GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0001525 | Angiogenesis |
| Process | GO:0001568 | Blood vessel development |
| Process | GO:0001666 | Response to hypoxia |
| Process | GO:0001817 | Regulation of cytokine production |
| Process | GO:0001819 | Positive regulation of cytokine production |
| Process | GO:0001932 | Regulation of protein phosphorylation |
| Process | GO:0001934 | Positive regulation of protein phosphorylation |
| Process | GO:0001936 | Regulation of endothelial cell proliferation |
| Process | GO:0001938 | Positive regulation of endothelial cell proliferation |
| Process | GO:0001944 | Vasculature development |
| Process | GO:0002040 | Sprouting angiogenesis |
| Process | GO:0002682 | Regulation of immune system process |
| Process | GO:0002684 | Positive regulation of immune system process |
| Process | GO:0002685 | Regulation of leukocyte migration |
| Process | GO:0002687 | Positive regulation of leukocyte migration |
| Process | GO:0002688 | Regulation of leukocyte chemotaxis |
| Process | GO:0002690 | Positive regulation of leukocyte chemotaxis |
| Process | GO:0006935 | Chemotaxis |
| Process | GO:0006950 | Response to stress |
| Process | GO:0007154 | Cell communication |
| Process | GO:0007165 | Signal transduction |
| Process | GO:0007166 | Cell surface receptor signaling pathway |
| Process | GO:0007167 | Enzyme-linked receptor protein signaling pathway |
| Process | GO:0007169 | Transmembrane receptor protein tyrosine kinase signaling pathway |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0007399 | Nervous system development |
| Process | GO:0008283 | Cell population proliferation |
| Process | GO:0008284 | Positive regulation of cell population proliferation |
| Process | GO:0009605 | Response to external stimulus |
| Process | GO:0009607 | Response to biotic stimulus |
| Process | GO:0009617 | Response to bacterium |
| Process | GO:0009628 | Response to abiotic stimulus |
| Process | GO:0009653 | Anatomical structure morphogenesis |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009966 | Regulation of signal transduction |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010033 | Response to organic substance |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010562 | Positive regulation of phosphorus metabolic process |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0010628 | Positive regulation of gene expression |
| Process | GO:0010646 | Regulation of cell communication |
| Process | GO:0019220 | Regulation of phosphate metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0022008 | Neurogenesis |
| Process | GO:0022603 | Regulation of anatomical structure morphogenesis |
| Process | GO:0023051 | Regulation of signaling |
| Process | GO:0023052 | Signaling |
| Process | GO:0030154 | Cell differentiation |
| Process | GO:0030182 | Neuron differentiation |
| Process | GO:0030334 | Regulation of cell migration |
| Process | GO:0030335 | Positive regulation of cell migration |
| Process | GO:0030947 | Regulation of vascular endothelial growth factor receptor signaling pathway |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0031325 | Positive regulation of cellular metabolic process |
| Process | GO:0031399 | Regulation of protein modification process |
| Process | GO:0031401 | Positive regulation of protein modification process |
| Process | GO:0032101 | Regulation of response to external stimulus |
| Process | GO:0032103 | Positive regulation of response to external stimulus |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032675 | Regulation of interleukin-6 production |
| Process | GO:0032755 | Positive regulation of interleukin-6 production |
| Process | GO:0035239 | Tube morphogenesis |
| Process | GO:0035295 | Tube development |
| Process | GO:0035924 | Cellular response to vascular endothelial growth factor stimulus |
| Process | GO:0036293 | Response to decreased oxygen levels |
| Process | GO:0038084 | Vascular endothelial growth factor signaling pathway |
| Process | GO:0040011 | Locomotion |
| Process | GO:0040012 | Regulation of locomotion |
| Process | GO:0040017 | Positive regulation of locomotion |
| Process | GO:0042127 | Regulation of cell population proliferation |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0042325 | Regulation of phosphorylation |
| Process | GO:0042327 | Positive regulation of phosphorylation |
| Process | GO:0042330 | Taxis |
| Process | GO:0043207 | Response to external biotic stimulus |
| Process | GO:0044419 | Biological process involved in interspecies interaction between organisms |
| Process | GO:0045765 | Regulation of angiogenesis |
| Process | GO:0045766 | Positive regulation of angiogenesis |
| Process | GO:0045937 | Positive regulation of phosphate metabolic process |
| Process | GO:0048010 | Vascular endothelial growth factor receptor signaling pathway |
| Process | GO:0048144 | Fibroblast proliferation |
| Process | GO:0048514 | Blood vessel morphogenesis |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048522 | Positive regulation of cellular process |
| Process | GO:0048583 | Regulation of response to stimulus |
| Process | GO:0048584 | Positive regulation of response to stimulus |
| Process | GO:0048646 | Anatomical structure formation involved in morphogenesis |
| Process | GO:0048699 | Generation of neurons |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0048869 | Cellular developmental process |
| Process | GO:0050678 | Regulation of epithelial cell proliferation |
| Process | GO:0050679 | Positive regulation of epithelial cell proliferation |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050793 | Regulation of developmental process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0050918 | Positive chemotaxis |
| Process | GO:0050920 | Regulation of chemotaxis |
| Process | GO:0050921 | Positive regulation of chemotaxis |
| Process | GO:0050926 | Regulation of positive chemotaxis |
| Process | GO:0050927 | Positive regulation of positive chemotaxis |
| Process | GO:0050930 | Induction of positive chemotaxis |
| Process | GO:0051094 | Positive regulation of developmental process |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051173 | Positive regulation of nitrogen compound metabolic process |
| Process | GO:0051174 | Regulation of phosphorus metabolic process |
| Process | GO:0051239 | Regulation of multicellular organismal process |
| Process | GO:0051240 | Positive regulation of multicellular organismal process |
| Process | GO:0051246 | Regulation of protein metabolic process |
| Process | GO:0051247 | Positive regulation of protein metabolic process |
| Process | GO:0051302 | Regulation of cell division |
| Process | GO:0051707 | Response to other organism |
| Process | GO:0051716 | Cellular response to stimulus |
| Process | GO:0051781 | Positive regulation of cell division |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0060753 | Regulation of mast cell chemotaxis |
| Process | GO:0060754 | Positive regulation of mast cell chemotaxis |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0070482 | Response to oxygen levels |
| Process | GO:0070848 | Response to growth factor |
| Process | GO:0070887 | Cellular response to chemical stimulus |
| Process | GO:0071310 | Cellular response to organic substance |
| Process | GO:0071363 | Cellular response to growth factor stimulus |
| Process | GO:0071542 | Dopaminergic neuron differentiation |
| Process | GO:0072359 | Circulatory system development |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0090287 | Regulation of cellular response to growth factor stimulus |
| Process | GO:1901342 | Regulation of vasculature development |
| Process | GO:1904018 | Positive regulation of vasculature development |
| Process | GO:2000026 | Regulation of multicellular organismal development |
| Process | GO:2000145 | Regulation of cell motility |
| Process | GO:2000147 | Positive regulation of cell motility |
| Component | GO:0005576 | Extracellular region |
| Component | GO:0005615 | Extracellular space |
| Component | GO:0016020 | Membrane |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0005102 | Signaling receptor binding |
| Function | GO:0005126 | Cytokine receptor binding |
| Function | GO:0005172 | Vascular endothelial growth factor receptor binding |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0008083 | Growth factor activity |
| Function | GO:0030545 | Signaling receptor regulator activity |
| Function | GO:0030546 | Signaling receptor activator activity |
| Function | GO:0042056 | Chemoattractant activity |
| Function | GO:0042802 | Identical protein binding |
| Function | GO:0043185 | Vascular endothelial growth factor receptor 3 binding |
| Function | GO:0048018 | Receptor ligand activity |
| Function | GO:0070851 | Growth factor receptor binding |
| Function | GO:0098772 | Molecular function regulator activity |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| RCTM | MMU-109582 | Hemostasis |
| RCTM | MMU-114608 | Platelet degranulation |
| RCTM | MMU-162582 | Signal Transduction |
| RCTM | MMU-194138 | Signaling by VEGF |
| RCTM | MMU-194313 | VEGF ligand-receptor interactions |
| RCTM | MMU-195399 | VEGF binds to VEGFR leading to receptor dimerization |
| RCTM | MMU-76002 | Platelet activation, signaling and aggregation |
| RCTM | MMU-76005 | Response to elevated platelet cytosolic Ca2+ |
| RCTM | MMU-9006934 | Signaling by Receptor Tyrosine Kinases |
| WikiPathways | WP2841 | Focal adhesion: PI3K-Akt-mTOR signaling pathway |
| WikiPathways | WP85 | Focal adhesion |
| KEGG | mmu04010 | MAPK signaling pathway |
| KEGG | mmu04014 | Ras signaling pathway |
| KEGG | mmu04015 | Rap1 signaling pathway |
| KEGG | mmu04020 | Calcium signaling pathway |
| KEGG | mmu04151 | PI3K-Akt signaling pathway |
| KEGG | mmu04510 | Focal adhesion |
| KEGG | mmu04668 | TNF signaling pathway |
| KEGG | mmu04926 | Relaxin signaling pathway |
| KEGG | mmu04933 | AGE-RAGE signaling pathway in diabetic complications |
| KEGG | mmu05200 | Pathways in cancer |
Expression profile of Vegfd in omics data
| ID | GSE109165 |
| Title | Suppression of RGSz1 function optimizes the actions of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway |
| Organism | Mus musculus |
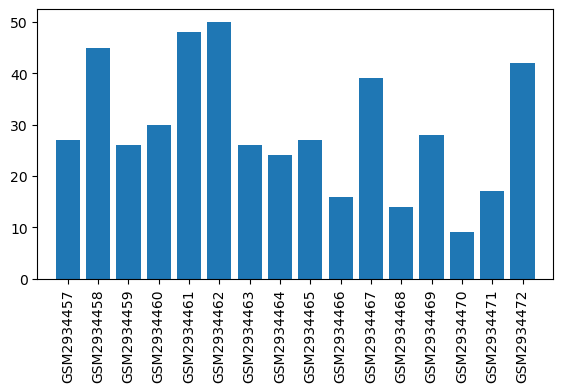
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2934457 | WT_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 27.0 |
| GSM2934458 | WT_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 45.0 |
| GSM2934459 | WT_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 26.0 |
| GSM2934460 | WT_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 30.0 |
| GSM2934461 | KO_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 48.0 |
| GSM2934462 | KO_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 50.0 |
| GSM2934463 | KO_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 26.0 |
| GSM2934464 | KO_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 24.0 |
| GSM2934465 | WT_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 27.0 |
| GSM2934466 | WT_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 16.0 |
| GSM2934467 | WT_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 39.0 |
| GSM2934468 | WT_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 14.0 |
| GSM2934469 | KO_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 28.0 |
| GSM2934470 | KO_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 9.0 |
| GSM2934471 | KO_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 17.0 |
| GSM2934472 | KO_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 42.0 |
| ID | GSE126662 |
| Title | Comparison between opioid (morphine) and vehicle treatments in mouse trigeminal ganglia and nucleus accumbens |
| Organism | Mus musculus |
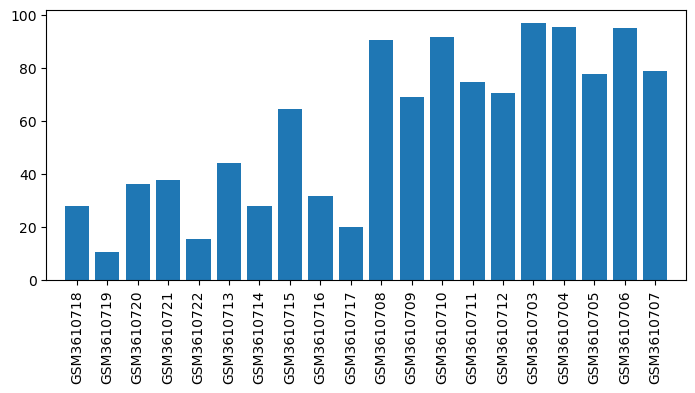
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM3610703 | TG_M1: OIH_M1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 96.9624 |
| GSM3610704 | TG_M2: OIH_M2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 95.4091 |
| GSM3610705 | TG_M3: OIH_M3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 77.8732 |
| GSM3610706 | TG_M4: OIH_M4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 94.94494999999999 |
| GSM3610707 | TG_M5: OIH_M5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 78.70645 |
| GSM3610708 | TG_V1: VehicleOIH_V1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 90.38995 |
| GSM3610709 | TG_V2: VehicleOIH_V2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 68.96645 |
| GSM3610710 | TG_V3: VehicleOIH_V3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 91.65315 |
| GSM3610711 | TG_V4: VehicleOIH_V4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 74.69785 |
| GSM3610712 | TG_V5: VehicleOIH_V5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 70.44315 |
| GSM3610713 | NAc_M1: OIH_M1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 43.9526 |
| GSM3610714 | NAc_M2: OIH_M2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 27.99998 |
| GSM3610715 | NAc_M3: OIH_M3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 64.6542 |
| GSM3610716 | NAc_M4: OIH_M4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 31.5 |
| GSM3610717 | NAc_M5: OIH_M5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 20.0 |
| GSM3610718 | NAc_V1: VehicleOIH_V1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 27.999995 |
| GSM3610719 | NAc_V2: VehicleOIH_V2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 10.57965 |
| GSM3610720 | NAc_V3: VehicleOIH_V3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 36.258095000000004 |
| GSM3610721 | NAc_V4: VehicleOIH_V4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 37.500015 |
| GSM3610722 | NAc_V5: VehicleOIH_V5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 15.231300000000001 |
| ID | GSE205201 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [mouse] |
| Organism | Mus musculus |
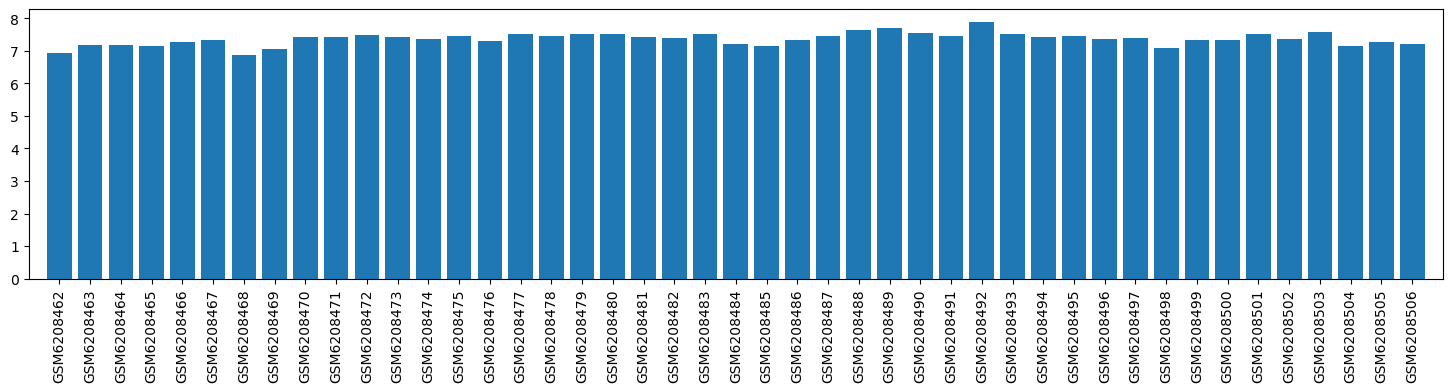
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208462 | Mouse_Baseline_0h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 6.9217 |
| GSM6208463 | Mouse_Baseline_0h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 7.166982 |
| GSM6208464 | Mouse_Baseline_0h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 7.174394 |
| GSM6208465 | Mouse_Baseline_0h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 7.154311 |
| GSM6208466 | Mouse_Baseline_0h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 7.274016 |
| GSM6208467 | Mouse_Vehicle_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.334744 |
| GSM6208468 | Mouse_Vehicle_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 6.884931 |
| GSM6208469 | Mouse_Vehicle_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.069827 |
| GSM6208470 | Mouse_Vehicle_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.439693 |
| GSM6208471 | Mouse_Vehicle_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.420033 |
| GSM6208472 | Mouse_Vehicle_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.474441 |
| GSM6208473 | Mouse_Vehicle_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.413324 |
| GSM6208474 | Mouse_Vehicle_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.360872 |
| GSM6208475 | Mouse_Vehicle_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.448611 |
| GSM6208476 | Mouse_Vehicle_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.315598 |
| GSM6208477 | Mouse_Vehicle_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.507977 |
| GSM6208478 | Mouse_Vehicle_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.447737 |
| GSM6208479 | Mouse_Vehicle_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.505931 |
| GSM6208480 | Mouse_Vehicle_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.529373 |
| GSM6208481 | Mouse_Vehicle_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.418226 |
| GSM6208482 | Mouse_Vehicle_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 7.409987 |
| GSM6208483 | Mouse_Vehicle_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 7.507345 |
| GSM6208484 | Mouse_Vehicle_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 7.197349 |
| GSM6208485 | Mouse_Vehicle_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 7.160123 |
| GSM6208486 | Mouse_Vehicle_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 7.32611 |
| GSM6208487 | Mouse_APAP_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.467089 |
| GSM6208488 | Mouse_APAP_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.632238 |
| GSM6208489 | Mouse_APAP_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.691271 |
| GSM6208490 | Mouse_APAP_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.560472 |
| GSM6208491 | Mouse_APAP_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.469183 |
| GSM6208492 | Mouse_APAP_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 7.887414 |
| GSM6208493 | Mouse_APAP_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 7.52723 |
| GSM6208494 | Mouse_APAP_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 7.425079 |
| GSM6208495 | Mouse_APAP_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 7.44466 |
| GSM6208496 | Mouse_APAP_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 7.367431 |
| GSM6208497 | Mouse_APAP_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.397217 |
| GSM6208498 | Mouse_APAP_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.095736 |
| GSM6208499 | Mouse_APAP_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.320354 |
| GSM6208500 | Mouse_APAP_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.341011 |
| GSM6208501 | Mouse_APAP_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.522354 |
| GSM6208502 | Mouse_APAP_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.376665 |
| GSM6208503 | Mouse_APAP_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.573605 |
| GSM6208504 | Mouse_APAP_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.143611 |
| GSM6208505 | Mouse_APAP_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.262807 |
| GSM6208506 | Mouse_APAP_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.197221 |
| ID | GSE223541 |
| Title | Oxycodone withdrawal induces HDAC1/2-dependent transcriptional maladaptations in the reward pathway in a murine model of peripheral nerve injury |
| Organism | Mus musculus |
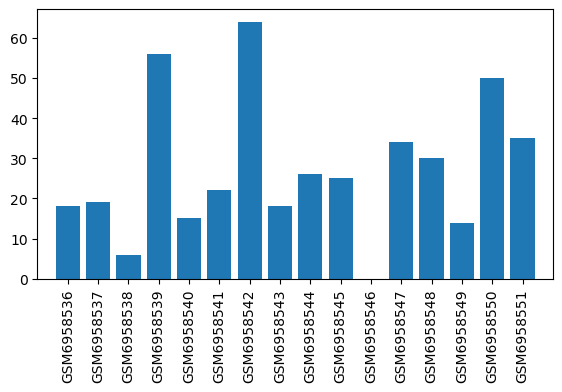
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6958536 | B1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sa | 18.0 |
| GSM6958537 | C1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 19.0 |
| GSM6958538 | F1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 6.0 |
| GSM6958539 | H1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 56.0 |
| GSM6958540 | B2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 15.0 |
| GSM6958541 | C2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 22.0 |
| GSM6958542 | F2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 64.0 |
| GSM6958543 | H2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 18.0 |
| GSM6958544 | B3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 26.0 |
| GSM6958545 | C3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 25.0 |
| GSM6958546 | F3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958547 | H3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 34.0 |
| GSM6958548 | B4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 30.0 |
| GSM6958549 | C4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 14.0 |
| GSM6958550 | F4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 50.0 |
| GSM6958551 | H4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 35.0 |
| ID | GSE26364 |
| Title | Gene expression changes in alpha GABA receptors in the mice brains after prolonged ketamine administration |
| Organism | Mus musculus |
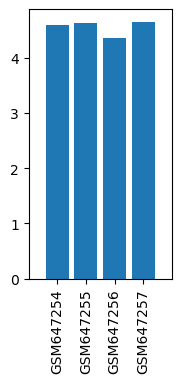
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM647254 | Saline PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 4.59102 |
| GSM647255 | Saline PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 4.62087 |
| GSM647256 | Ketamine PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 4.35316 |
| GSM647257 | Ketamine PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 4.64701 |

