Gene Information
|
Gene Name
|
FGFR2 |
|
Gene ID
|
2263
|
|
Gene Full Name
|
fibroblast growth factor receptor 2 |
|
Gene Alias
|
BBDS|BEK|BFR-1|CD332|CEK3|CFD1|ECT1|JWS|K-SAM|KGFR|TK14|TK25 |
|
Transcripts
|
ENSG00000066468
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Cell Junctions;Nucleoplasm, Vesicles;Secreted - unknown location; 
|
|
Membrane Info
|
Cancer-related genes, CD markers, Disease related genes, Enzymes, FDA approved drug targets, Human disease related genes, Plasma proteins, Predicted intracellular proteins, Predicted membrane proteins, Predicted secreted proteins, RAS pathway related proteins |
|
Uniport_ID
|
P21802
|
|
HGNC ID
|
HGNC:3689
|
|
OMIM ID
|
176943 |
|
Summary
|
The protein encoded by this gene is a member of the fibroblast growth factor receptor family, where amino acid sequence is highly conserved between members and throughout evolution. FGFR family members differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region, composed of three immunoglobulin-like domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. This particular family member is a high-affinity receptor for acidic, basic and/or keratinocyte growth factor, depending on the isoform. Mutations in this gene are associated with Crouzon syndrome, Pfeiffer syndrome, Craniosynostosis, Apert syndrome, Jackson-Weiss syndrome, Beare-Stevenson cutis gyrata syndrome, Saethre-Chotzen syndrome, and syndromic craniosynostosis. Multiple alternatively spliced transcript variants encoding different isoforms have been noted for this gene. [provided by RefSeq, Jan 2009] |
Target gene [FGFR2] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1001957 |
chr10 |
246 |
101 |
49 |
396 |
View |
| 1006111 |
chr10 |
3 |
30 |
10 |
43 |
View |
Target gene [FGFR2] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
E GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
When the query gene is differentially changed in the dataset, a volcano/bar plot will be displayed.
> Dataset: GSE100400 - FGFR2 expression across samples
|
Volcano Plot
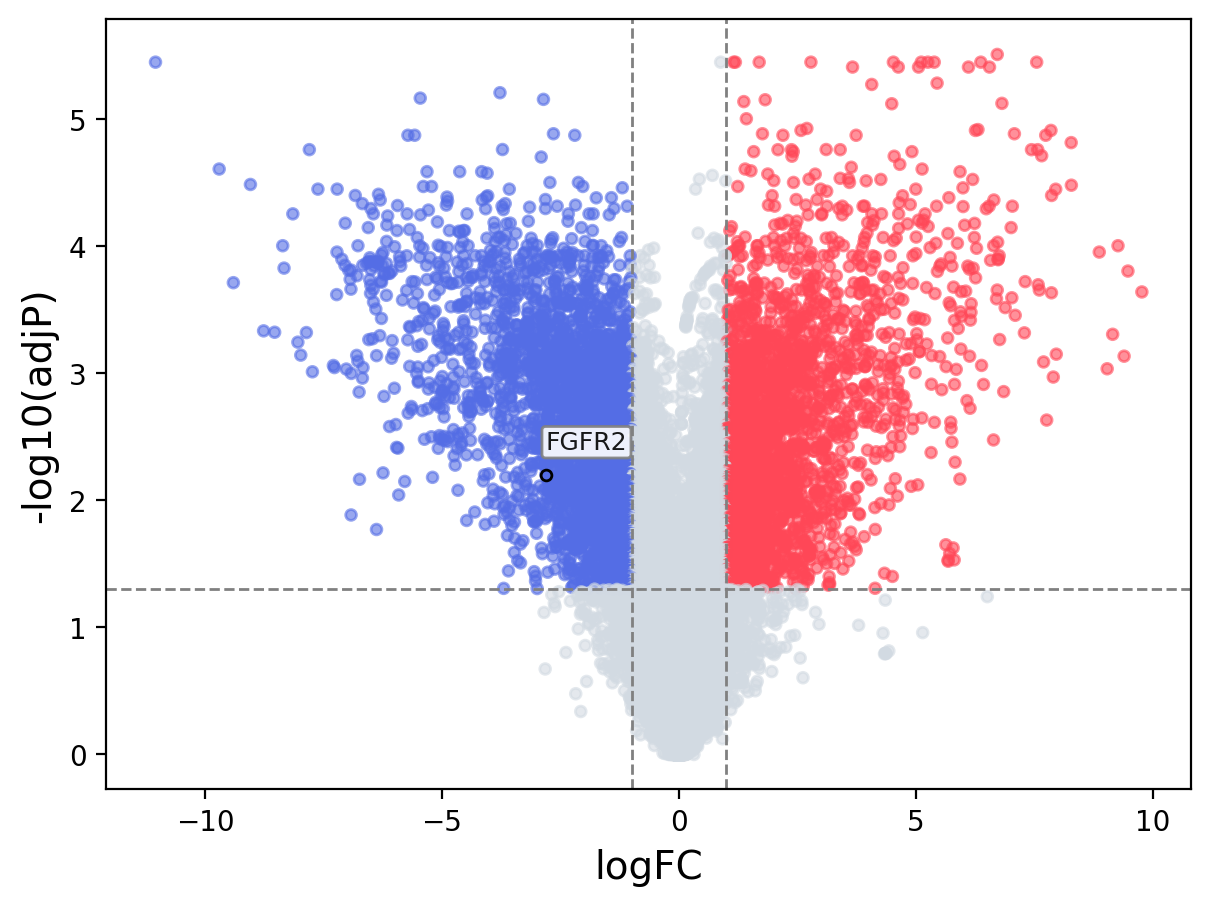
|
Bar Plot
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information