Gene Information
|
Gene Name
|
MET |
|
Gene ID
|
4233
|
|
Gene Full Name
|
MET proto-oncogene, receptor tyrosine kinase |
|
Gene Alias
|
AUTS9|DA11|DFNB97|HGFR|RCCP2|c-Met |
|
Transcripts
|
ENSG00000105976
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Plasma membrane;Cytosol;Secreted to blood; 
|
|
Membrane Info
|
Cancer-related genes, Disease related genes, Enzymes, FDA approved drug targets, Human disease related genes, Plasma proteins, Predicted intracellular proteins, Predicted membrane proteins, Predicted secreted proteins, RAS pathway related proteins, Transporters |
|
Uniport_ID
|
P08581
|
|
HGNC ID
|
HGNC:7029
|
|
OMIM ID
|
164860 |
|
Summary
|
This gene encodes a member of the receptor tyrosine kinase family of proteins and the product of the proto-oncogene MET. The encoded preproprotein is proteolytically processed to generate alpha and beta subunits that are linked via disulfide bonds to form the mature receptor. Further processing of the beta subunit results in the formation of the M10 peptide, which has been shown to reduce lung fibrosis. Binding of its ligand, hepatocyte growth factor, induces dimerization and activation of the receptor, which plays a role in cellular survival, embryogenesis, and cellular migration and invasion. Mutations in this gene are associated with papillary renal cell carcinoma, hepatocellular carcinoma, and various head and neck cancers. Amplification and overexpression of this gene are also associated with multiple human cancers. [provided by RefSeq, May 2016] |
Target gene [MET] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1009601 |
chr7 |
1 |
14 |
0 |
15 |
View |
Target gene [MET] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
GSE199850
|
scRNA-seq |
1 |
HiSeq X Ten (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
When the gene can detect a peak in the dataset, a peak plot will be displayed.
> Dataset: GSE35465 - MET peak across samples
|
Peak Plot
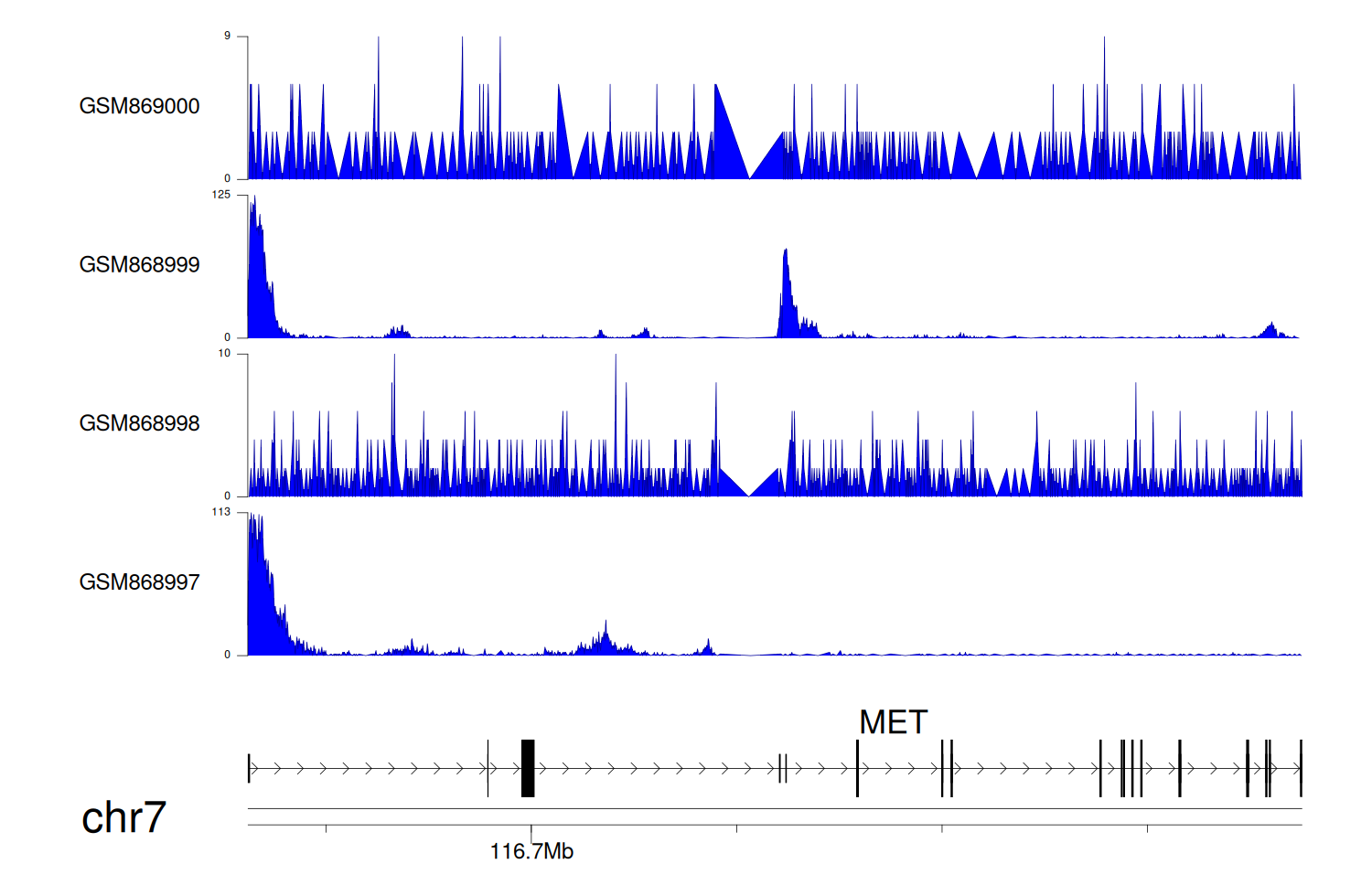
|
> Dataset: GSE68402 - MET peak across samples
|
Peak Plot
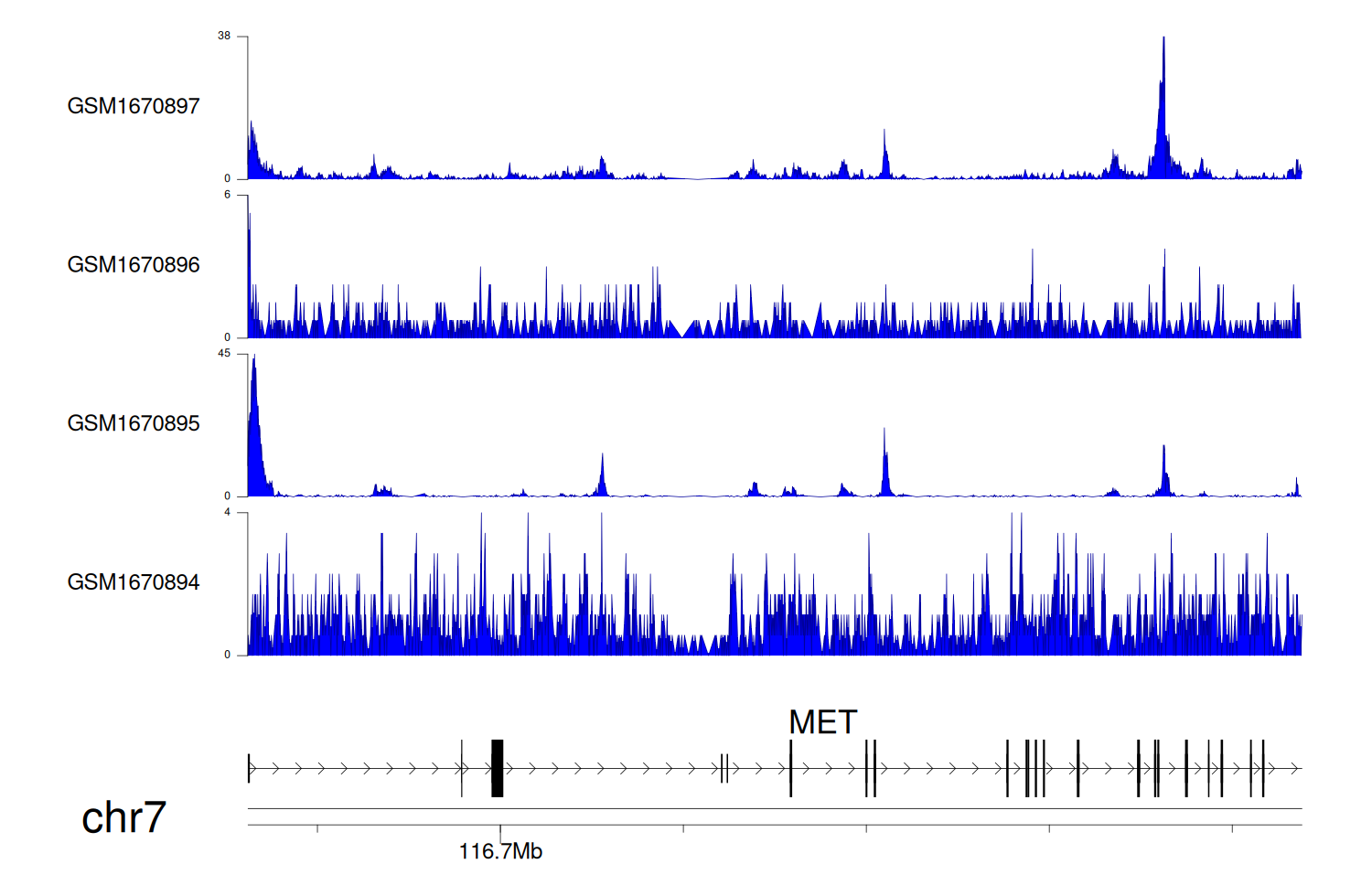
|
> Dataset: GSE270130 - MET peak across samples
|
Peak Plot
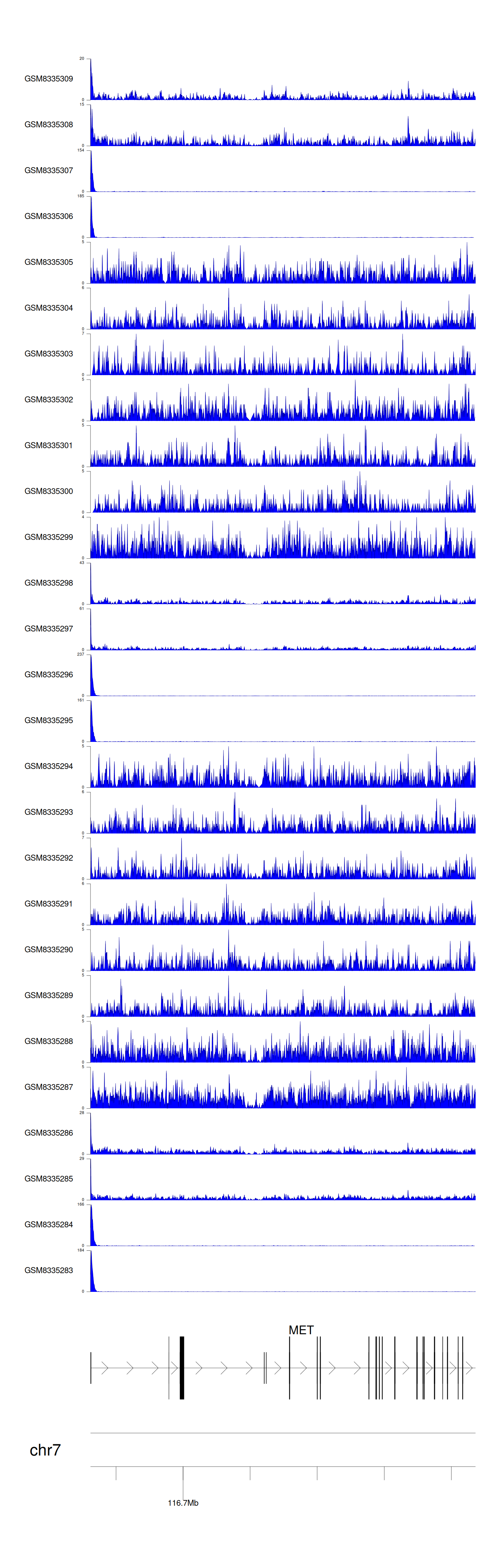
|
> Dataset: GSE131257 - MET peak across samples
|
Peak Plot
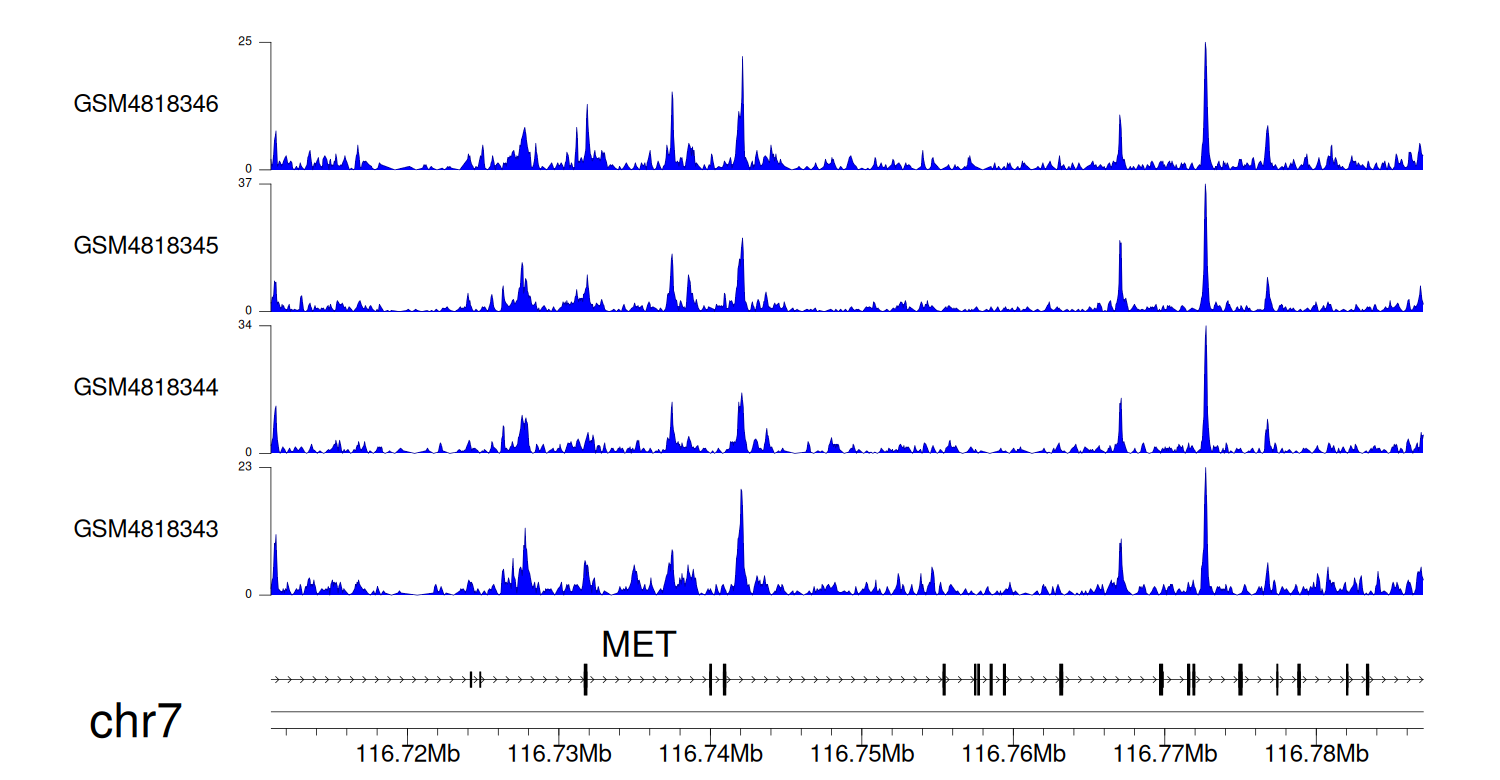
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information