Gene Information
|
Gene Name
|
TUSC3 |
|
Gene ID
|
7991
|
|
Gene Full Name
|
tumor suppressor candidate 3 |
|
Gene Alias
|
D8S1992|M33|MRT22|MRT7|MagT2|N33|OST3A|SLC58A2 |
|
Transcripts
|
ENSG00000104723
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|

|
|
Membrane Info
|
Disease related genes, Human disease related genes, Metabolic proteins, Plasma proteins, Potential drug targets, Predicted membrane proteins, Transporters |
|
Uniport_ID
|
Q13454
|
|
HGNC ID
|
HGNC:30242
|
|
OMIM ID
|
601385 |
|
Summary
|
This gene encodes a protein that has been associated with several biological functions including cellular magnesium uptake, protein glycosylation and embryonic development. This protein localizes to the endoplasmic reticulum and acts as a component of the oligosaccharyl transferase complex which is responsible for N-linked protein glycosylation. This gene is a candidate tumor suppressor gene. Homozygous mutations in this gene are associated with autosomal recessive nonsyndromic mental retardation-7 and in the proliferation and invasiveness of several cancers including metastatic pancreatic cancer, ovarian cancer and glioblastoma multiform. [provided by RefSeq, Oct 2017] |
Target gene [TUSC3] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1009065 |
chr8 |
6 |
11 |
0 |
17 |
View |
| 1016070 |
chr8 |
0 |
2 |
0 |
2 |
View |
| 1017876 |
chr8 |
0 |
0 |
0 |
0 |
View |
| 1021603 |
chr8 |
0 |
0 |
0 |
0 |
View |
Target gene [TUSC3] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
E C GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
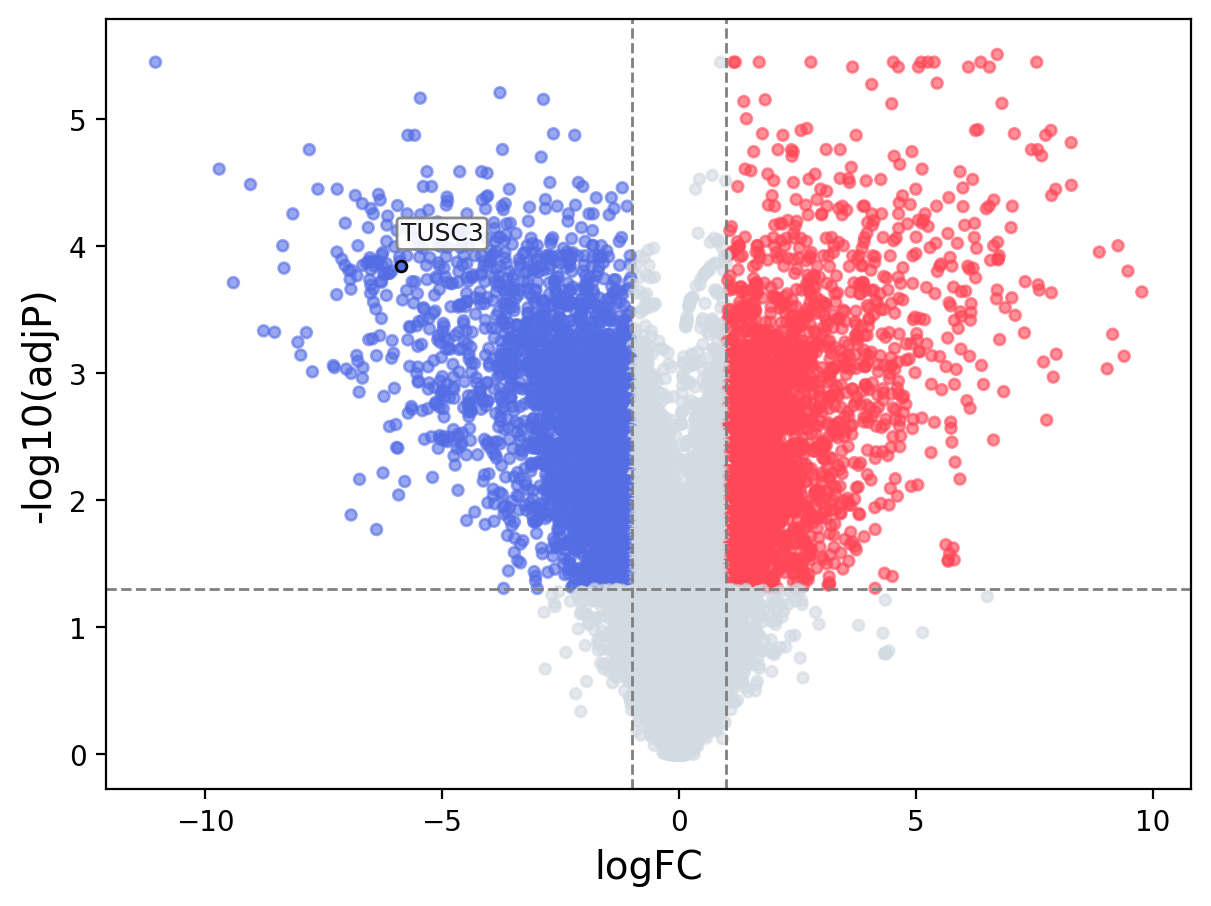
When the query gene is differentially changed in the dataset, a volcano/bar plot will be displayed.
> Dataset: GSE100400 - TUSC3 expression across samples
|
Volcano Plot
|
Bar Plot
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information