Gene Information
|
Gene Name
|
PTPRO |
|
Gene ID
|
5800
|
|
Gene Full Name
|
protein tyrosine phosphatase receptor type O |
|
Gene Alias
|
GLEPP1|NPHS6|PTP-OC|PTP-U2|PTPROT|PTPU2|R-PTP-O |
|
Transcripts
|
ENSG00000151490
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Intracellular and membrane; 
|
|
Membrane Info
|
Cancer-related genes, Disease related genes, Enzymes, Human disease related genes, Potential drug targets, Predicted intracellular proteins, Predicted membrane proteins |
|
Uniport_ID
|
Q16827
|
|
HGNC ID
|
HGNC:9678
|
|
OMIM ID
|
600579 |
|
Summary
|
This gene encodes a member of the R3 subtype family of receptor-type protein tyrosine phosphatases. These proteins are localized to the apical surface of polarized cells and may have tissue-specific functions through activation of Src family kinases. This gene contains two distinct promoters, and alternatively spliced transcript variants encoding multiple isoforms have been observed. The encoded proteins may have multiple isoform-specific and tissue-specific functions, including the regulation of osteoclast production and activity, inhibition of cell proliferation and facilitation of apoptosis. This gene is a candidate tumor suppressor, and decreased expression of this gene has been observed in several types of cancer. [provided by RefSeq, May 2011] |
Target gene [PTPRO] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1015814 |
chr12 |
2 |
0 |
0 |
2 |
View |
| 1017079 |
chr12 |
1 |
0 |
0 |
1 |
View |
Target gene [PTPRO] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
S GSE199850
|
scRNA-seq |
1 |
HiSeq X Ten (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
C GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
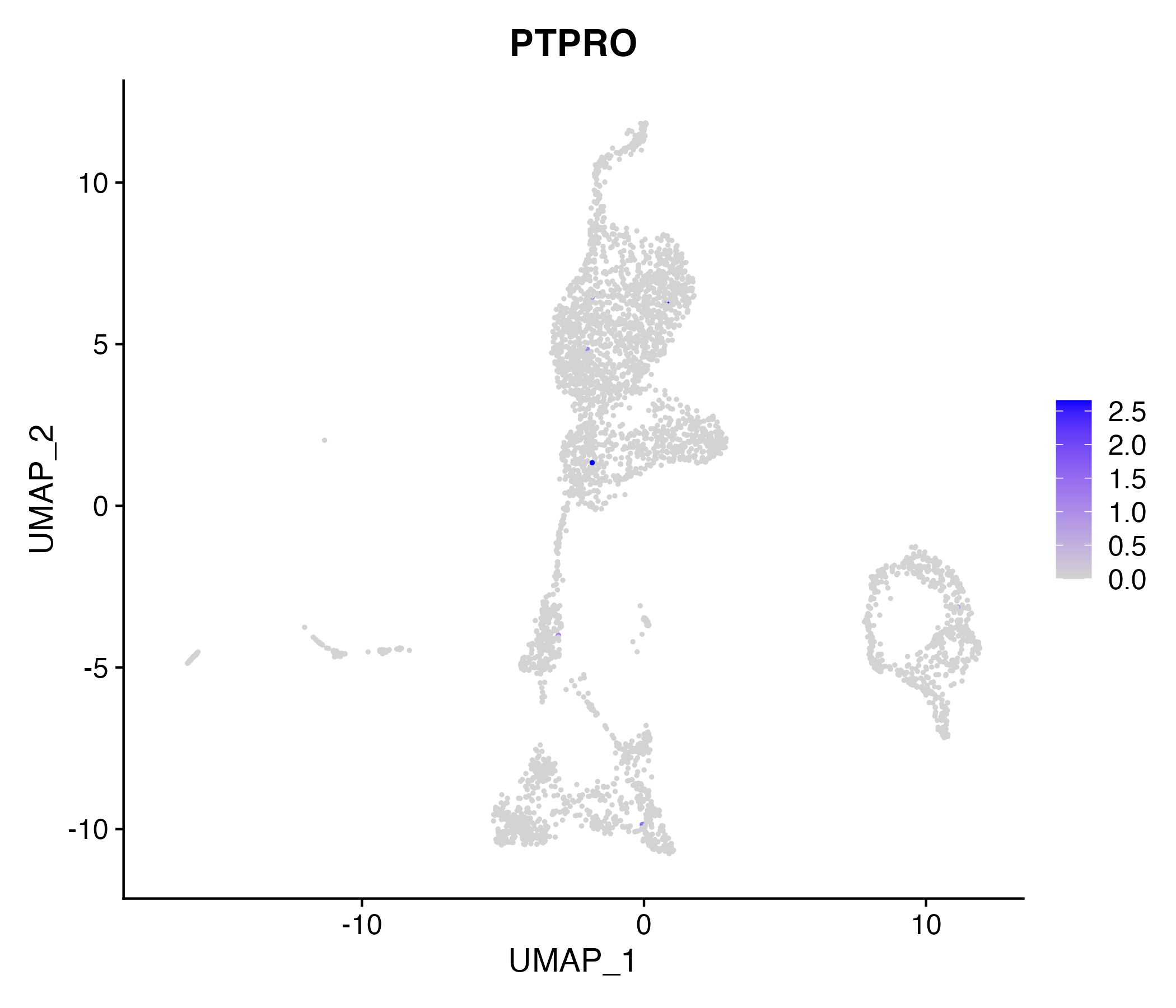
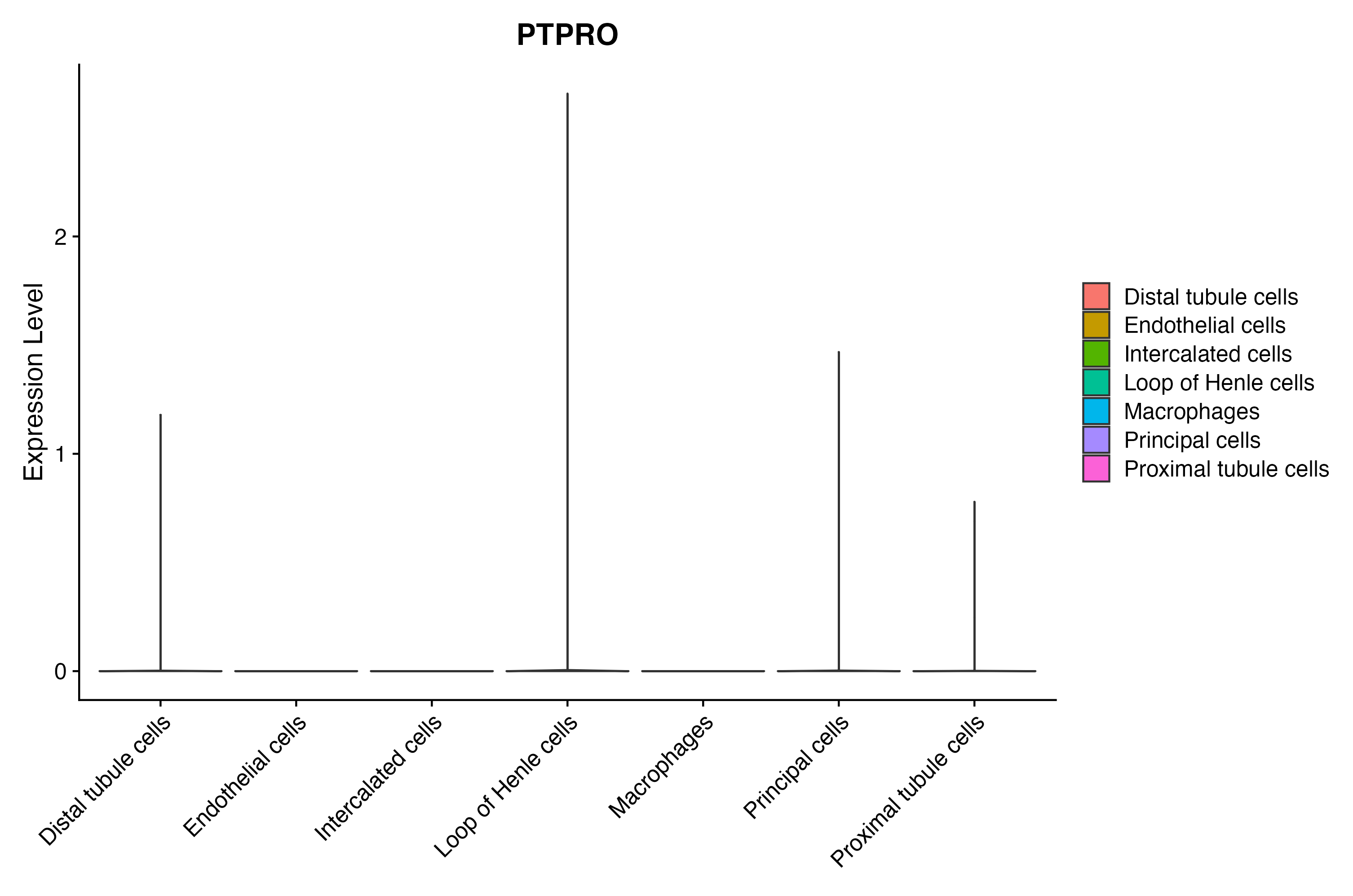
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
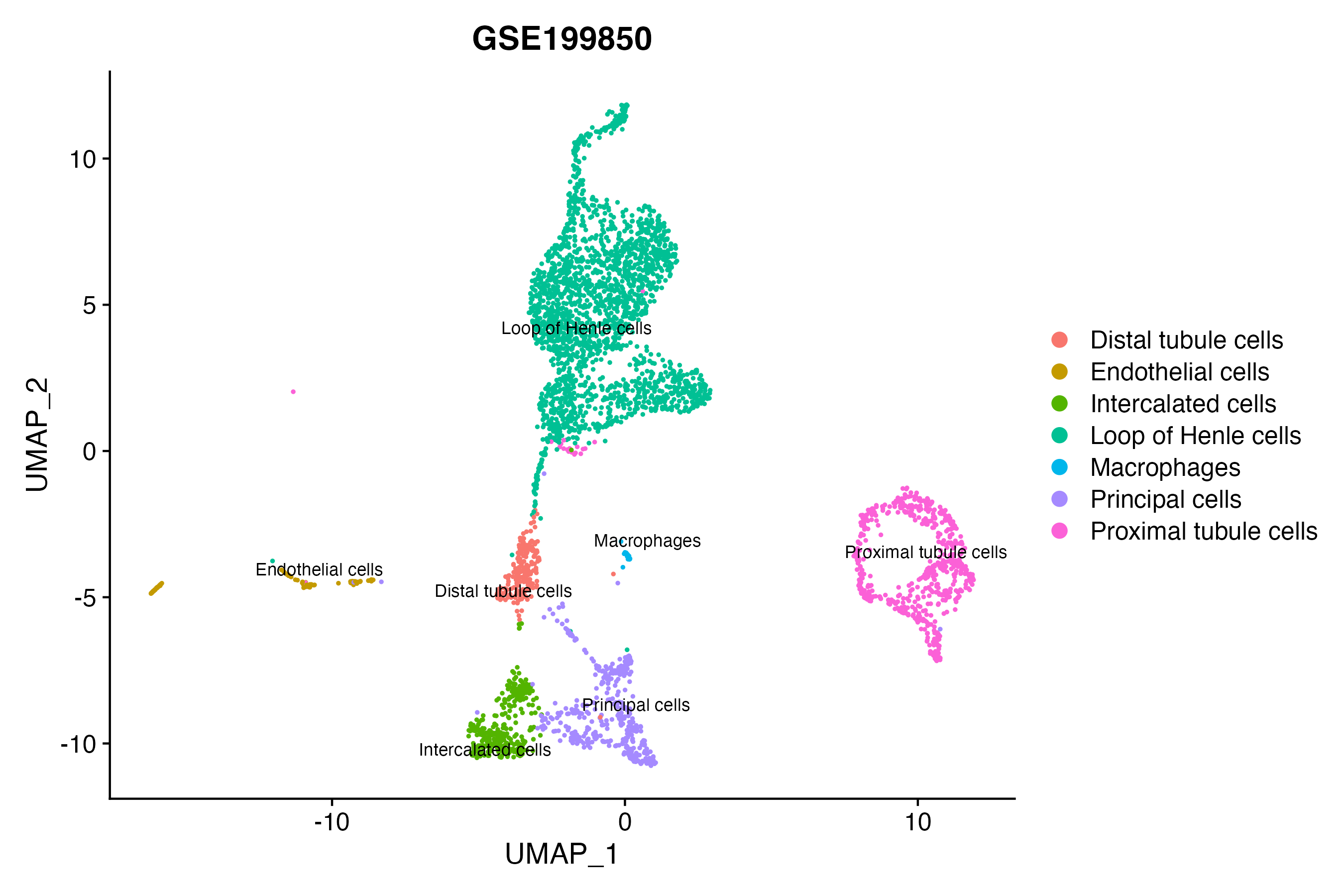
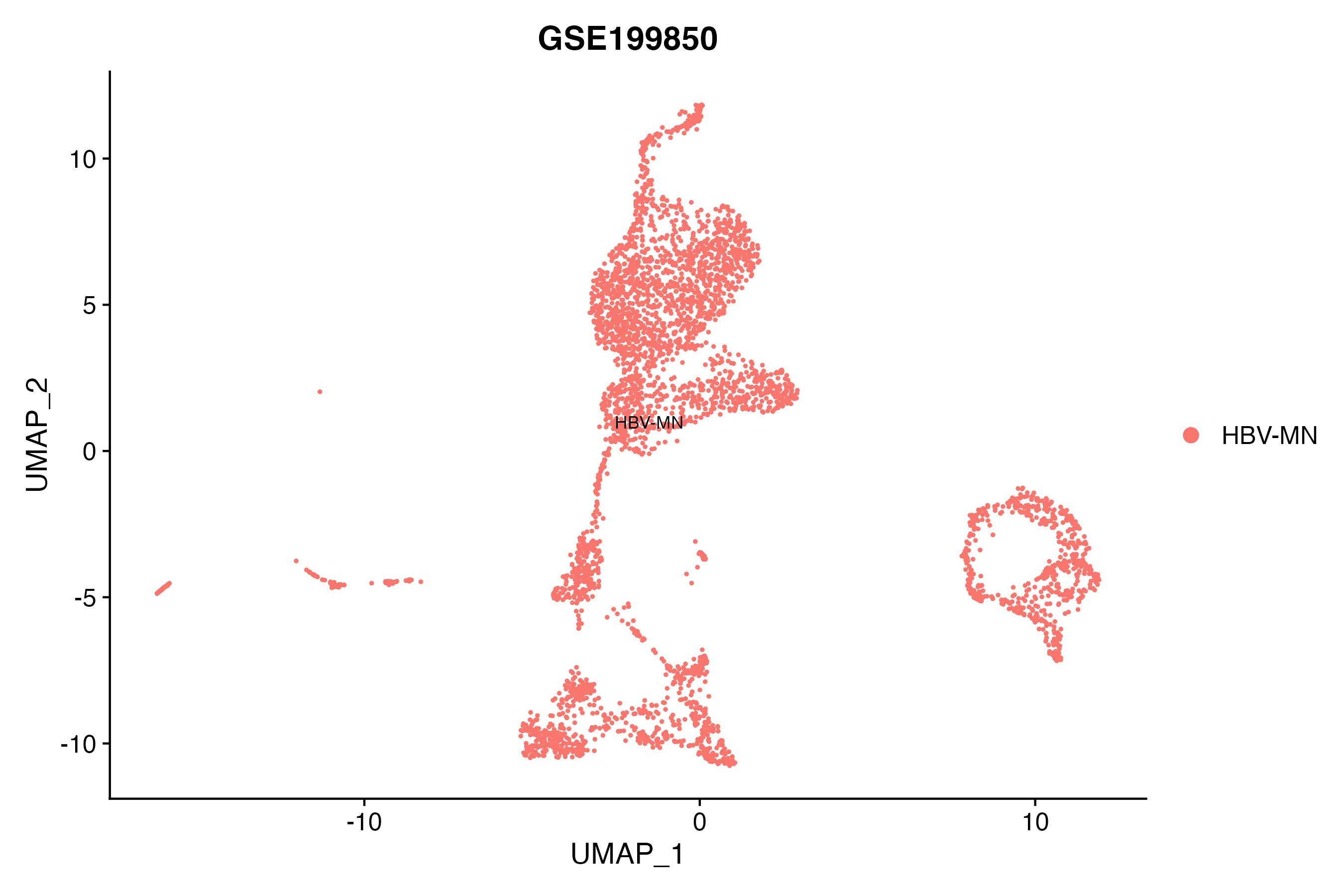
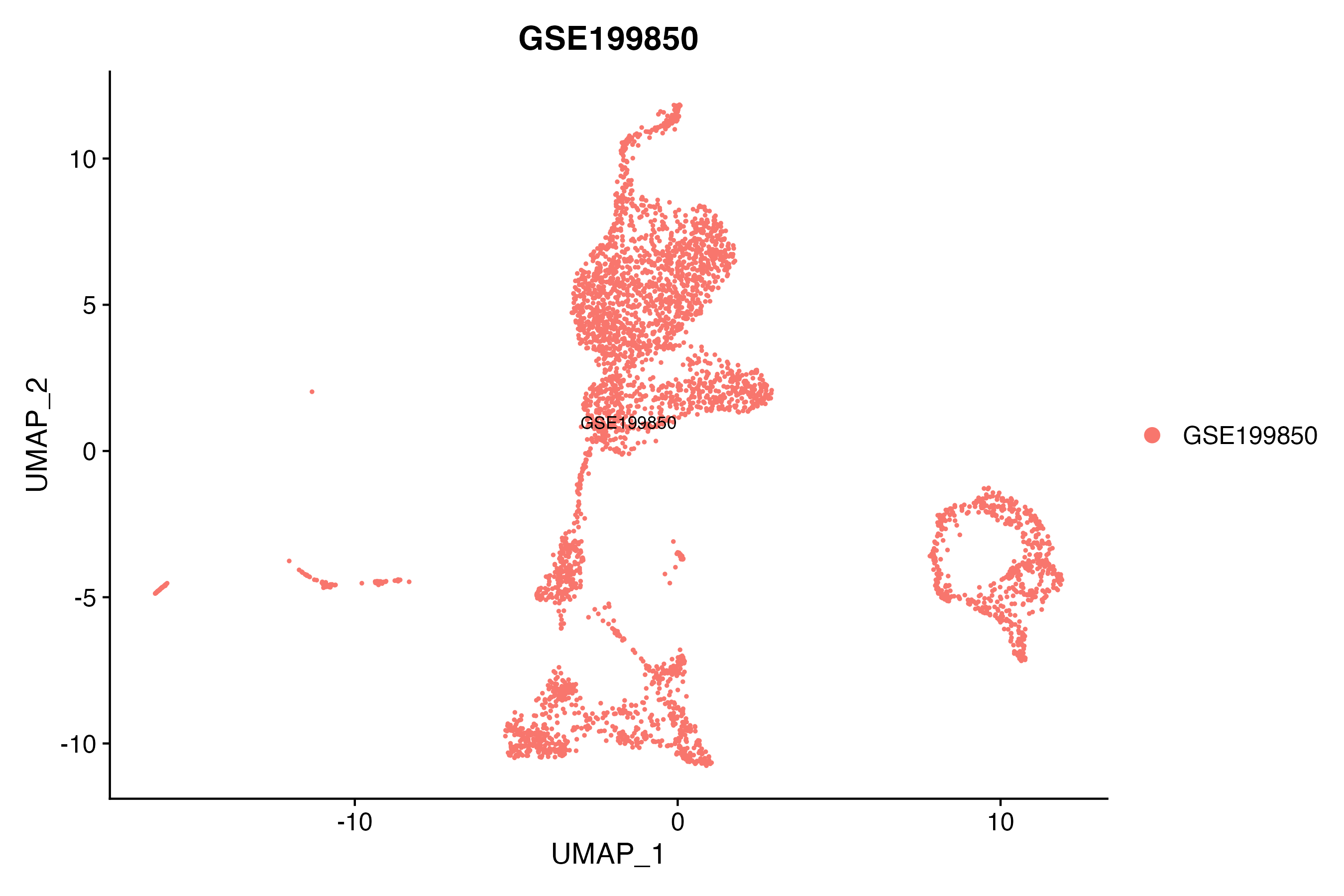
> Dataset: GSE199850 - Gene expression in cell subsets
|
|
|
|
|
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information