GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0006629 | Lipid metabolic process |
| Process | GO:0007154 | Cell communication |
| Process | GO:0007165 | Signal transduction |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0010628 | Positive regulation of gene expression |
| Process | GO:0010954 | Positive regulation of protein processing |
| Process | GO:0019216 | Regulation of lipid metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0021700 | Developmental maturation |
| Process | GO:0023052 | Signaling |
| Process | GO:0030154 | Cell differentiation |
| Process | GO:0030162 | Regulation of proteolysis |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0042592 | Homeostatic process |
| Process | GO:0043086 | Negative regulation of catalytic activity |
| Process | GO:0044092 | Negative regulation of molecular function |
| Process | GO:0044237 | Cellular metabolic process |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0044255 | Cellular lipid metabolic process |
| Process | GO:0045444 | Fat cell differentiation |
| Process | GO:0045862 | Positive regulation of proteolysis |
| Process | GO:0048468 | Cell development |
| Process | GO:0048469 | Cell maturation |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0048869 | Cellular developmental process |
| Process | GO:0048878 | Chemical homeostasis |
| Process | GO:0050746 | Regulation of lipoprotein metabolic process |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050790 | Regulation of catalytic activity |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0051004 | Regulation of lipoprotein lipase activity |
| Process | GO:0051005 | Negative regulation of lipoprotein lipase activity |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051173 | Positive regulation of nitrogen compound metabolic process |
| Process | GO:0051246 | Regulation of protein metabolic process |
| Process | GO:0051247 | Positive regulation of protein metabolic process |
| Process | GO:0051336 | Regulation of hydrolase activity |
| Process | GO:0051346 | Negative regulation of hydrolase activity |
| Process | GO:0051716 | Cellular response to stimulus |
| Process | GO:0055088 | Lipid homeostasis |
| Process | GO:0055090 | Acylglycerol homeostasis |
| Process | GO:0060191 | Regulation of lipase activity |
| Process | GO:0060192 | Negative regulation of lipase activity |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065008 | Regulation of biological quality |
| Process | GO:0065009 | Regulation of molecular function |
| Process | GO:0070328 | Triglyceride homeostasis |
| Process | GO:0070613 | Regulation of protein processing |
| Process | GO:0071695 | Anatomical structure maturation |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:1903317 | Regulation of protein maturation |
| Process | GO:1903319 | Positive regulation of protein maturation |
| Component | GO:0005576 | Extracellular region |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0005102 | Signaling receptor binding |
| Function | GO:0005179 | Hormone activity |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0030545 | Signaling receptor regulator activity |
| Function | GO:0030546 | Signaling receptor activator activity |
| Function | GO:0048018 | Receptor ligand activity |
| Function | GO:0098772 | Molecular function regulator activity |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| RCTM | MMU-174824 | Plasma lipoprotein assembly, remodeling, and clearance |
| RCTM | MMU-382551 | Transport of small molecules |
| RCTM | MMU-8963889 | Assembly of active LPL and LIPC lipase complexes |
| RCTM | MMU-8963899 | Plasma lipoprotein remodeling |
| KEGG | mmu04979 | Cholesterol metabolism |
Expression profile of Angptl8 in omics data
| ID | GSE109165 |
| Title | Suppression of RGSz1 function optimizes the actions of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway |
| Organism | Mus musculus |
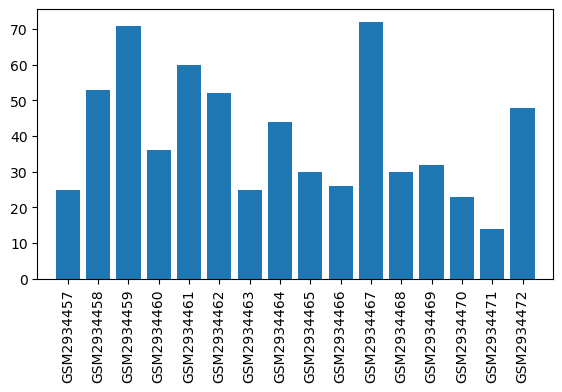
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2934457 | WT_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 25.0 |
| GSM2934458 | WT_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 53.0 |
| GSM2934459 | WT_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 71.0 |
| GSM2934460 | WT_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 36.0 |
| GSM2934461 | KO_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 60.0 |
| GSM2934462 | KO_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 52.0 |
| GSM2934463 | KO_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 25.0 |
| GSM2934464 | KO_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 44.0 |
| GSM2934465 | WT_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 30.0 |
| GSM2934466 | WT_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 26.0 |
| GSM2934467 | WT_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 72.0 |
| GSM2934468 | WT_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 30.0 |
| GSM2934469 | KO_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 32.0 |
| GSM2934470 | KO_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 23.0 |
| GSM2934471 | KO_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 14.0 |
| GSM2934472 | KO_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 48.0 |
| ID | GSE205201 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [mouse] |
| Organism | Mus musculus |
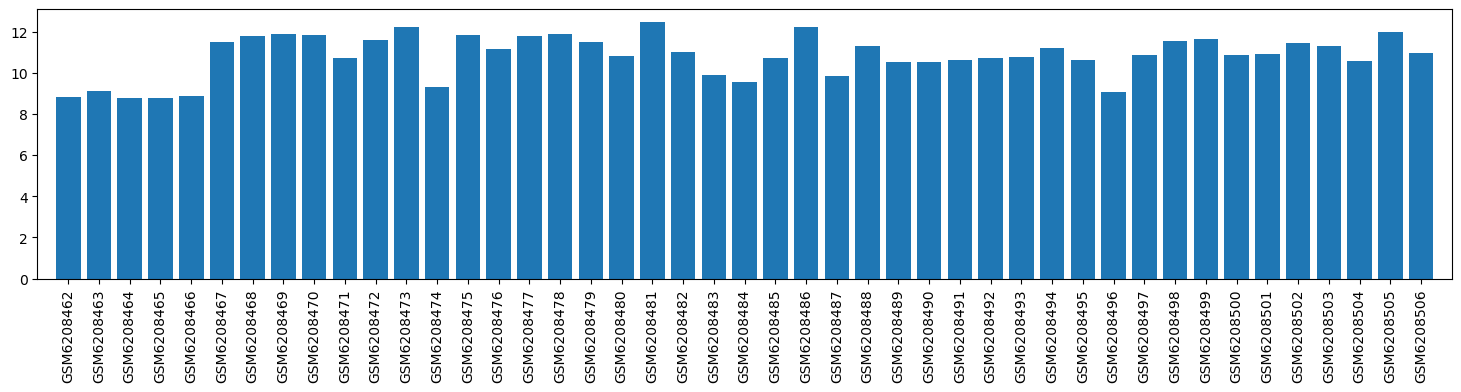
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208462 | Mouse_Baseline_0h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.82491 |
| GSM6208463 | Mouse_Baseline_0h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 9.110324 |
| GSM6208464 | Mouse_Baseline_0h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.785923 |
| GSM6208465 | Mouse_Baseline_0h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.773776 |
| GSM6208466 | Mouse_Baseline_0h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.876376 |
| GSM6208467 | Mouse_Vehicle_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.467117 |
| GSM6208468 | Mouse_Vehicle_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.767739 |
| GSM6208469 | Mouse_Vehicle_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.890827 |
| GSM6208470 | Mouse_Vehicle_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 11.835215 |
| GSM6208471 | Mouse_Vehicle_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 10.700607 |
| GSM6208472 | Mouse_Vehicle_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.565396 |
| GSM6208473 | Mouse_Vehicle_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 12.203166 |
| GSM6208474 | Mouse_Vehicle_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 9.30652 |
| GSM6208475 | Mouse_Vehicle_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.854845 |
| GSM6208476 | Mouse_Vehicle_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 11.167183 |
| GSM6208477 | Mouse_Vehicle_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.793583 |
| GSM6208478 | Mouse_Vehicle_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.895704 |
| GSM6208479 | Mouse_Vehicle_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 11.512185 |
| GSM6208480 | Mouse_Vehicle_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 10.818287 |
| GSM6208481 | Mouse_Vehicle_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 12.462568 |
| GSM6208482 | Mouse_Vehicle_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 10.992562 |
| GSM6208483 | Mouse_Vehicle_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 9.89067 |
| GSM6208484 | Mouse_Vehicle_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 9.540933 |
| GSM6208485 | Mouse_Vehicle_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 10.726744 |
| GSM6208486 | Mouse_Vehicle_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 12.203492 |
| GSM6208487 | Mouse_APAP_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 9.858638 |
| GSM6208488 | Mouse_APAP_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 11.298261 |
| GSM6208489 | Mouse_APAP_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 10.526203 |
| GSM6208490 | Mouse_APAP_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 10.506255 |
| GSM6208491 | Mouse_APAP_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 10.636904 |
| GSM6208492 | Mouse_APAP_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 10.73139 |
| GSM6208493 | Mouse_APAP_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 10.743199 |
| GSM6208494 | Mouse_APAP_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 11.215814 |
| GSM6208495 | Mouse_APAP_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 10.621399 |
| GSM6208496 | Mouse_APAP_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 9.079881 |
| GSM6208497 | Mouse_APAP_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.868337 |
| GSM6208498 | Mouse_APAP_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 11.539703 |
| GSM6208499 | Mouse_APAP_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 11.62756 |
| GSM6208500 | Mouse_APAP_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.88168 |
| GSM6208501 | Mouse_APAP_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 10.912539 |
| GSM6208502 | Mouse_APAP_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.425129 |
| GSM6208503 | Mouse_APAP_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.272624 |
| GSM6208504 | Mouse_APAP_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 10.555348 |
| GSM6208505 | Mouse_APAP_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 11.960985 |
| GSM6208506 | Mouse_APAP_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 10.971051 |
| ID | GSE223541 |
| Title | Oxycodone withdrawal induces HDAC1/2-dependent transcriptional maladaptations in the reward pathway in a murine model of peripheral nerve injury |
| Organism | Mus musculus |

| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6958536 | B1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sa | 0.0 |
| GSM6958537 | C1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958538 | F1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958539 | H1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| GSM6958540 | B2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 0.0 |
| GSM6958541 | C2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958542 | F2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958543 | H2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| GSM6958544 | B3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 0.0 |
| GSM6958545 | C3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958546 | F3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958547 | H3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| GSM6958548 | B4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 0.0 |
| GSM6958549 | C4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958550 | F4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958551 | H4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 0.0 |
| ID | GSE26364 |
| Title | Gene expression changes in alpha GABA receptors in the mice brains after prolonged ketamine administration |
| Organism | Mus musculus |
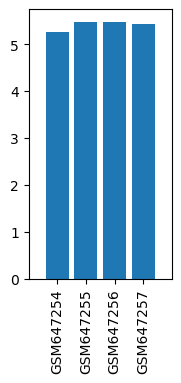
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM647254 | Saline PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 5.25025 |
| GSM647255 | Saline PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 5.47197 |
| GSM647256 | Ketamine PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 5.47416 |
| GSM647257 | Ketamine PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 5.41898 |

