Gene Information
|
Gene Name
|
AXL |
|
Gene ID
|
558
|
|
Gene Full Name
|
AXL receptor tyrosine kinase |
|
Gene Alias
|
ARK|AXL3|JTK11|Tyro7|UFO |
|
Transcripts
|
ENSG00000167601
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Plasma membrane;Vesicles, Actin filaments; 
|
|
Membrane Info
|
Disease related genes, Enzymes, FDA approved drug targets, Plasma proteins, Predicted membrane proteins |
|
Uniport_ID
|
P30530
|
|
HGNC ID
|
HGNC:905
|
|
OMIM ID
|
109135 |
|
Summary
|
The protein encoded by this gene is a member of the Tyro3-Axl-Mer (TAM) receptor tyrosine kinase subfamily. The encoded protein possesses an extracellular domain which is composed of two immunoglobulin-like motifs at the N-terminal, followed by two fibronectin type-III motifs. It transduces signals from the extracellular matrix into the cytoplasm by binding to the vitamin K-dependent protein growth arrest-specific 6 (Gas6). This gene may be involved in several cellular functions including growth, migration, aggregation and anti-inflammation in multiple cell types. The encoded protein acts as a host cell receptor for multiple viruses, including Marburg, Ebola and Lassa viruses and is a candidate receptor for the SARS-CoV2 virus. [provided by RefSeq, Sep 2021] |
Target gene [AXL] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1041255 |
chr19 |
19 |
3 |
0 |
22 |
View |
| 1042434 |
chr19 |
53 |
17 |
3 |
73 |
View |
Target gene [AXL] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
S GSE252863
|
scRNA-seq |
10 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
S GSE199850
|
scRNA-seq |
1 |
HiSeq X Ten (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE252863 - Gene expression in cell subsets
|
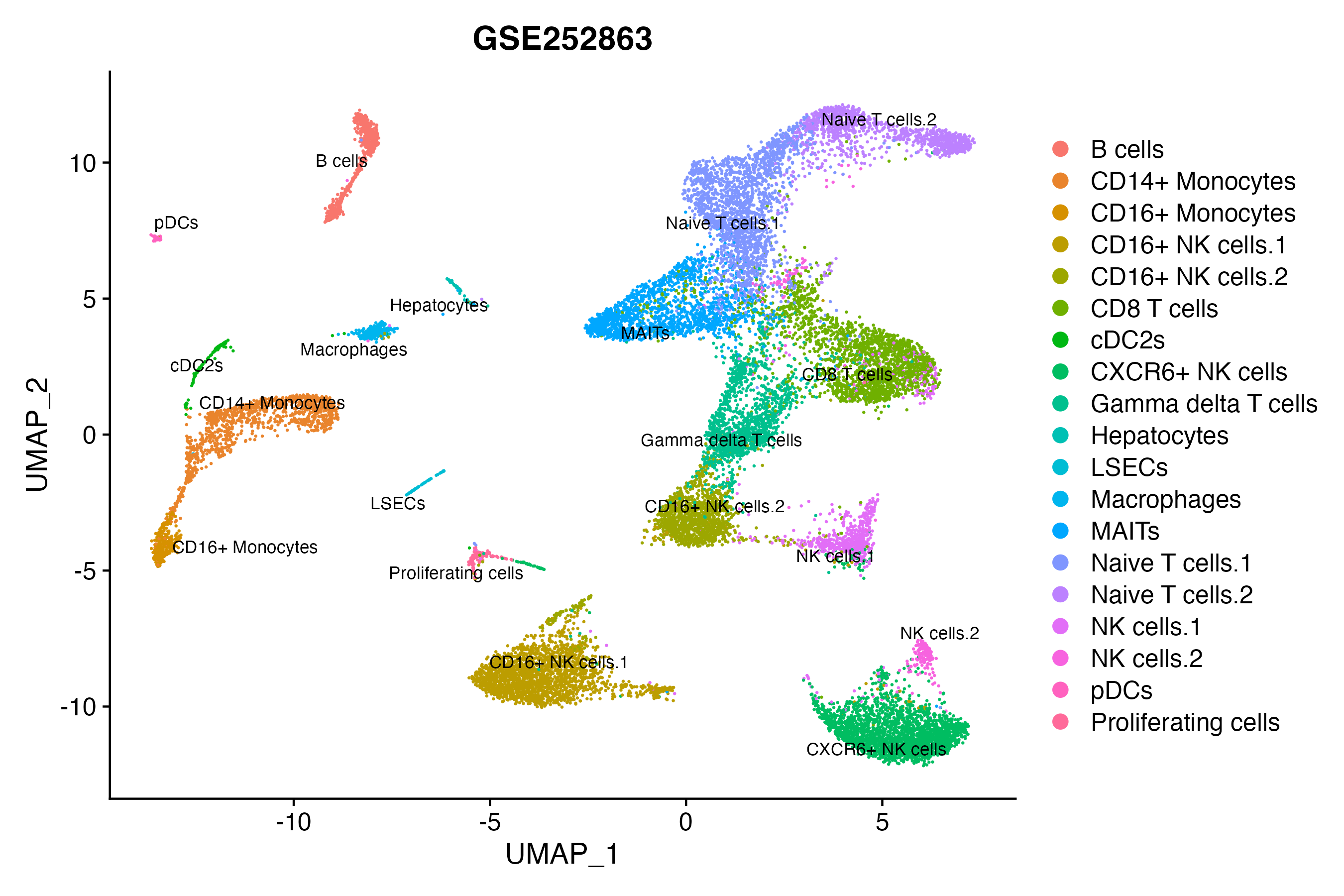
|
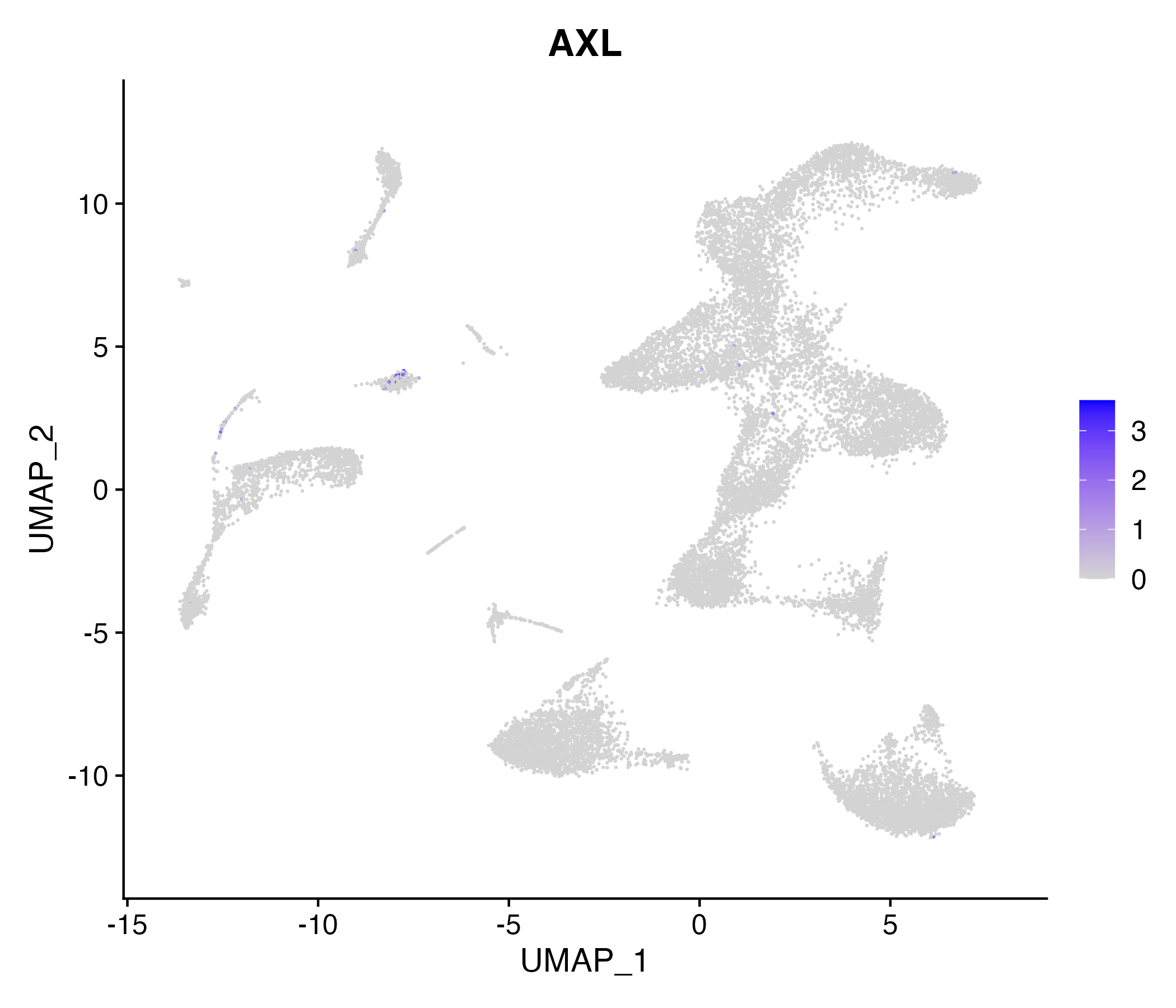
|
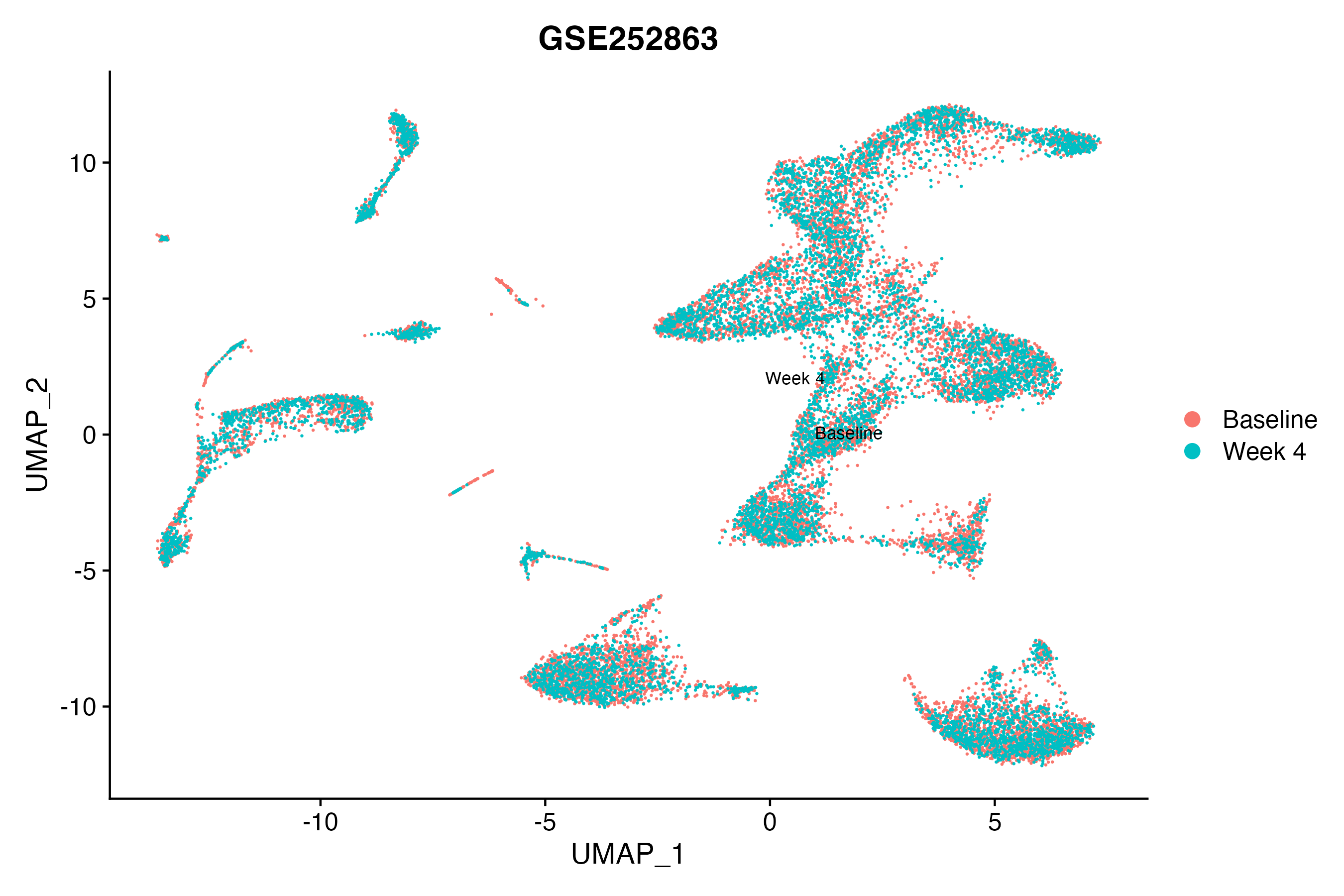
|
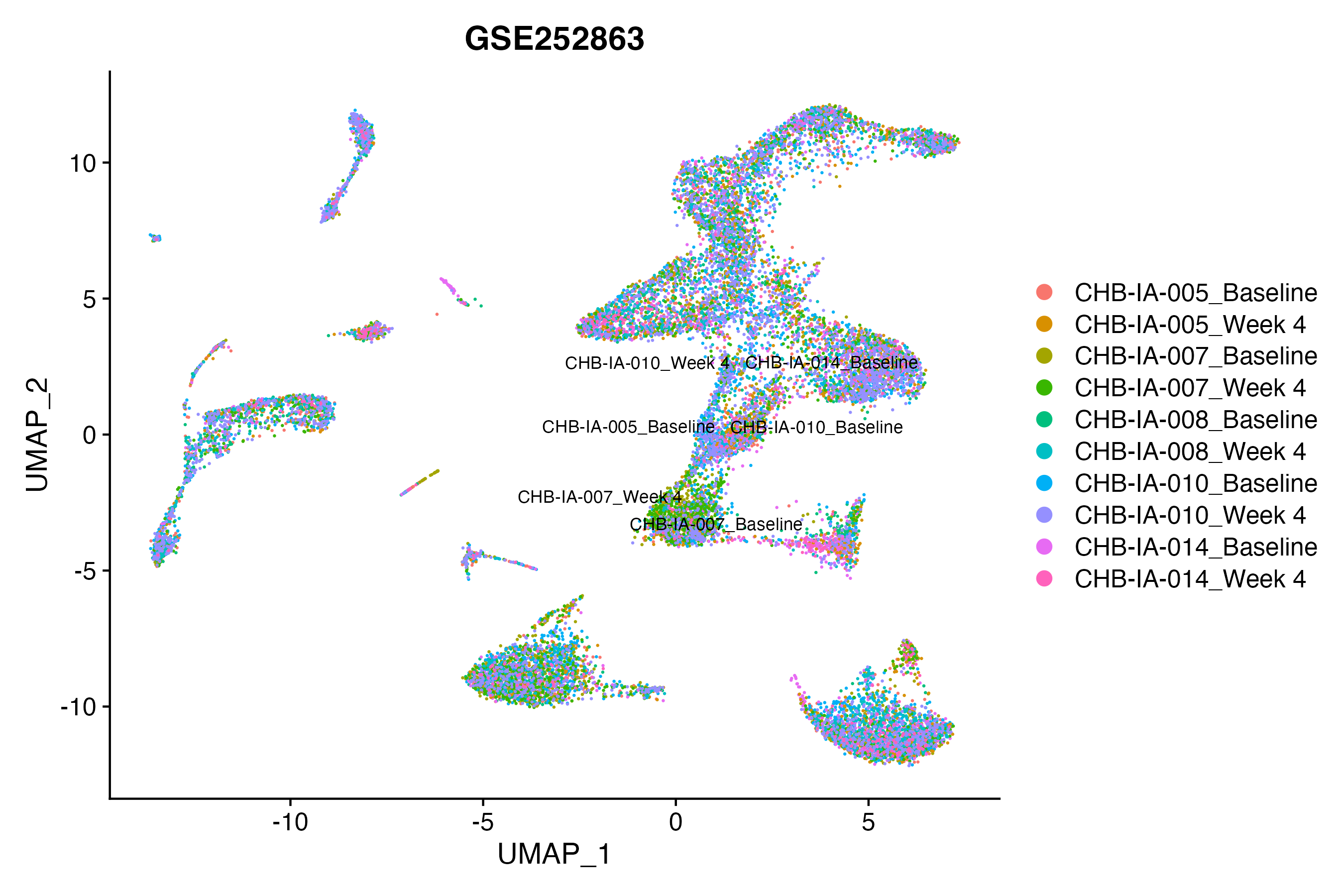
|
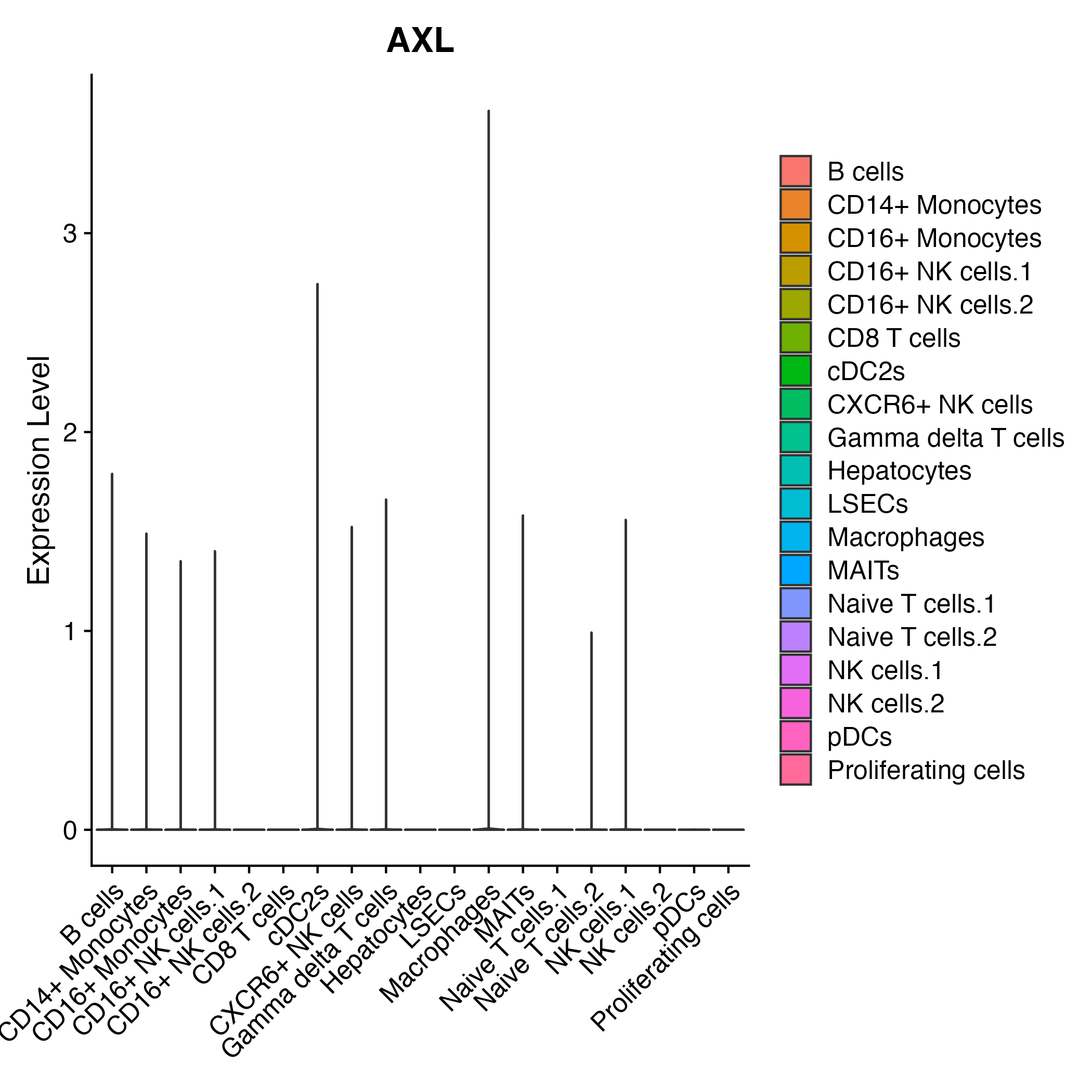
|
> Dataset: GSE199850 - Gene expression in cell subsets
|
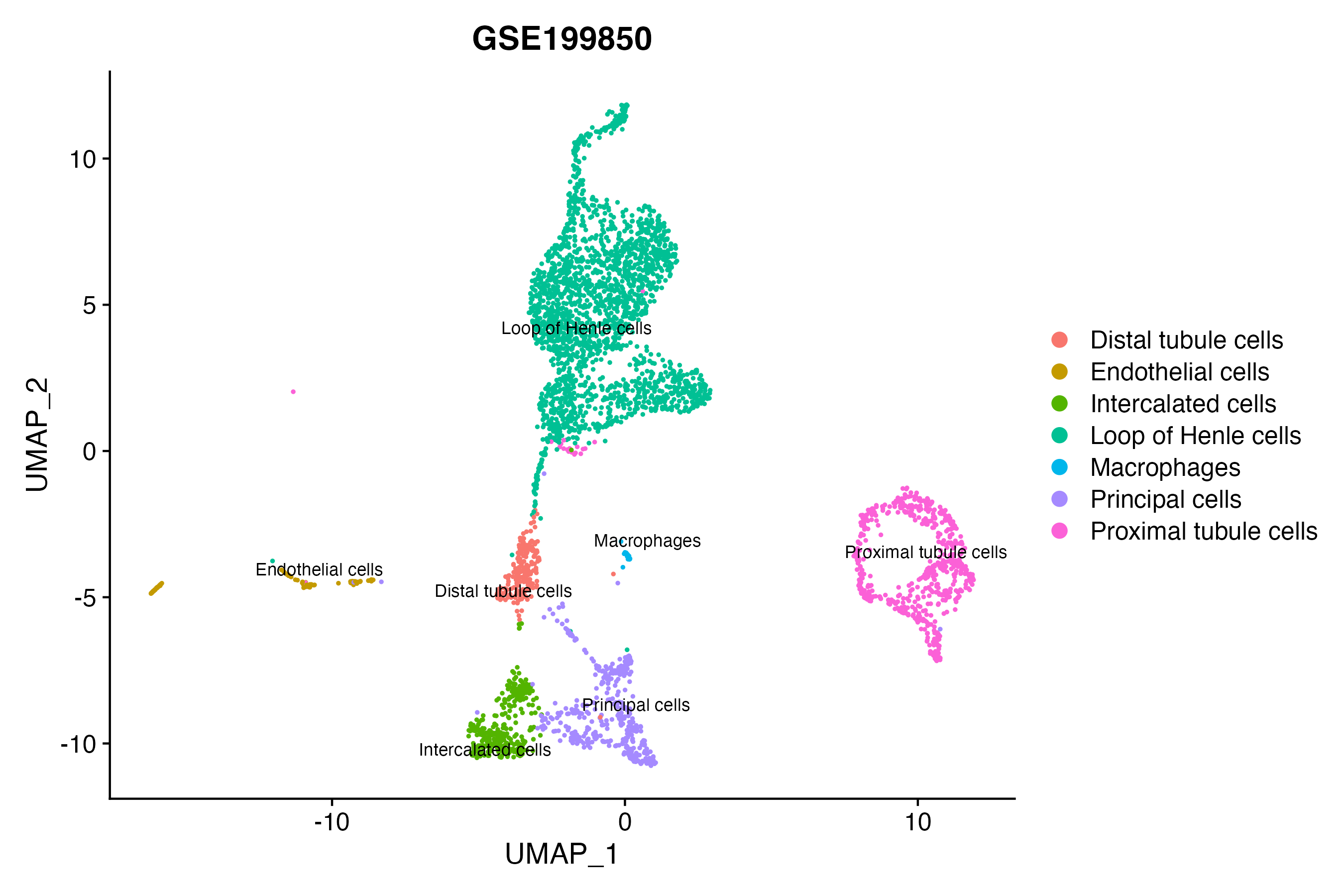
|
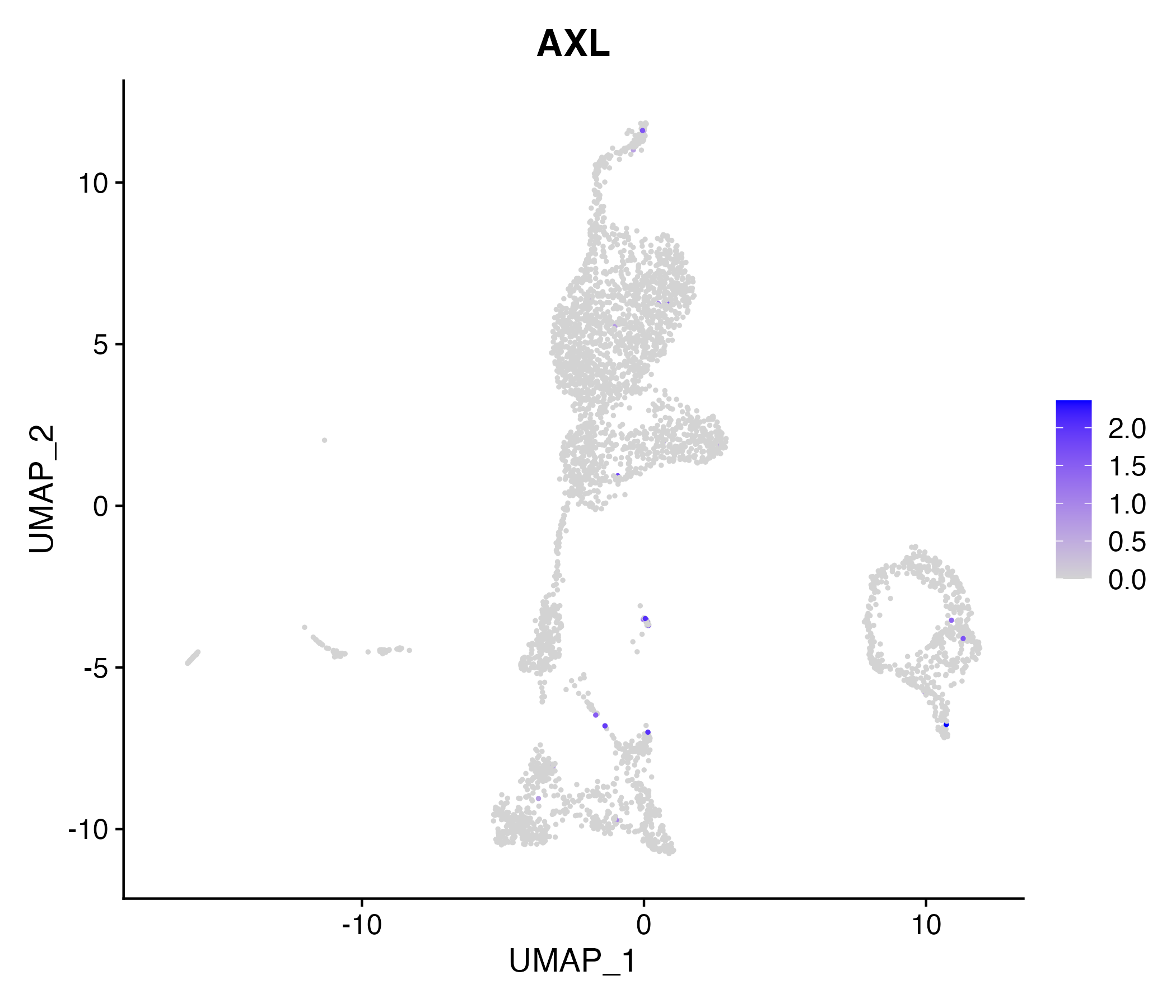
|
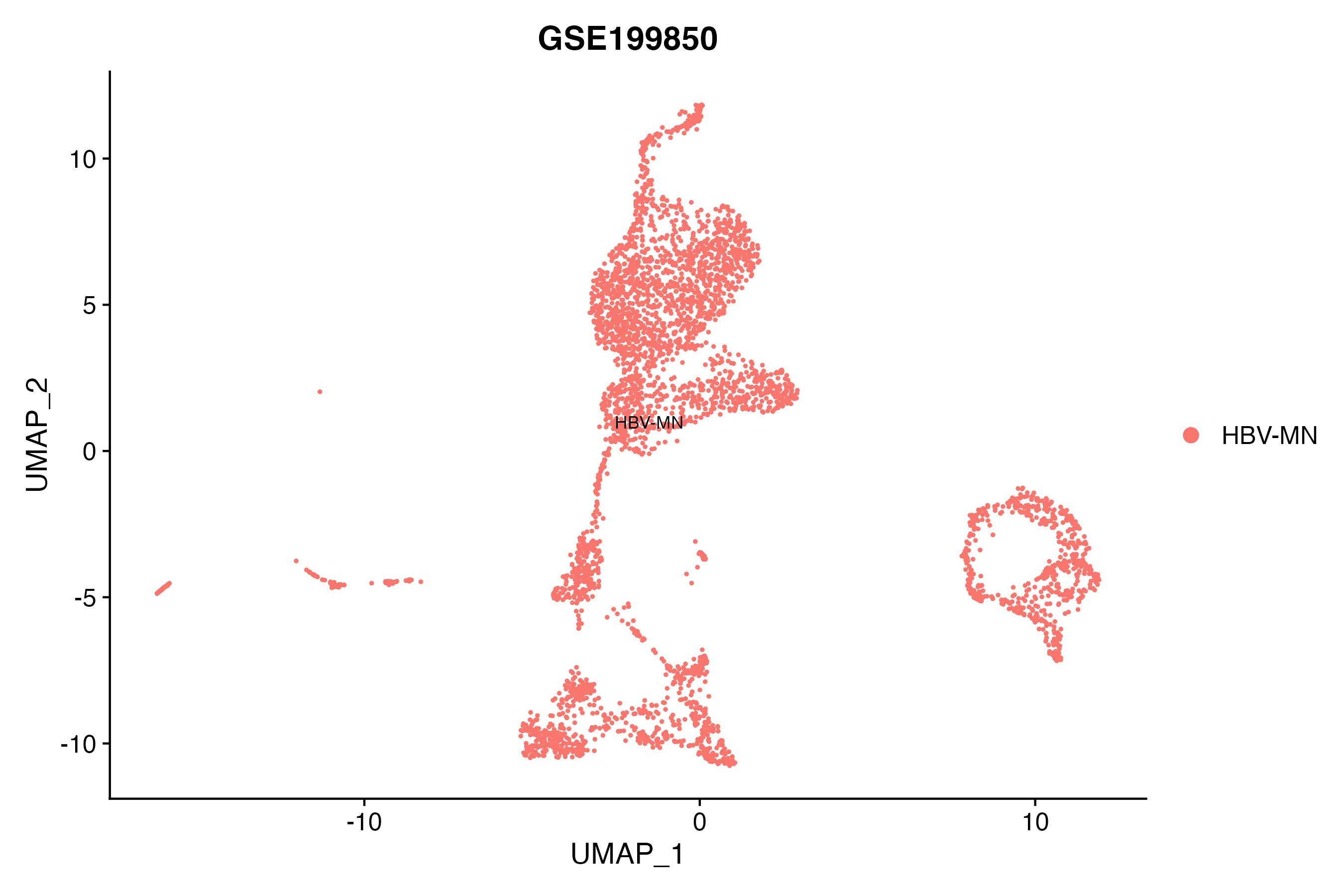
|
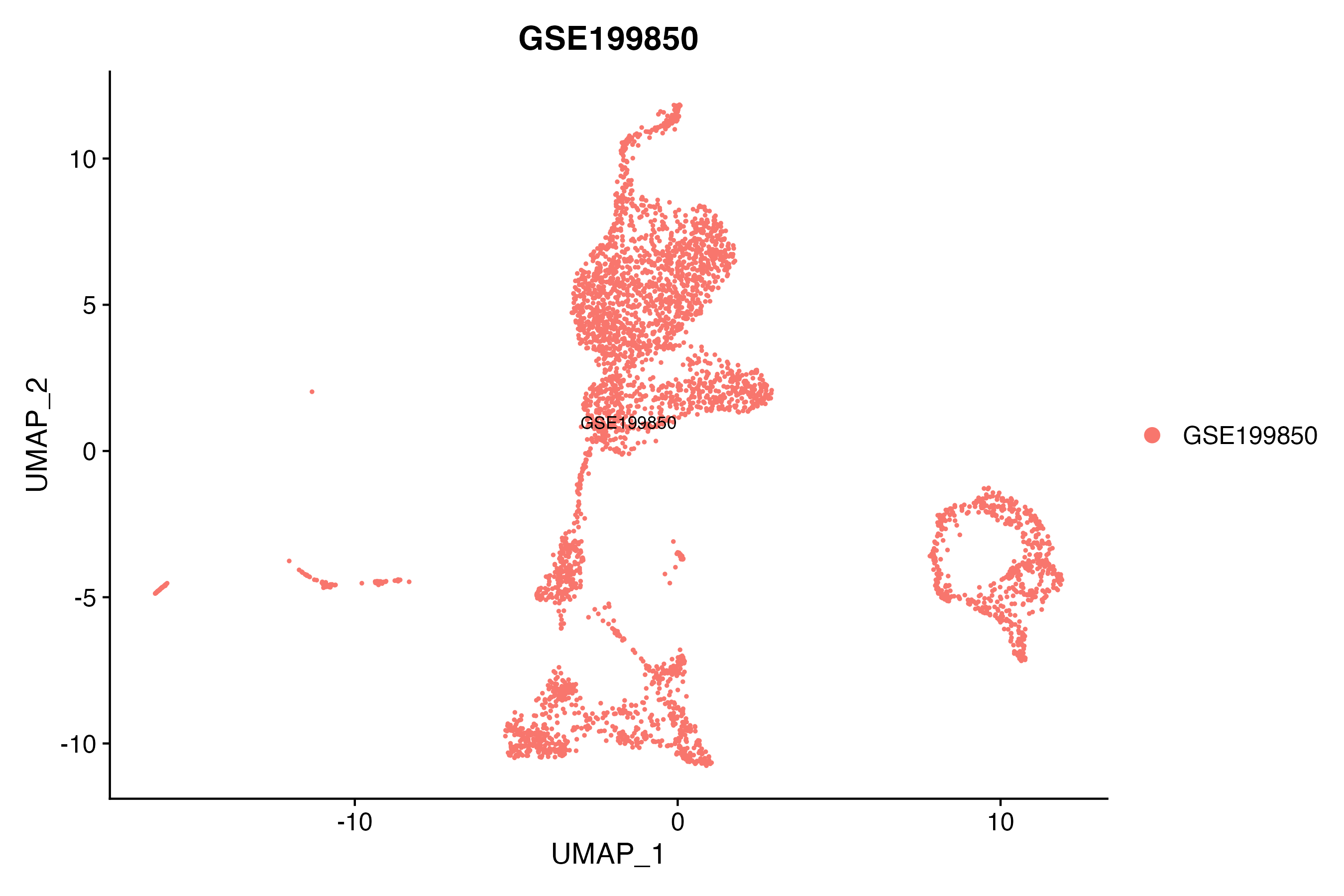
|
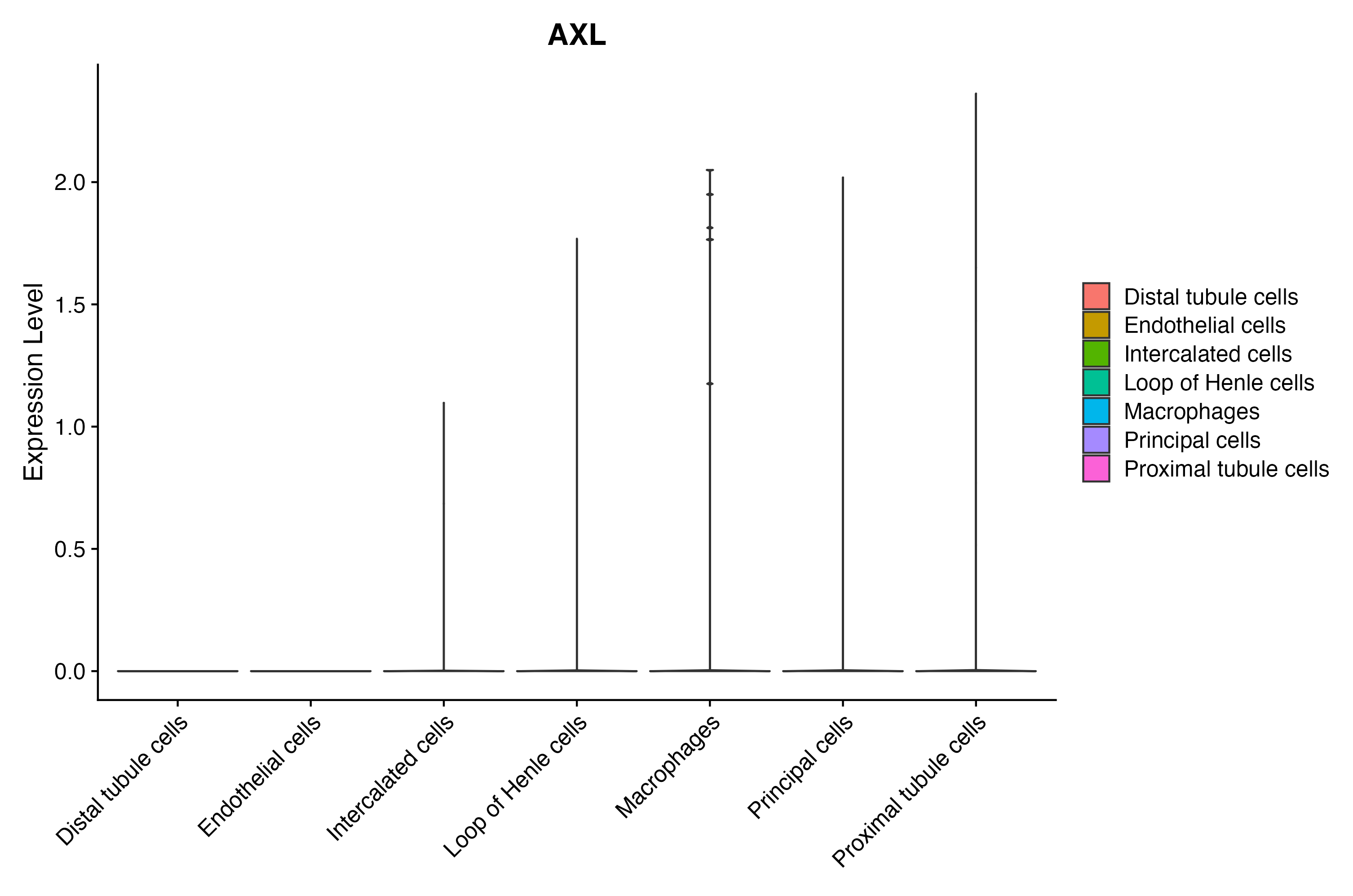
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information