Gene Information
|
Gene Name
|
ANK3 |
|
Gene ID
|
288
|
|
Gene Full Name
|
ankyrin 3 |
|
Gene Alias
|
ANKYRIN-G|MRT37 |
|
Transcripts
|
ENSG00000151150
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Plasma membrane; 
|
|
Membrane Info
|
Cancer-related genes, Disease related genes, Human disease related genes, Plasma proteins, Potential drug targets, Predicted intracellular proteins, Transporters |
|
Uniport_ID
|
Q12955
|
|
HGNC ID
|
HGNC:494
|
|
OMIM ID
|
600465 |
|
Summary
|
Ankyrins are a family of proteins that are believed to link the integral membrane proteins to the underlying spectrin-actin cytoskeleton and play key roles in activities such as cell motility, activation, proliferation, contact, and the maintenance of specialized membrane domains. Multiple isoforms of ankyrin with different affinities for various target proteins are expressed in a tissue-specific, developmentally regulated manner. Most ankyrins are typically composed of three structural domains: an amino-terminal domain containing multiple ankyrin repeats; a central region with a highly conserved spectrin binding domain; and a carboxy-terminal regulatory domain which is the least conserved and subject to variation. Ankyrin 3 is an immunologically distinct gene product from ankyrins 1 and 2, and was originally found at the axonal initial segment and nodes of Ranvier of neurons in the central and peripheral nervous systems. Multiple transcript variants encoding different isoforms have been found for this gene.[provided by RefSeq, Feb 2011] |
Target gene [ANK3] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1011481 |
chr10 |
6 |
3 |
0 |
9 |
View |
| 1011749 |
chr10 |
6 |
2 |
0 |
8 |
View |
| 1011909 |
chr10 |
5 |
3 |
0 |
8 |
View |
| 1012488 |
chr10 |
5 |
1 |
0 |
6 |
View |
| 1015564 |
chr10 |
2 |
0 |
0 |
2 |
View |
| 1016003 |
chr10 |
1 |
1 |
0 |
2 |
View |
| 1041818 |
chr10 |
88 |
34 |
65 |
187 |
View |
Target gene [ANK3] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
GSE199850
|
scRNA-seq |
1 |
HiSeq X Ten (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
S GSE247322
|
scRNA-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
E C GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
When the query gene is differentially changed in the dataset, a volcano/bar plot will be displayed.
> Dataset: GSE100400 - ANK3 expression across samples
|
Volcano Plot
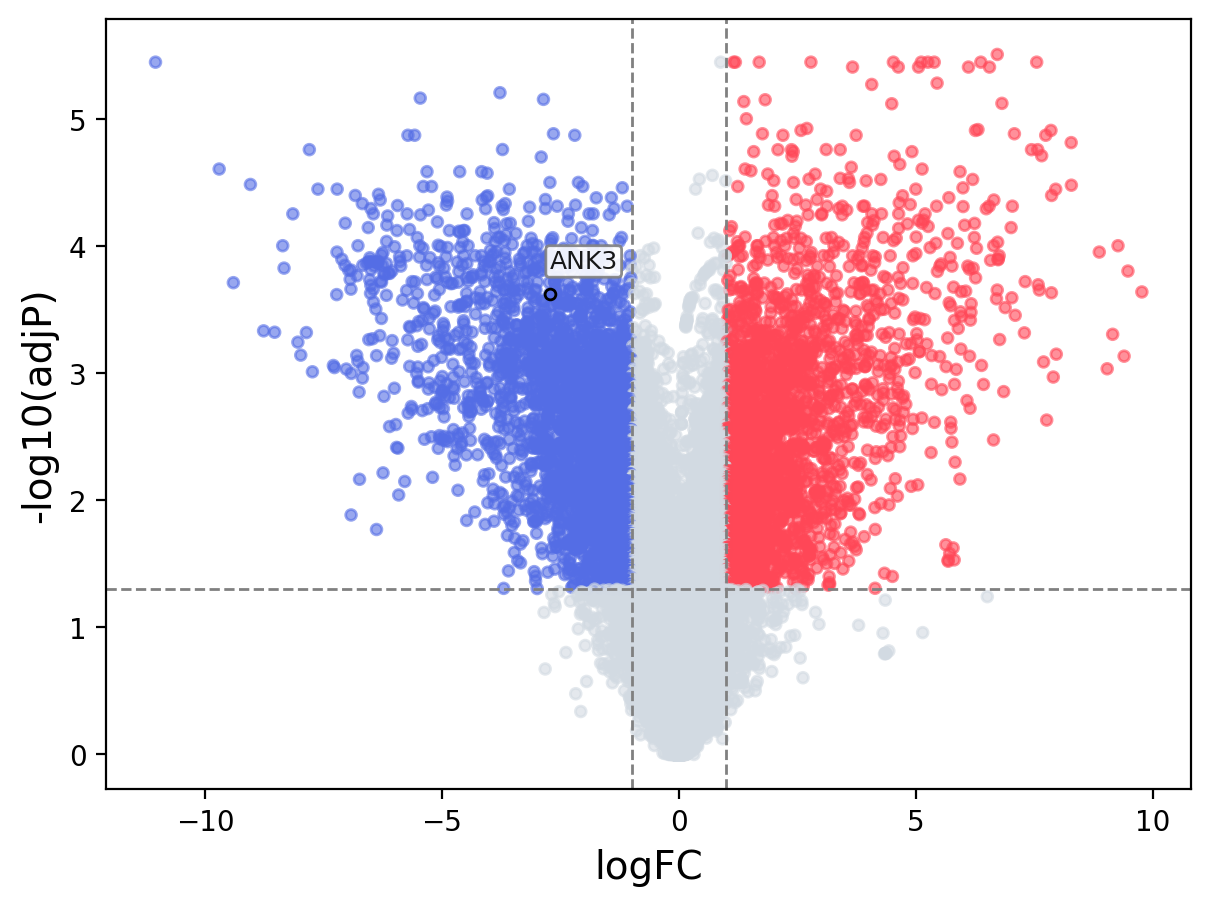
|
Bar Plot
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information