GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0000375 | RNA splicing, via transesterification reactions |
| Process | GO:0000377 | RNA splicing, via transesterification reactions with bulged adenosine as nucleophile |
| Process | GO:0000381 | Regulation of alternative mRNA splicing, via spliceosome |
| Process | GO:0000398 | mRNA splicing, via spliceosome |
| Process | GO:0006139 | Nucleobase-containing compound metabolic process |
| Process | GO:0006396 | RNA processing |
| Process | GO:0006397 | mRNA processing |
| Process | GO:0006725 | Cellular aromatic compound metabolic process |
| Process | GO:0006807 | Nitrogen compound metabolic process |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0007399 | Nervous system development |
| Process | GO:0007417 | Central nervous system development |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0008380 | RNA splicing |
| Process | GO:0009892 | Negative regulation of metabolic process |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010467 | Gene expression |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0016070 | RNA metabolic process |
| Process | GO:0016071 | mRNA metabolic process |
| Process | GO:0019219 | Regulation of nucleobase-containing compound metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0021510 | Spinal cord development |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0034641 | Cellular nitrogen compound metabolic process |
| Process | GO:0043170 | Macromolecule metabolic process |
| Process | GO:0043484 | Regulation of RNA splicing |
| Process | GO:0044237 | Cellular metabolic process |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0046483 | Heterocycle metabolic process |
| Process | GO:0048024 | Regulation of mRNA splicing, via spliceosome |
| Process | GO:0048513 | Animal organ development |
| Process | GO:0048519 | Negative regulation of biological process |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0050684 | Regulation of mRNA processing |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051239 | Regulation of multicellular organismal process |
| Process | GO:0051241 | Negative regulation of multicellular organismal process |
| Process | GO:0051252 | Regulation of RNA metabolic process |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065008 | Regulation of biological quality |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0090304 | Nucleic acid metabolic process |
| Process | GO:0120161 | Regulation of cold-induced thermogenesis |
| Process | GO:0120163 | Negative regulation of cold-induced thermogenesis |
| Process | GO:1901360 | Organic cyclic compound metabolic process |
| Process | GO:1903311 | Regulation of mRNA metabolic process |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005634 | Nucleus |
| Component | GO:0005737 | Cytoplasm |
| Component | GO:0043226 | Organelle |
| Component | GO:0043227 | Membrane-bounded organelle |
| Component | GO:0043229 | Intracellular organelle |
| Component | GO:0043231 | Intracellular membrane-bounded organelle |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0003676 | Nucleic acid binding |
| Function | GO:0003723 | RNA binding |
| Function | GO:0003729 | mRNA binding |
| Function | GO:0003730 | mRNA 3-UTR binding |
| Function | GO:0005488 | Binding |
| Function | GO:0097159 | Organic cyclic compound binding |
| Function | GO:1901363 | Heterocyclic compound binding |
| Function | GO:1990825 | Sequence-specific mRNA binding |
Expression profile of Nova1 in omics data
| ID | GSE100499 |
| Title | Exploring acupuncture mechanism in preconditioning of IRI |
| Organism | Rattus norvegicus |
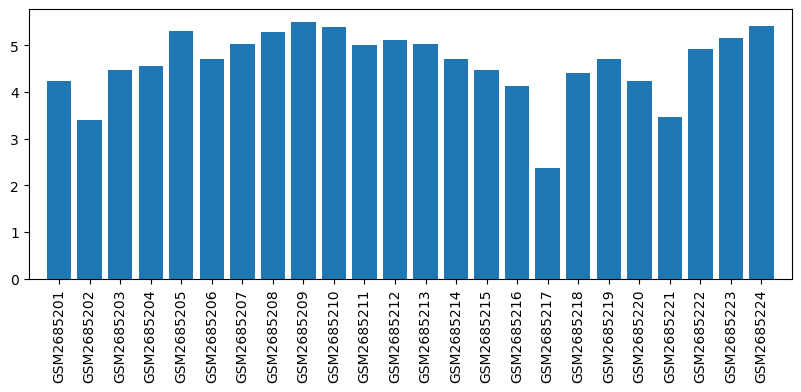
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2685201 | MX1 | tissue: Myocardium gender: male | 4.226157 |
| GSM2685202 | MX2 | tissue: Myocardium gender: female | 3.3906634 |
| GSM2685203 | MX3 | tissue: Myocardium gender: female | 4.459609 |
| GSM2685204 | MX4 | tissue: Myocardium gender: male | 4.554419 |
| GSM2685205 | DZ1 | tissue: Myocardium gender: male | 5.3085012 |
| GSM2685206 | DZ2 | tissue: Myocardium gender: male | 4.7062163 |
| GSM2685207 | DZ3 | tissue: Myocardium gender: female | 5.02386 |
| GSM2685208 | DZ4 | tissue: Myocardium gender: female | 5.277398 |
| GSM2685209 | JJ1 | tissue: Myocardium gender: male | 5.4980865 |
| GSM2685210 | JJ2 | tissue: Myocardium gender: female | 5.381919 |
| GSM2685211 | JJ3 | tissue: Myocardium gender: female | 5.0057645 |
| GSM2685212 | JJ4 | tissue: Myocardium gender: male | 5.106628 |
| GSM2685213 | NG1 | tissue: Myocardium gender: male | 5.0169144 |
| GSM2685214 | NG2 | tissue: Myocardium gender: female | 4.7138777 |
| GSM2685215 | NG3 | tissue: Myocardium gender: male | 4.4649315 |
| GSM2685216 | NG4 | tissue: Myocardium gender: female | 4.121854 |
| GSM2685217 | YL1 | tissue: Myocardium gender: male | 2.3701637 |
| GSM2685218 | YL2 | tissue: Myocardium gender: male | 4.4018936 |
| GSM2685219 | YL3 | tissue: Myocardium gender: female | 4.707291 |
| GSM2685220 | YL4 | tissue: Myocardium gender: female | 4.23035 |
| GSM2685221 | QC1 | tissue: Myocardium gender: male | 3.4646583 |
| GSM2685222 | QC2 | tissue: Myocardium gender: male | 4.9111195 |
| GSM2685223 | QC3 | tissue: Myocardium gender: female | 5.1648183 |
| GSM2685224 | QC4 | tissue: Myocardium gender: female | 5.4041567 |
| ID | GSE136833 |
| Title | Effects of lidocaine on the expression of rat spinal cord |
| Organism | Rattus norvegicus |
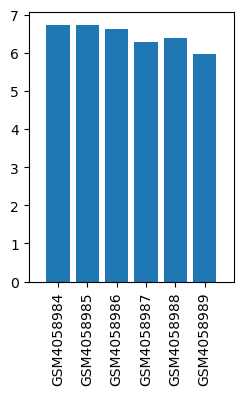
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4058984 | Control_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.715056398 |
| GSM4058985 | Control_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.726529851 |
| GSM4058986 | Control_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.62371424 |
| GSM4058987 | Lidocaine_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.277280585 |
| GSM4058988 | Lidocaine_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 6.373857146 |
| GSM4058989 | Lidocaine_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 5.97857986 |
| ID | GSE138454 |
| Title | Impact of inflammation on hepatic subcellular energetics in anesthetized rats |
| Organism | Rattus norvegicus |
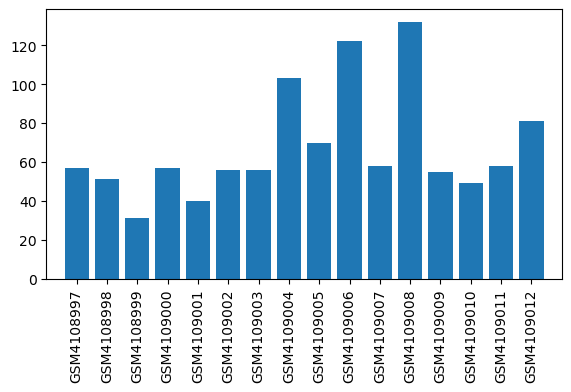
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4108997 | Sample 1 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 57.0 |
| GSM4108998 | Sample 2 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 51.0 |
| GSM4108999 | Sample 3 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 31.0 |
| GSM4109000 | Sample 4 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 57.0 |
| GSM4109001 | Sample 5 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 40.0 |
| GSM4109002 | Sample 6 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 56.0 |
| GSM4109003 | Sample 7 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 56.0 |
| GSM4109004 | Sample 8 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 103.0 |
| GSM4109005 | Sample9 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 70.0 |
| GSM4109006 | Sample10 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 122.0 |
| GSM4109007 | Sample11 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 58.0 |
| GSM4109008 | Sample12 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 132.0 |
| GSM4109009 | Sample13 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 55.0 |
| GSM4109010 | Sample14 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 49.0 |
| GSM4109011 | Sample 15 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 58.0 |
| GSM4109012 | Sample16 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 81.0 |
| ID | GSE139220 |
| Title | Expression data from sevoflurane anesthesia treated rat |
| Organism | Rattus norvegicus |
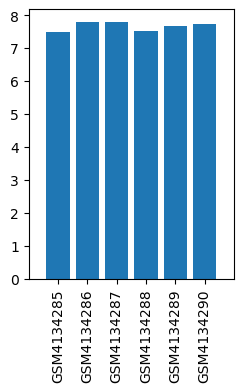
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4134285 | Brain_Control_rep1 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 7.489034 |
| GSM4134286 | Brain_Control_rep2 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 7.78331 |
| GSM4134287 | Brain_Control_rep3 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 7.794203 |
| GSM4134288 | Brain_Sevoflurane_rep1 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 7.531178 |
| GSM4134289 | Brain_Sevoflurane_rep2 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 7.662706 |
| GSM4134290 | Brain_Sevoflurane_rep3 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 7.736571 |
| ID | GSE143895 |
| Title | GPR160 de-orphanization reveals critical roles in neuropathic pain in rodents |
| Organism | Rattus norvegicus |
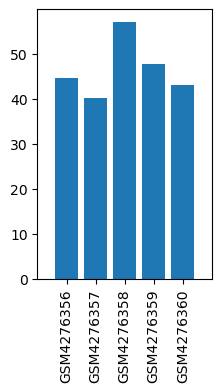
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4276356 | dhsc01 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 44.504 |
| GSM4276357 | dhsc03 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 40.266 |
| GSM4276358 | dhsc05 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 57.06 |
| GSM4276359 | dhsc13 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 47.768 |
| GSM4276360 | dhsc15 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 43.157 |
| ID | GSE192399 |
| Title | Gene expression of brain regions activated by sevoflurane and propofol |
| Organism | Rattus norvegicus |
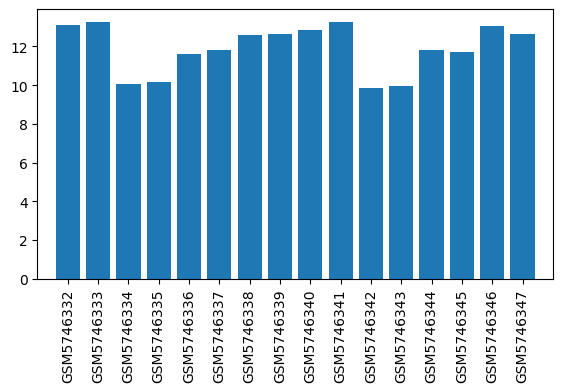
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5746332 | LHb, Sevoflurane, 60mins | brain region: the lateral habenula (LHb) treatment: Sevoflurane | 13.1031 |
| GSM5746333 | LHb, O2, 60mins | brain region: the lateral habenula (LHb) treatment: O2 | 13.2652 |
| GSM5746334 | MHb, Sevoflurane, 60mins | brain region: the medial habenula (MHb) treatment: Sevoflurane | 10.0489 |
| GSM5746335 | MHb, O2, 60mins | brain region: the medial habenula (MHb) treatment: O2 | 10.1804 |
| GSM5746336 | Sol, Sevoflurane, 60mins | brain region: the solitary nucleus (Sol) treatment: Sevoflurane | 11.5901 |
| GSM5746337 | Sol, O2, 60mins | brain region: the solitary nucleus (Sol) treatment: O2 | 11.8049 |
| GSM5746338 | Mve, Sevoflurane, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Sevoflurane | 12.5789 |
| GSM5746339 | Mve, O2, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: O2 | 12.6317 |
| GSM5746340 | LHb, Propofol, 60mins | brain region: the lateral habenula (LHb) treatment: Propofol | 12.8675 |
| GSM5746341 | LHb, Intralipos, 60mins | brain region: the lateral habenula (LHb) treatment: Intralipos | 13.251 |
| GSM5746342 | MHb, Propofol, 60mins | brain region: the medial habenula (MHb) treatment: Propofol | 9.8297 |
| GSM5746343 | MHb, Intralipos, 60mins | brain region: the medial habenula (MHb) treatment: Intralipos | 9.95543 |
| GSM5746344 | Sol, Propofol, 60mins | brain region: the solitary nucleus (Sol) treatment: Propofol | 11.8048 |
| GSM5746345 | Sol, Intralipos, 60mins | brain region: the solitary nucleus (Sol) treatment: Intralipos | 11.7216 |
| GSM5746346 | MVe, Propofol, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Propofol | 13.07 |
| GSM5746347 | MVe, Intralipos, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Intralipos | 12.653 |
| ID | GSE232804 |
| Title | EGR3 regulates opioid-related nociception and motivation in male rats |
| Organism | Rattus norvegicus |
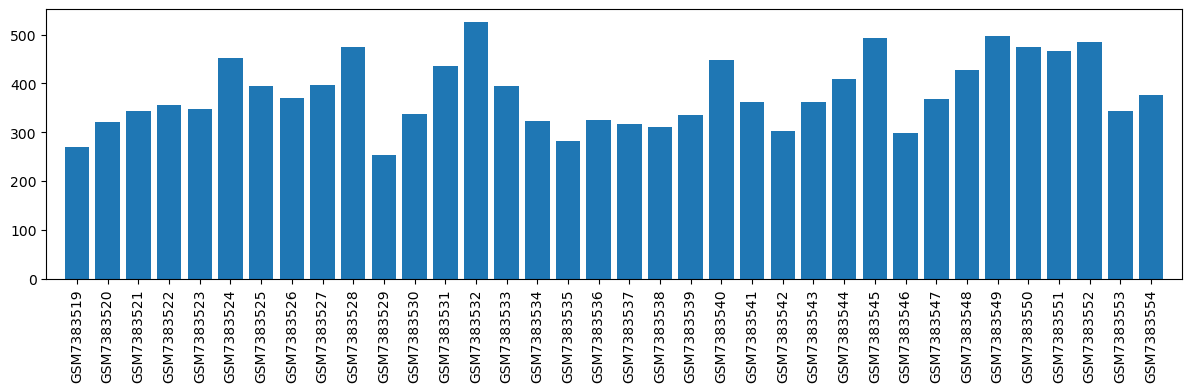
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7383519 | ww01_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Salin | 269.0 |
| GSM7383520 | ww02_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Saline | 321.0 |
| GSM7383521 | ww03_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Salin | 344.0 |
| GSM7383522 | ww05_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Saline | 356.0 |
| GSM7383523 | ww06_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Salin | 348.0 |
| GSM7383524 | ww07_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Saline | 453.0 |
| GSM7383525 | ww08_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Salin | 395.0 |
| GSM7383526 | ww10_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Saline | 370.0 |
| GSM7383527 | ww12_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 396.0 |
| GSM7383528 | ww13_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 474.0 |
| GSM7383529 | ww14_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 254.0 |
| GSM7383530 | ww15_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 338.0 |
| GSM7383531 | ww16_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 435.0 |
| GSM7383532 | ww17_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 526.0 |
| GSM7383533 | ww18_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 395.0 |
| GSM7383534 | ww20_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 324.0 |
| GSM7383535 | ww21_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 283.0 |
| GSM7383536 | ww22_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 326.0 |
| GSM7383537 | ww23_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 317.0 |
| GSM7383538 | ww24_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 310.0 |
| GSM7383539 | ww26_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 336.0 |
| GSM7383540 | ww27_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 449.0 |
| GSM7383541 | ww28_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 362.0 |
| GSM7383542 | ww30_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 302.0 |
| GSM7383543 | ww31_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 363.0 |
| GSM7383544 | ww32_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 409.0 |
| GSM7383545 | ww34_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 493.0 |
| GSM7383546 | ww35_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 298.0 |
| GSM7383547 | ww36_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 368.0 |
| GSM7383548 | ww37_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 427.0 |
| GSM7383549 | ww38_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 498.0 |
| GSM7383550 | www09_Saline_Saline | tissue: Brain / mPFC pain: Saline drug: Saline | 474.0 |
| GSM7383551 | www19_Saline_OXY | tissue: Brain / mPFC pain: Saline drug: OXY | 466.0 |
| GSM7383552 | www29_CFA_Saline | tissue: Brain / mPFC pain: CFA drug: Saline | 484.0 |
| GSM7383553 | www33_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 343.0 |
| GSM7383554 | www39_CFA_OXY | tissue: Brain / mPFC pain: CFA drug: OXY | 377.0 |
| ID | GSE244436 |
| Title | Similarity and dissimilarity in alteration of gene expression profile associated with inhalational anesthesia between sevoflurane and desflurane |
| Organism | Rattus norvegicus |
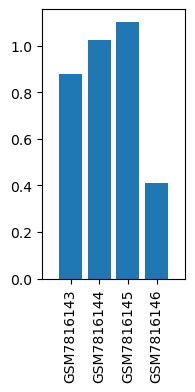
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7816143 | Liver, Control for SevoAnes | tissue: liver genotype: wild type treatment: Control for SevoAnes | 0.879075 |
| GSM7816144 | Liver, Anesthetized with sevoflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with sevoflurane for 6 h | 1.024049 |
| GSM7816145 | Liver, Control for DesAnes | tissue: liver genotype: wild type treatment: Control for DesAnes | 1.101806 |
| GSM7816146 | Liver, Anesthetized with desflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with desflurane for 6 h | 0.412238 |
| ID | GSE2982 |
| Title | Effect of inhaled anesthetic to cultured cortical neurons |
| Organism | Rattus norvegicus |
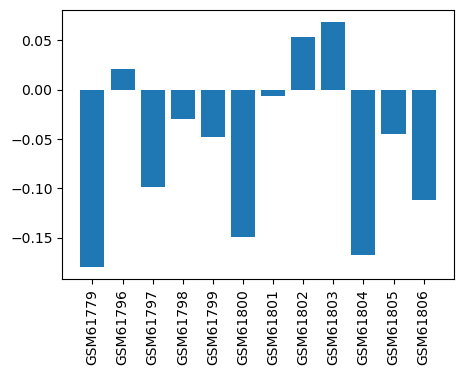
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM61779 | Cell Culture Exposure (control1) | Cy3 label;Cy5; | -0.18 |
| GSM61796 | Cell Culture Exposure (control2) | Cy3; Cy5 | 0.0214 |
| GSM61797 | Cell Culture Exposure (control3) | Cy3; Cy5 | -0.0985 |
| GSM61798 | Cell Culture Exposure (1MAC Halo1) | Cy3; Cy5 | -0.0292 |
| GSM61799 | Cell Culture Exposure (1MAC Halo2) | Cy3; Cy5 | -0.0483 |
| GSM61800 | Cell Culture Exposure (1MAC Halo3) | Cy3; Cy5 | -0.149 |
| GSM61801 | Cell Culture Exposure (3MAC Halo1) | Cy3; Cy5 | -0.00583 |
| GSM61802 | Cell Culture Exposure (3MAC Halo2) | Cy3; Cy5 | 0.0535 |
| GSM61803 | Cell Culture Exposure (3MAC Halo3) | Cy3; Cy5 | 0.0686 |
| GSM61804 | Cell Culture Exposure (3MAC ISO1) | Cy3; Cy5 | -0.168 |
| GSM61805 | Cell Culture Exposure (3MAC ISO2) | Cy3; Cy5 | -0.0445 |
| GSM61806 | Cell Culture Exposure (3MAC ISO3) | Cy3; Cy5 | -0.112 |
| ID | GSE53897 |
| Title | Genome-wide analysis of dorsal root ganglion (DRG) gene expression after morphine or oxycodone treatment |
| Organism | Rattus norvegicus |
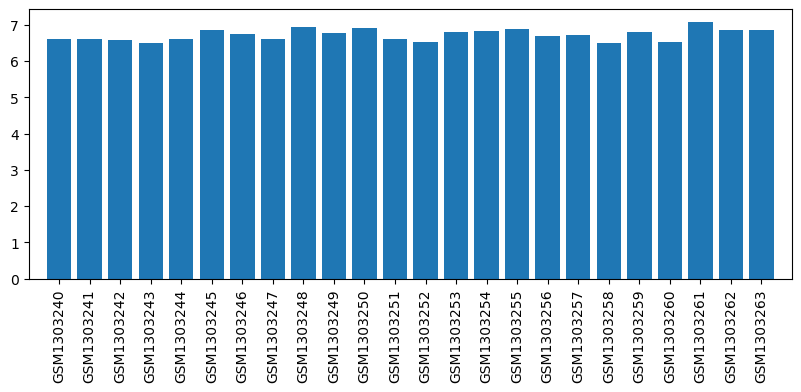
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1303240 | DRG RVM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.609609 |
| GSM1303241 | DRG RVM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.617867 |
| GSM1303242 | DRG RVM3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.582419 |
| GSM1303243 | DRG RVM7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.48812 |
| GSM1303244 | DRG RVOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.619108 |
| GSM1303245 | DRG RVOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.847206 |
| GSM1303246 | DRG RVOx5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.742211 |
| GSM1303247 | DRG RVOx7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.61373 |
| GSM1303248 | DRG RVS4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.930413 |
| GSM1303249 | DRG RVS5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.762799 |
| GSM1303250 | DRG RVS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.922582 |
| GSM1303251 | DRG RVS7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.603728 |
| GSM1303252 | DRG RSM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.513164 |
| GSM1303253 | DRG RSM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.796594 |
| GSM1303254 | DRG RSM4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.826217 |
| GSM1303255 | DRG RSM5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.895822 |
| GSM1303256 | DRG RSOx1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.702743 |
| GSM1303257 | DRG RSOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.720526 |
| GSM1303258 | DRG RSOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.493901 |
| GSM1303259 | DRG RSOx4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.803527 |
| GSM1303260 | DRG RSS1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.513821 |
| GSM1303261 | DRG RSS2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 7.077507 |
| GSM1303262 | DRG RSS3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.861835 |
| GSM1303263 | DRG RSS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 6.843575 |
| ID | GSE55573 |
| Title | Using DNA microarray to screen intracellular signal pathways of hypoglycemic activity by electroacupuncture of Zusanli (ST36) acupoints in streptozotocin-induced rats with diabetes |
| Organism | Rattus norvegicus |
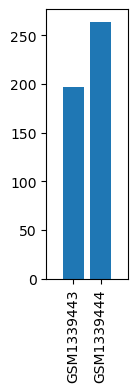
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1339443 | Mock skeletal muscle | strain: Wistar age: 8-10 week-old gender: Male | 196.7114 |
| GSM1339444 | EA (electroacupuncture) skeletal muscle | strain: Wistar age: 8-10 week-old gender: Male | 263.8223 |

