Gene Information
|
Gene Name
|
RASA1 |
|
Gene ID
|
5921
|
|
Gene Full Name
|
RAS p21 protein activator 1 |
|
Gene Alias
|
CM-AVM|CMAVM|CMAVM1|GAP|PKWS|RASA|RASGAP|p120|p120GAP|p120RASGAP |
|
Transcripts
|
ENSG00000145715
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Vesicles, Basal body;Cytosol; 
|
|
Membrane Info
|
Cancer-related genes, Disease related genes, Human disease related genes, Plasma proteins, Predicted intracellular proteins, RAS pathway related proteins |
|
Uniport_ID
|
P20936
|
|
HGNC ID
|
HGNC:9871
|
|
OMIM ID
|
139150 |
|
Summary
|
The protein encoded by this gene is located in the cytoplasm and is part of the GAP1 family of GTPase-activating proteins. The gene product stimulates the GTPase activity of normal RAS p21 but not its oncogenic counterpart. Acting as a suppressor of RAS function, the protein enhances the weak intrinsic GTPase activity of RAS proteins resulting in the inactive GDP-bound form of RAS, thereby allowing control of cellular proliferation and differentiation. Mutations leading to changes in the binding sites of either protein are associated with basal cell carcinomas. Mutations also have been associated with hereditary capillary malformations (CM) with or without arteriovenous malformations (AVM) and Parkes Weber syndrome. Alternative splicing results in two isoforms where the shorter isoform, lacking the N-terminal hydrophobic region but retaining the same activity, appears to be abundantly expressed in placental but not adult tissues. [provided by RefSeq, May 2012] |
Target gene [RASA1] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1020123 |
chr5 |
42 |
116 |
3 |
161 |
View |
Target gene [RASA1] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
GSE199850
|
scRNA-seq |
1 |
HiSeq X Ten (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
S GSE247322
|
scRNA-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
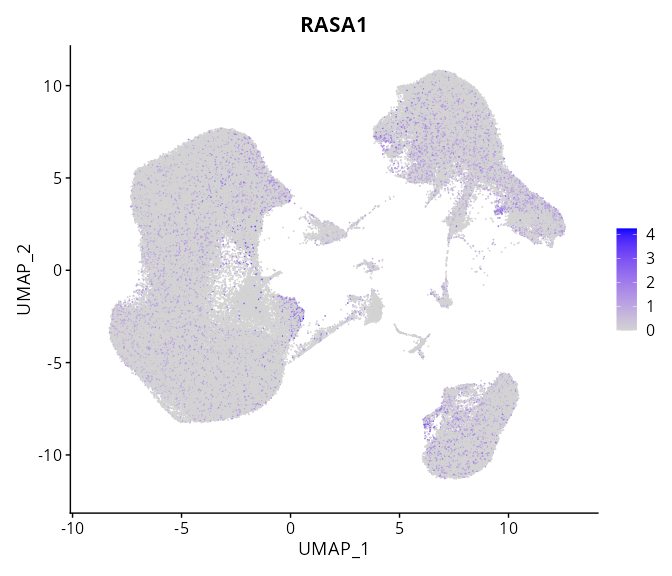
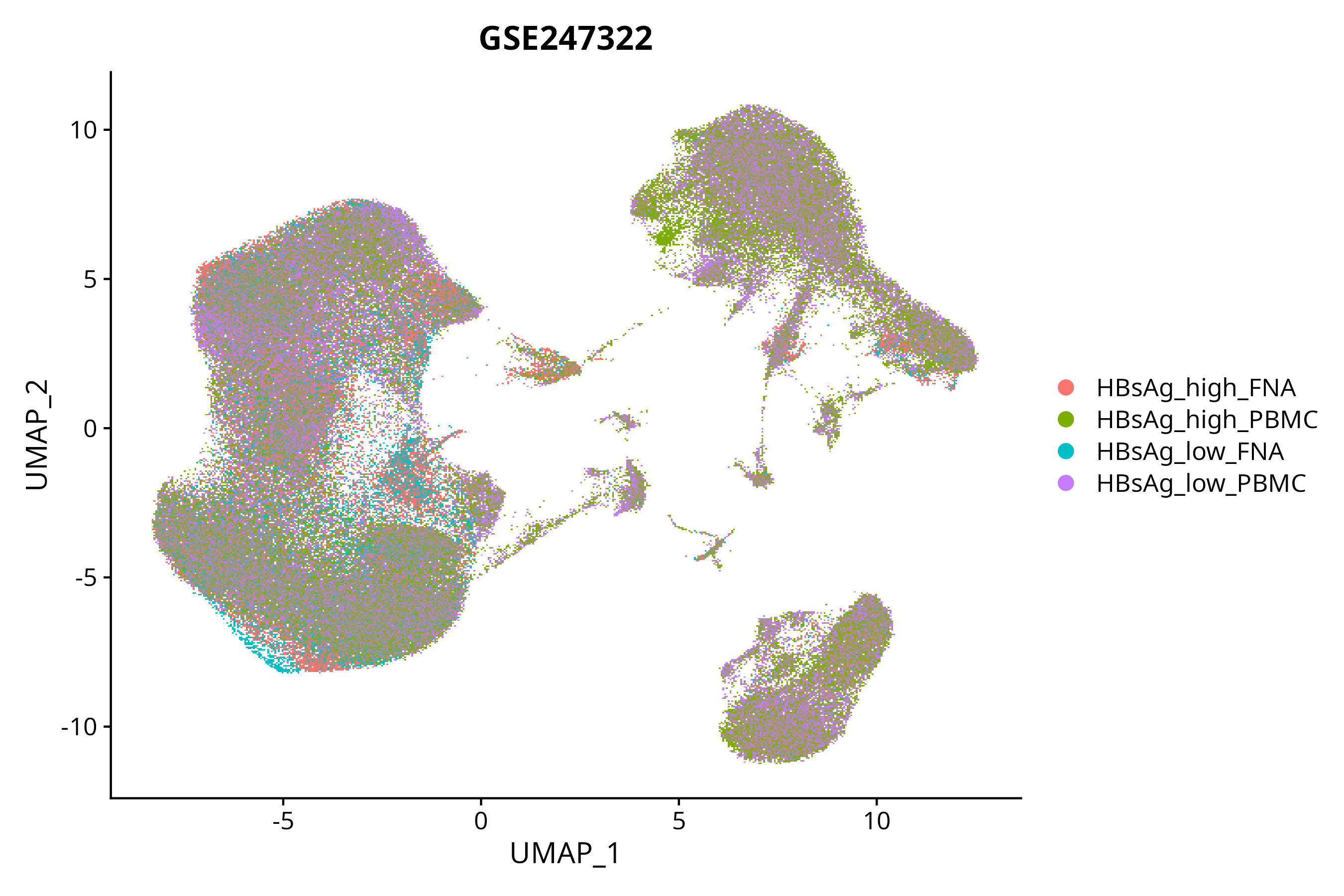
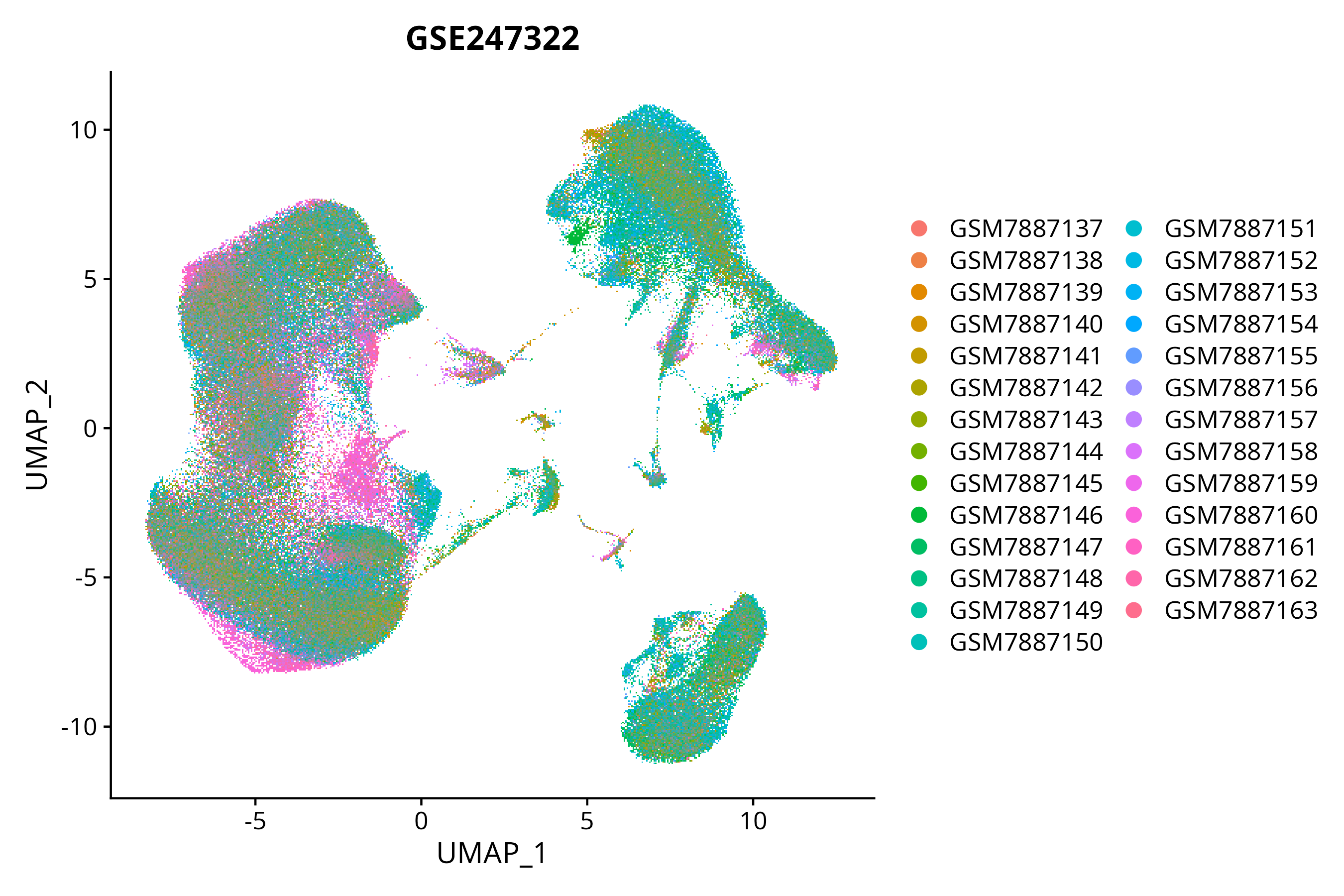
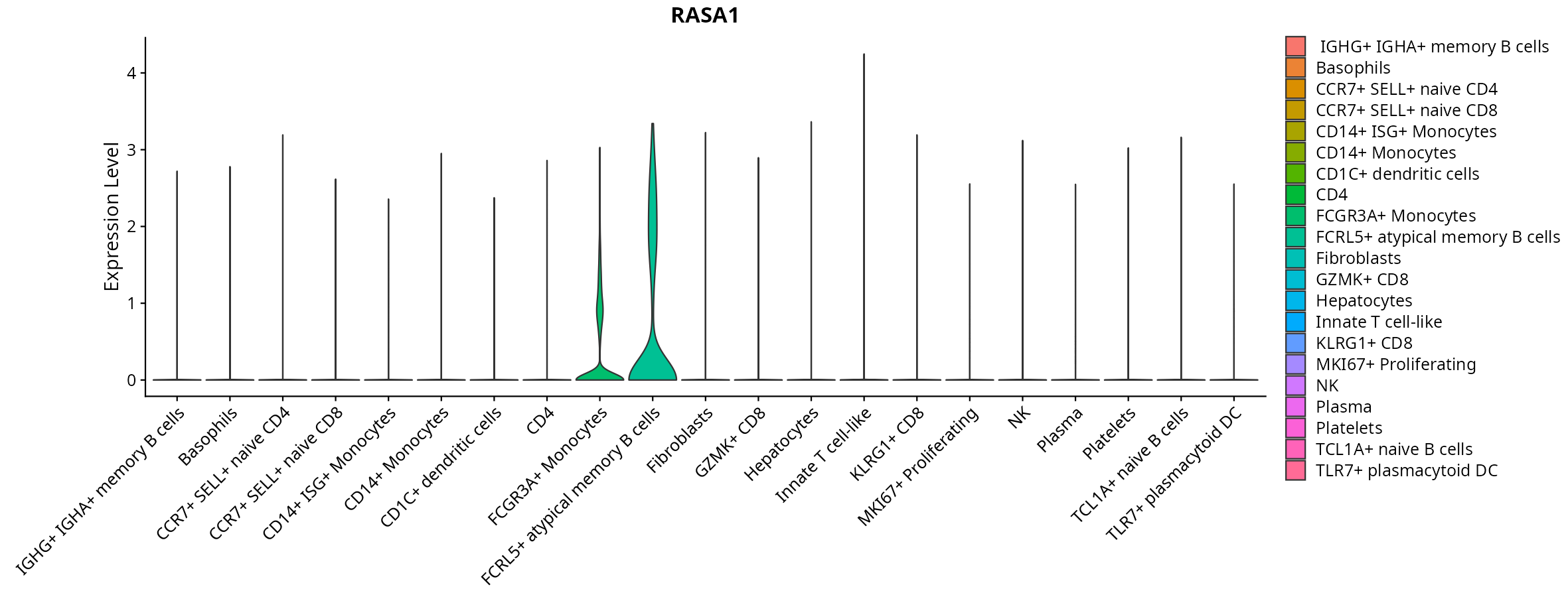
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE247322 - Gene expression in cell subsets
|
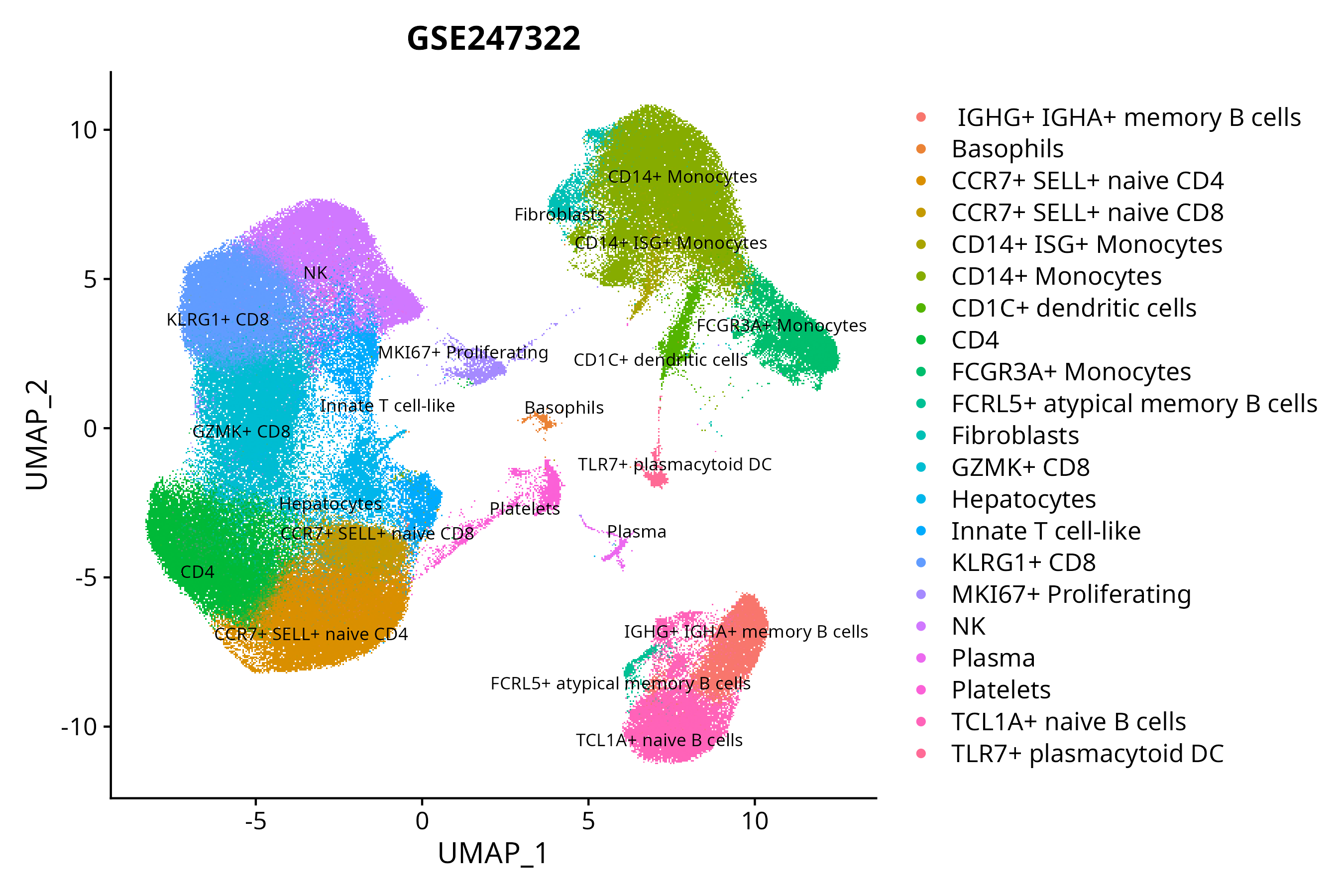

|
|
|
|
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information