Gene Information
|
Gene Name
|
MASP1 |
|
Gene ID
|
5648
|
|
Gene Full Name
|
MBL associated serine protease 1 |
|
Gene Alias
|
3MC1|CRARF|CRARF1|MAP-1|MAP1|MASP|MASP-3|MASP3|MAp44|PRSS5|RaRF |
|
Transcripts
|
ENSG00000127241
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Nucleoplasm, Cytosol;Secreted to blood; 
|
|
Membrane Info
|
Disease related genes, Enzymes, Human disease related genes, Plasma proteins, Potential drug targets, Predicted intracellular proteins, Predicted secreted proteins |
|
Uniport_ID
|
P48740
|
|
HGNC ID
|
HGNC:6901
|
|
OMIM ID
|
600521 |
|
Summary
|
This gene encodes a serine protease that functions as a component of the lectin pathway of complement activation. The complement pathway plays an essential role in the innate and adaptive immune response. The encoded protein is synthesized as a zymogen and is activated when it complexes with the pathogen recognition molecules of lectin pathway, the mannose-binding lectin and the ficolins. This protein is not directly involved in complement activation but may play a role as an amplifier of complement activation by cleaving complement C2 or by activating another complement serine protease, MASP-2. The encoded protein is also able to cleave fibrinogen and factor XIII and may may be involved in coagulation. A splice variant of this gene which lacks the serine protease domain functions as an inhibitor of the complement pathway. Alternate splicing results in multiple transcript variants.[provided by RefSeq, Apr 2010] |
Target gene [MASP1] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1010248 |
chr3 |
12 |
1 |
0 |
13 |
View |
| 1010249 |
chr3 |
13 |
0 |
0 |
13 |
View |
Target gene [MASP1] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE252863
|
scRNA-seq |
10 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
E GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
When the query gene is differentially changed in the dataset, a volcano/bar plot will be displayed.
> Dataset: GSE94660 - MASP1 expression across samples
|
Volcano Plot
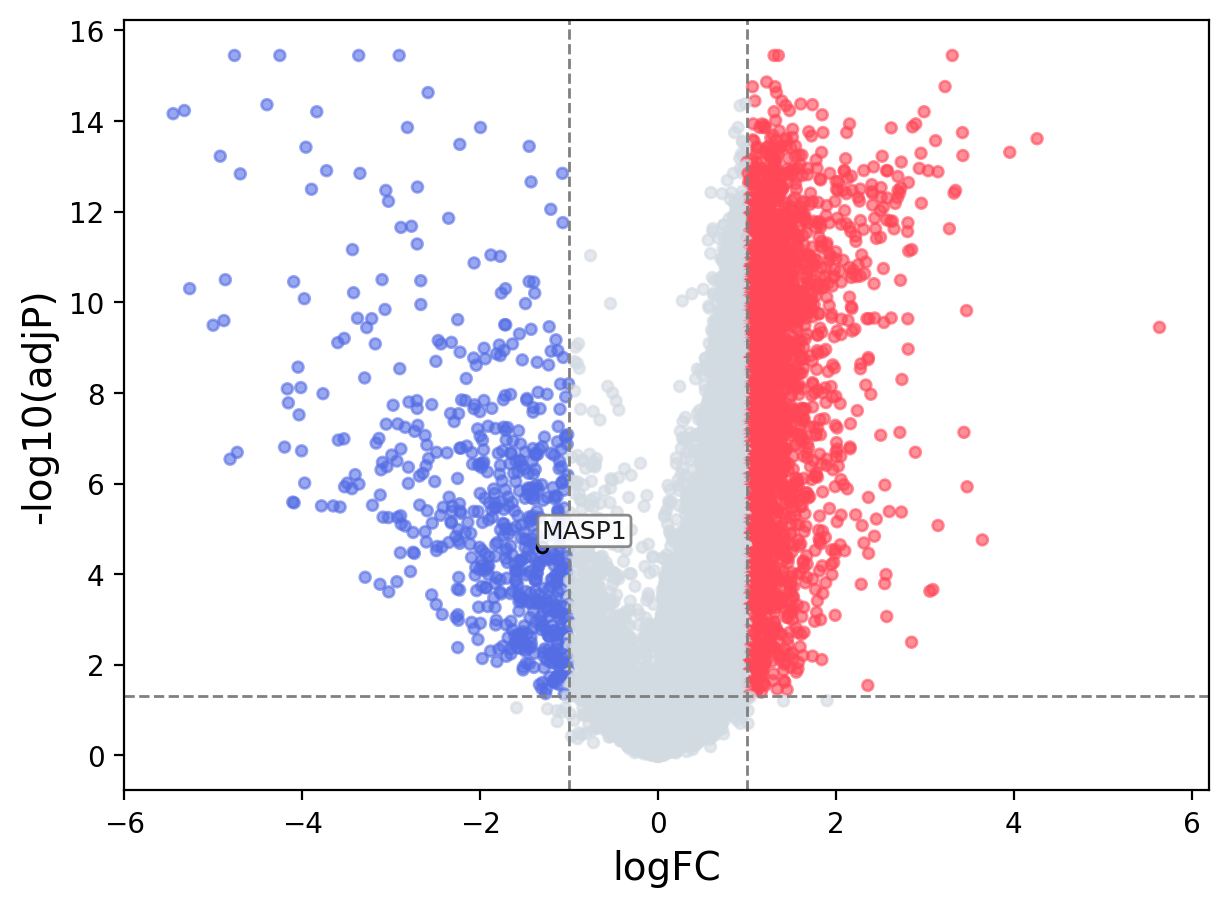
|
Bar Plot
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information