process | GO:0019725 | Cellular homeostasis |
| Process | GO:0019835 | Cytolysis |
| Process | GO:0022603 | Regulation of anatomical structure morphogenesis |
| Process | GO:0030003 | Cellular cation homeostasis |
| Process | GO:0030162 | Regulation of proteolysis |
| Process | GO:0030334 | Regulation of cell migration |
| Process | GO:0030336 | Negative regulation of cell migration |
| Process | GO:0032101 | Regulation of response to external stimulus |
| Process | GO:0032102 | Negative regulation of response to external stimulus |
| Process | GO:0032103 | Positive regulation of response to external stimulus |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032642 | Regulation of chemokine production |
| Process | GO:0032722 | Positive regulation of chemokine production |
| Process | GO:0032835 | Glomerulus development |
| Process | GO:0032879 | Regulation of localization |
| Process | GO:0033500 | Carbohydrate homeostasis |
| Process | GO:0033602 | Negative regulation of dopamine secretion |
| Process | GO:0033604 | Negative regulation of catecholamine secretion |
| Process | GO:0040011 | Locomotion |
| Process | GO:0040012 | Regulation of locomotion |
| Process | GO:0040013 | Negative regulation of locomotion |
| Process | GO:0040017 | Positive regulation of locomotion |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0042330 | Taxis |
| Process | GO:0042592 | Homeostatic process |
| Process | GO:0042593 | Glucose homeostasis |
| Process | GO:0043009 | Chordate embryonic development |
| Process | GO:0043086 | Negative regulation of catalytic activity |
| Process | GO:0043207 | Response to external biotic stimulus |
| Process | GO:0043269 | Regulation of ion transport |
| Process | GO:0043271 | Negative regulation of ion transport |
| Process | GO:0044092 | Negative regulation of molecular function |
| Process | GO:0044419 | Biological process involved in interspecies interaction between organisms |
| Process | GO:0045087 | Innate immune response |
| Process | GO:0045765 | Regulation of angiogenesis |
| Process | GO:0045766 | Positive regulation of angiogenesis |
| Process | GO:0045861 | Negative regulation of proteolysis |
| Process | GO:0048513 | Animal organ development |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048519 | Negative regulation of biological process |
| Process | GO:0048523 | Negative regulation of cellular process |
| Process | GO:0048583 | Regulation of response to stimulus |
| Process | GO:0048584 | Positive regulation of response to stimulus |
| Process | GO:0048585 | Negative regulation of response to stimulus |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0048870 | Cell motility |
| Process | GO:0048872 | Homeostasis of number of cells |
| Process | GO:0048878 | Chemical homeostasis |
| Process | GO:0050433 | Regulation of catecholamine secretion |
| Process | GO:0050776 | Regulation of immune response |
| Process | GO:0050778 | Positive regulation of immune response |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050790 | Regulation of catalytic activity |
| Process | GO:0050793 | Regulation of developmental process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050801 | Ion homeostasis |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0050900 | Leukocyte migration |
| Process | GO:0050920 | Regulation of chemotaxis |
| Process | GO:0050921 | Positive regulation of chemotaxis |
| Process | GO:0050922 | Negative regulation of chemotaxis |
| Process | GO:0051046 | Regulation of secretion |
| Process | GO:0051048 | Negative regulation of secretion |
| Process | GO:0051049 | Regulation of transport |
| Process | GO:0051051 | Negative regulation of transport |
| Process | GO:0051094 | Positive regulation of developmental process |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051172 | Negative regulation of nitrogen compound metabolic process |
| Process | GO:0051239 | Regulation of multicellular organismal process |
| Process | GO:0051240 | Positive regulation of multicellular organismal process |
| Process | GO:0051246 | Regulation of protein metabolic process |
| Process | GO:0051248 | Negative regulation of protein metabolic process |
| Process | GO:0051336 | Regulation of hydrolase activity |
| Process | GO:0051346 | Negative regulation of hydrolase activity |
| Process | GO:0051707 | Response to other organism |
| Process | GO:0051952 | Regulation of amine transport |
| Process | GO:0051953 | Negative regulation of amine transport |
| Process | GO:0052547 | Regulation of peptidase activity |
| Process | GO:0052548 | Regulation of endopeptidase activity |
| Process | GO:0055065 | Metal ion homeostasis |
| Process | GO:0055074 | Calcium ion homeostasis |
| Process | GO:0055080 | Cation homeostasis |
| Process | GO:0055082 | Cellular chemical homeostasis |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065008 | Regulation of biological quality |
| Process | GO:0065009 | Regulation of molecular function |
| Process | GO:0072001 | Renal system development |
| Process | GO:0072006 | Nephron development |
| Process | GO:0072503 | Cellular divalent inorganic cation homeostasis |
| Process | GO:0072507 | Divalent inorganic cation homeostasis |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0090594 | Inflammatory response to wounding |
| Process | GO:0097273 | Creatinine homeostasis |
| Process | GO:0098542 | Defense response to other organism |
| Process | GO:0098771 | Inorganic ion homeostasis |
| Process | GO:1901342 | Regulation of vasculature development |
| Process | GO:1903530 | Regulation of secretion by cell |
| Process | GO:1903531 | Negative regulation of secretion by cell |
| Process | GO:1904018 | Positive regulation of vasculature development |
| Process | GO:1905521 | Regulation of macrophage migration |
| Process | GO:1905522 | Negative regulation of macrophage migration |
| Process | GO:2000026 | Regulation of multicellular organismal development |
| Process | GO:2000145 | Regulation of cell motility |
| Process | GO:2000146 | Negative regulation of cell motility |
| Component | GO:0005576 | Extracellular region |
| Component | GO:0005579 | Membrane attack complex |
| Component | GO:0005615 | Extracellular space |
| Component | GO:0005886 | Plasma membrane |
| Component | GO:0005887 | Integral component of plasma membrane |
| Component | GO:0016020 | Membrane |
| Component | GO:0016021 | Integral component of membrane |
| Component | GO:0031224 | Intrinsic component of membrane |
| Component | GO:0031226 | Intrinsic component of plasma membrane |
| Component | GO:0032991 | Protein-containing complex |
| Component | GO:0046930 | Pore complex |
| Component | GO:0071944 | Cell periphery |
| Component | GO:0098796 | Membrane protein complex |
| Component | GO:0098797 | Plasma membrane protein complex |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0001664 | G protein-coupled receptor binding |
| Function | GO:0004857 | Enzyme inhibitor activity |
| Function | GO:0004866 | Endopeptidase inhibitor activity |
| Function | GO:0005102 | Signaling receptor binding |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0030234 | Enzyme regulator activity |
| Function | GO:0030414 | Peptidase inhibitor activity |
| Function | GO:0031714 | C5a anaphylatoxin chemotactic receptor binding |
| Function | GO:0061134 | Peptidase regulator activity |
| Function | GO:0061135 | Endopeptidase regulator activity |
| Function | GO:0098772 | Molecular function regulator activity |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| WikiPathways | WP2433 | Spinal cord injury |
| KEGG | rno04080 | Neuroactive ligand-receptor interaction |
| KEGG | rno04610 | Complement and coagulation cascades |
| KEGG | rno04613 | Neutrophil extracellular trap formation |
| KEGG | rno04810 | Regulation of actin cytoskeleton |
| KEGG | rno04936 | Alcoholic liver disease |
| KEGG | rno05020 | Prion disease |
| KEGG | rno05133 | Pertussis |
| KEGG | rno05150 | Staphylococcus aureus infection |
| KEGG | rno05168 | Herpes simplex virus 1 infection |
| KEGG | rno05171 | Coronavirus disease - COVID-19 |
| KEGG | rno05322 | Systemic lupus erythematosus |
Expression profile of C5 in omics data
| ID | GSE100499 |
| Title | Exploring acupuncture mechanism in preconditioning of IRI |
| Organism | Rattus norvegicus |
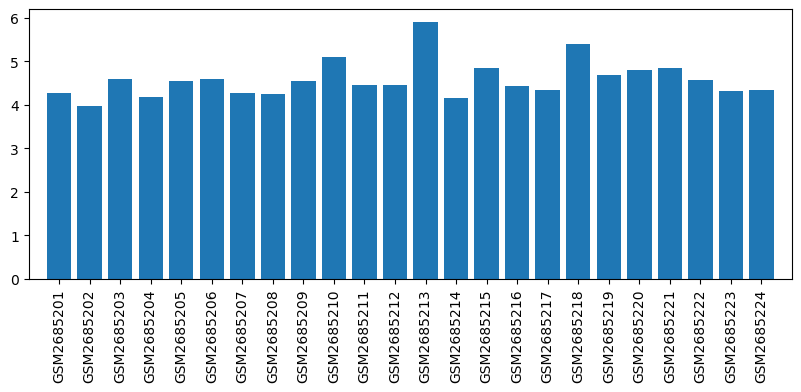
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2685201 | MX1 | tissue: Myocardium gender: male | 4.2644987 |
| GSM2685202 | MX2 | tissue: Myocardium gender: female | 3.9609184 |
| GSM2685203 | MX3 | tissue: Myocardium gender: female | 4.5994506 |
| GSM2685204 | MX4 | tissue: Myocardium gender: male | 4.1812134 |
| GSM2685205 | DZ1 | tissue: Myocardium gender: male | 4.542303 |
| GSM2685206 | DZ2 | tissue: Myocardium gender: male | 4.6008987 |
| GSM2685207 | DZ3 | tissue: Myocardium gender: female | 4.274541 |
| GSM2685208 | DZ4 | tissue: Myocardium gender: female | 4.257958 |
| GSM2685209 | JJ1 | tissue: Myocardium gender: male | 4.549946 |
| GSM2685210 | JJ2 | tissue: Myocardium gender: female | 5.0963874 |
| GSM2685211 | JJ3 | tissue: Myocardium gender: female | 4.461265 |
| GSM2685212 | JJ4 | tissue: Myocardium gender: male | 4.4599295 |
| GSM2685213 | NG1 | tissue: Myocardium gender: male | 5.9051437 |
| GSM2685214 | NG2 | tissue: Myocardium gender: female | 4.1470304 |
| GSM2685215 | NG3 | tissue: Myocardium gender: male | 4.839599 |
| GSM2685216 | NG4 | tissue: Myocardium gender: female | 4.4329686 |
| GSM2685217 | YL1 | tissue: Myocardium gender: male | 4.3516073 |
| GSM2685218 | YL2 | tissue: Myocardium gender: male | 5.3903003 |
| GSM2685219 | YL3 | tissue: Myocardium gender: female | 4.6931686 |
| GSM2685220 | YL4 | tissue: Myocardium gender: female | 4.7997046 |
| GSM2685221 | QC1 | tissue: Myocardium gender: male | 4.8531923 |
| GSM2685222 | QC2 | tissue: Myocardium gender: male | 4.575076 |
| GSM2685223 | QC3 | tissue: Myocardium gender: female | 4.3191004 |
| GSM2685224 | QC4 | tissue: Myocardium gender: female | 4.349281 |
| ID | GSE136833 |
| Title | Effects of lidocaine on the expression of rat spinal cord |
| Organism | Rattus norvegicus |
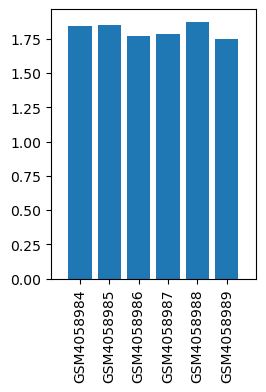
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4058984 | Control_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 1.840143462 |
| GSM4058985 | Control_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 1.852951241 |
| GSM4058986 | Control_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 1.771573046 |
| GSM4058987 | Lidocaine_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 1.786045777 |
| GSM4058988 | Lidocaine_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 1.872961414 |
| GSM4058989 | Lidocaine_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 1.750187338 |
| ID | GSE139220 |
| Title | Expression data from sevoflurane anesthesia treated rat |
| Organism | Rattus norvegicus |
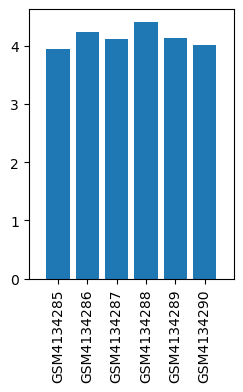
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4134285 | Brain_Control_rep1 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 3.954015 |
| GSM4134286 | Brain_Control_rep2 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 4.23808 |
| GSM4134287 | Brain_Control_rep3 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 4.11619 |
| GSM4134288 | Brain_Sevoflurane_rep1 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 4.412125 |
| GSM4134289 | Brain_Sevoflurane_rep2 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 4.139254 |
| GSM4134290 | Brain_Sevoflurane_rep3 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 4.01146 |
| ID | GSE205202 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [rat] |
| Organism | Rattus norvegicus |
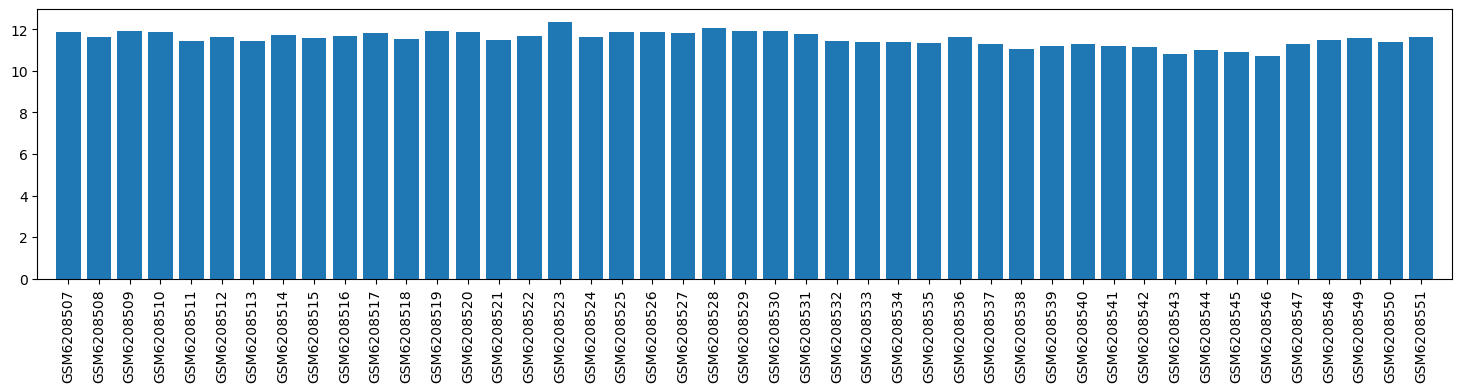
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208507 | Rat_Baseline_0h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 11.85082 |
| GSM6208508 | Rat_Baseline_0h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 11.61389 |
| GSM6208509 | Rat_Baseline_0h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 11.92355 |
| GSM6208510 | Rat_Baseline_0h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 11.89044 |
| GSM6208511 | Rat_Baseline_0h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 11.43537 |
| GSM6208512 | Rat_Vehicle_03h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.65193 |
| GSM6208513 | Rat_Vehicle_03h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.43752 |
| GSM6208514 | Rat_Vehicle_03h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.71639 |
| GSM6208515 | Rat_Vehicle_03h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.5863 |
| GSM6208516 | Rat_Vehicle_03h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.66903 |
| GSM6208517 | Rat_Vehicle_06h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.84601 |
| GSM6208518 | Rat_Vehicle_06h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.55521 |
| GSM6208519 | Rat_Vehicle_06h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.91783 |
| GSM6208520 | Rat_Vehicle_06h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.85208 |
| GSM6208521 | Rat_Vehicle_06h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.48482 |
| GSM6208522 | Rat_Vehicle_09h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.66237 |
| GSM6208523 | Rat_Vehicle_09h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 12.35313 |
| GSM6208524 | Rat_Vehicle_09h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.63226 |
| GSM6208525 | Rat_Vehicle_09h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.8665 |
| GSM6208526 | Rat_Vehicle_09h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.85227 |
| GSM6208527 | Rat_Vehicle_24h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 11.83181 |
| GSM6208528 | Rat_Vehicle_24h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 12.05671 |
| GSM6208529 | Rat_Vehicle_24h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 11.92893 |
| GSM6208530 | Rat_Vehicle_24h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 11.90984 |
| GSM6208531 | Rat_Vehicle_24h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 11.79556 |
| GSM6208532 | Rat_APAP_03h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 11.44258 |
| GSM6208533 | Rat_APAP_03h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 11.37001 |
| GSM6208534 | Rat_APAP_03h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 11.38005 |
| GSM6208535 | Rat_APAP_03h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 11.33857 |
| GSM6208536 | Rat_APAP_03h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 11.62448 |
| GSM6208537 | Rat_APAP_06h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 11.31483 |
| GSM6208538 | Rat_APAP_06h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 11.05874 |
| GSM6208539 | Rat_APAP_06h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 11.21603 |
| GSM6208540 | Rat_APAP_06h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 11.29992 |
| GSM6208541 | Rat_APAP_06h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 11.20885 |
| GSM6208542 | Rat_APAP_09h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 11.15213 |
| GSM6208543 | Rat_APAP_09h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.78958 |
| GSM6208544 | Rat_APAP_09h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.99763 |
| GSM6208545 | Rat_APAP_09h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.90836 |
| GSM6208546 | Rat_APAP_09h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.70404 |
| GSM6208547 | Rat_APAP_24h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.29389 |
| GSM6208548 | Rat_APAP_24h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.49957 |
| GSM6208549 | Rat_APAP_24h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.5991 |
| GSM6208550 | Rat_APAP_24h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.40933 |
| GSM6208551 | Rat_APAP_24h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.64217 |
| ID | GSE55573 |
| Title | Using DNA microarray to screen intracellular signal pathways of hypoglycemic activity by electroacupuncture of Zusanli (ST36) acupoints in streptozotocin-induced rats with diabetes |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1339443 | Mock skeletal muscle | strain: Wistar age: 8-10 week-old gender: Male | 126.20948999999999 |
| GSM1339444 | EA (electroacupuncture) skeletal muscle | strain: Wistar age: 8-10 week-old gender: Male | 124.07785 |
| ID | GSE245201 |
| Title | Transcriptional Reprogramming of Brain Cells Under Anesthesia in wt and Alzheimer's Disease mouse model |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7847823 | 1_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847824 | 2_APP/PS1_Wake | tissue: Hippocampus genotype: APP/PS1 treatment: Wake | |
| GSM7847825 | 3_WT_Isofluran | tissue: Hippocampus genotype: WT treatment: Isofluran | |
| GSM7847826 | 4_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847827 | 10_APP/PS1_Wake | tissue: Hippocampus genotype: APP/PS1 treatment: Wake | |
| GSM7847828 | 11_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847829 | 12_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847830 | 13_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847831 | 14_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847832 | 15_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847833 | 16_WT_Isofluran | tissue: Hippocampus genotype: WT treatment: Isofluran | |
| GSM7847834 | 17_APP/PS1_Wake | tissue: Hippocampus genotype: APP/PS1 treatment: Wake | |
| GSM7847835 | 18_WT_Isofluran | tissue: Hippocampus genotype: WT treatment: Isofluran | |
| GSM7847836 | 19_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847837 | 20_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847838 | 25_APP/PS1_Ketamine | tissue: Hippocampus genotype: APP/PS1 treatment: Ketamine | |
| GSM7847839 | 26_APP/PS1_Ketamine | tissue: Hippocampus genotype: APP/PS1 treatment: Ketamine | |
| GSM7847840 | 27_WT_Ketamine | tissue: Hippocampus genotype: WT treatment: Ketamine | |
| GSM7847841 | 28_WT_Ketamine | tissue: Hippocampus genotype: WT treatment: Ketamine | |
| GSM7847842 | 29_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake |
| ID | GSE245201 |
| Title | Transcriptional Reprogramming of Brain Cells Under Anesthesia in wt and Alzheimer's Disease mouse model |
| Organism | Mus musculus |

| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7847823 | 1_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847824 | 2_APP/PS1_Wake | tissue: Hippocampus genotype: APP/PS1 treatment: Wake | |
| GSM7847825 | 3_WT_Isofluran | tissue: Hippocampus genotype: WT treatment: Isofluran | |
| GSM7847826 | 4_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847827 | 10_APP/PS1_Wake | tissue: Hippocampus genotype: APP/PS1 treatment: Wake | |
| GSM7847828 | 11_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847829 | 12_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847830 | 13_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847831 | 14_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847832 | 15_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847833 | 16_WT_Isofluran | tissue: Hippocampus genotype: WT treatment: Isofluran | |
| GSM7847834 | 17_APP/PS1_Wake | tissue: Hippocampus genotype: APP/PS1 treatment: Wake | |
| GSM7847835 | 18_WT_Isofluran | tissue: Hippocampus genotype: WT treatment: Isofluran | |
| GSM7847836 | 19_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake | |
| GSM7847837 | 20_APP/PS1_Isofluran | tissue: Hippocampus genotype: APP/PS1 treatment: Isofluran | |
| GSM7847838 | 25_APP/PS1_Ketamine | tissue: Hippocampus genotype: APP/PS1 treatment: Ketamine | |
| GSM7847839 | 26_APP/PS1_Ketamine | tissue: Hippocampus genotype: APP/PS1 treatment: Ketamine | |
| GSM7847840 | 27_WT_Ketamine | tissue: Hippocampus genotype: WT treatment: Ketamine | |
| GSM7847841 | 28_WT_Ketamine | tissue: Hippocampus genotype: WT treatment: Ketamine | |
| GSM7847842 | 29_WT_Wake | tissue: Hippocampus genotype: WT treatment: Wake |

