Gene Information
|
Gene Name
|
SLC12A3 |
|
Gene ID
|
6559
|
|
Gene Full Name
|
solute carrier family 12 member 3 |
|
Gene Alias
|
NCC|NCCT|TSC |
|
Transcripts
|
ENSG00000070915
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|

|
|
Membrane Info
|
Disease related genes, FDA approved drug targets, Human disease related genes, Metabolic proteins, Plasma proteins, Predicted membrane proteins, Transporters |
|
Uniport_ID
|
P55017
|
|
HGNC ID
|
HGNC:10912
|
|
OMIM ID
|
600968 |
|
Summary
|
This gene encodes a renal thiazide-sensitive sodium-chloride cotransporter that is important for electrolyte homeostasis. This cotransporter mediates sodium and chloride reabsorption in the distal convoluted tubule. Mutations in this gene cause Gitelman syndrome, a disease similar to Bartter's syndrome, that is characterized by hypokalemic alkalosis combined with hypomagnesemia, low urinary calcium, and increased renin activity associated with normal blood pressure. This cotransporter is the target for thiazide diuretics that are used for treating high blood pressure. Multiple transcript variants encoding different isoforms have been found for this gene. [provided by RefSeq, Jul 2008] |
Target gene [SLC12A3] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1012152 |
chr16 |
6 |
1 |
0 |
7 |
View |
| 1014459 |
chr16 |
2 |
1 |
0 |
3 |
View |
| 1021835 |
chr16 |
2 |
1 |
0 |
3 |
View |
| 1042196 |
chr16 |
2 |
1 |
0 |
3 |
View |
Target gene [SLC12A3] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
S GSE199850
|
scRNA-seq |
1 |
HiSeq X Ten (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
GSE247322
|
scRNA-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
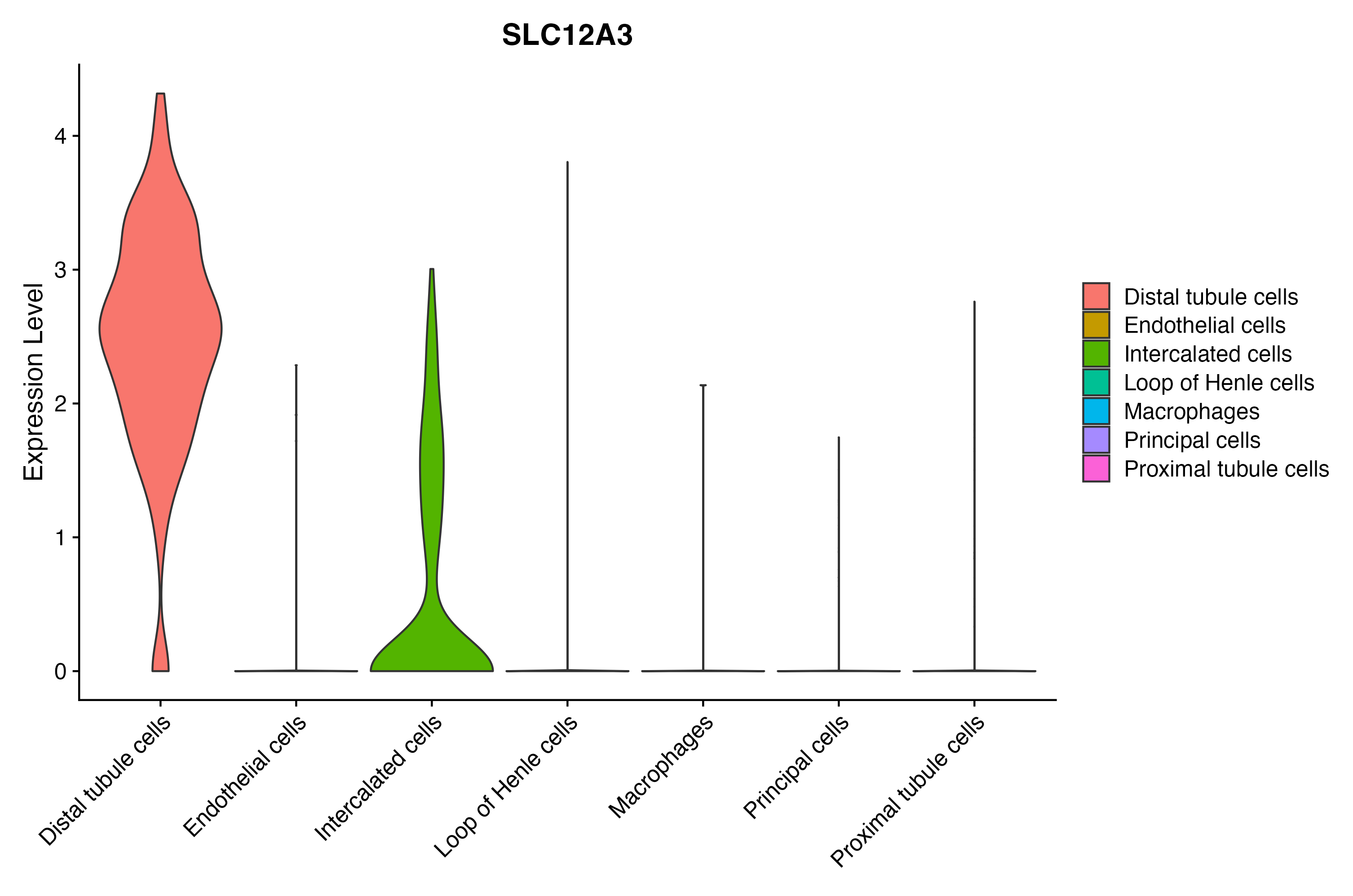
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE199850 - Gene expression in cell subsets
|
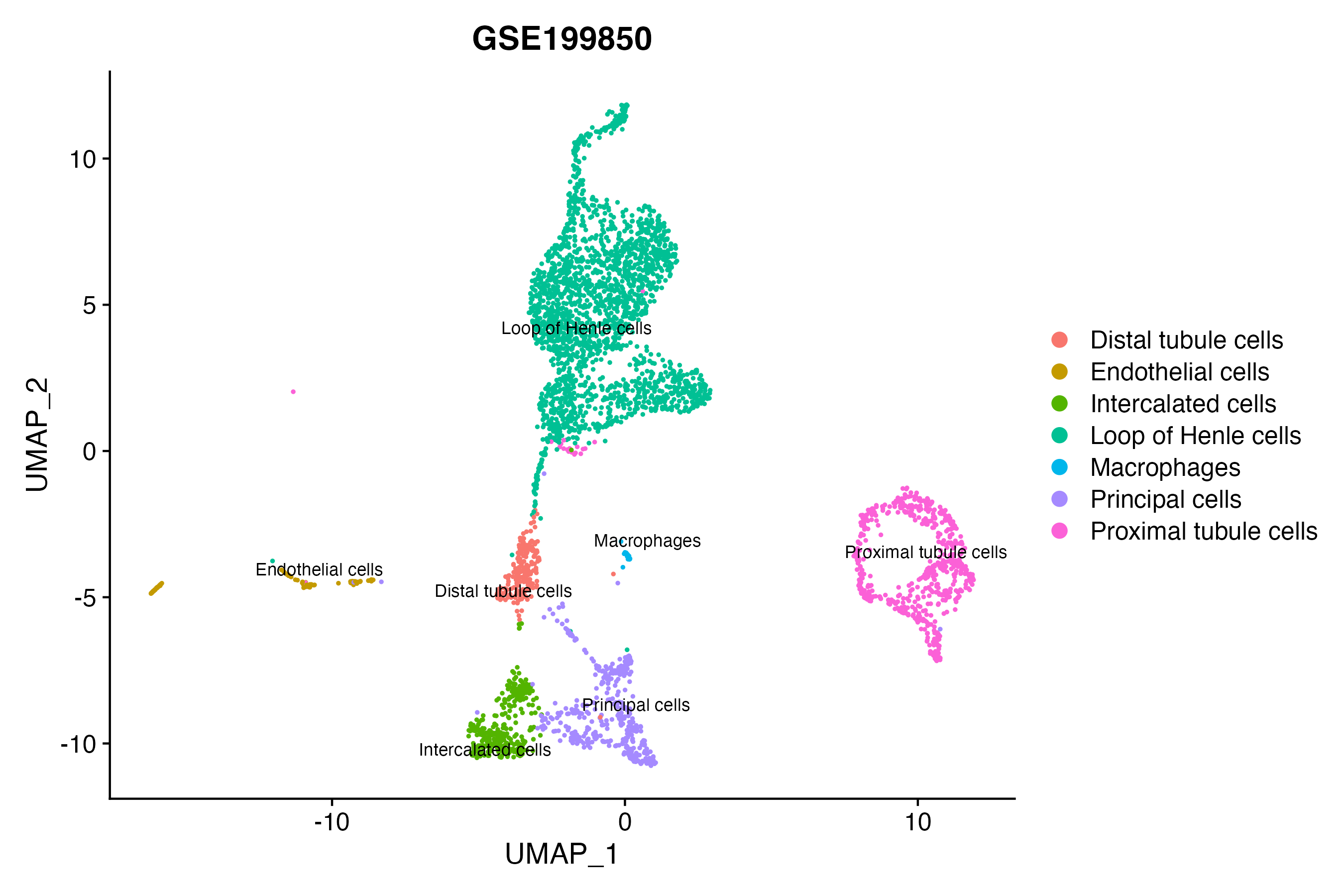
|
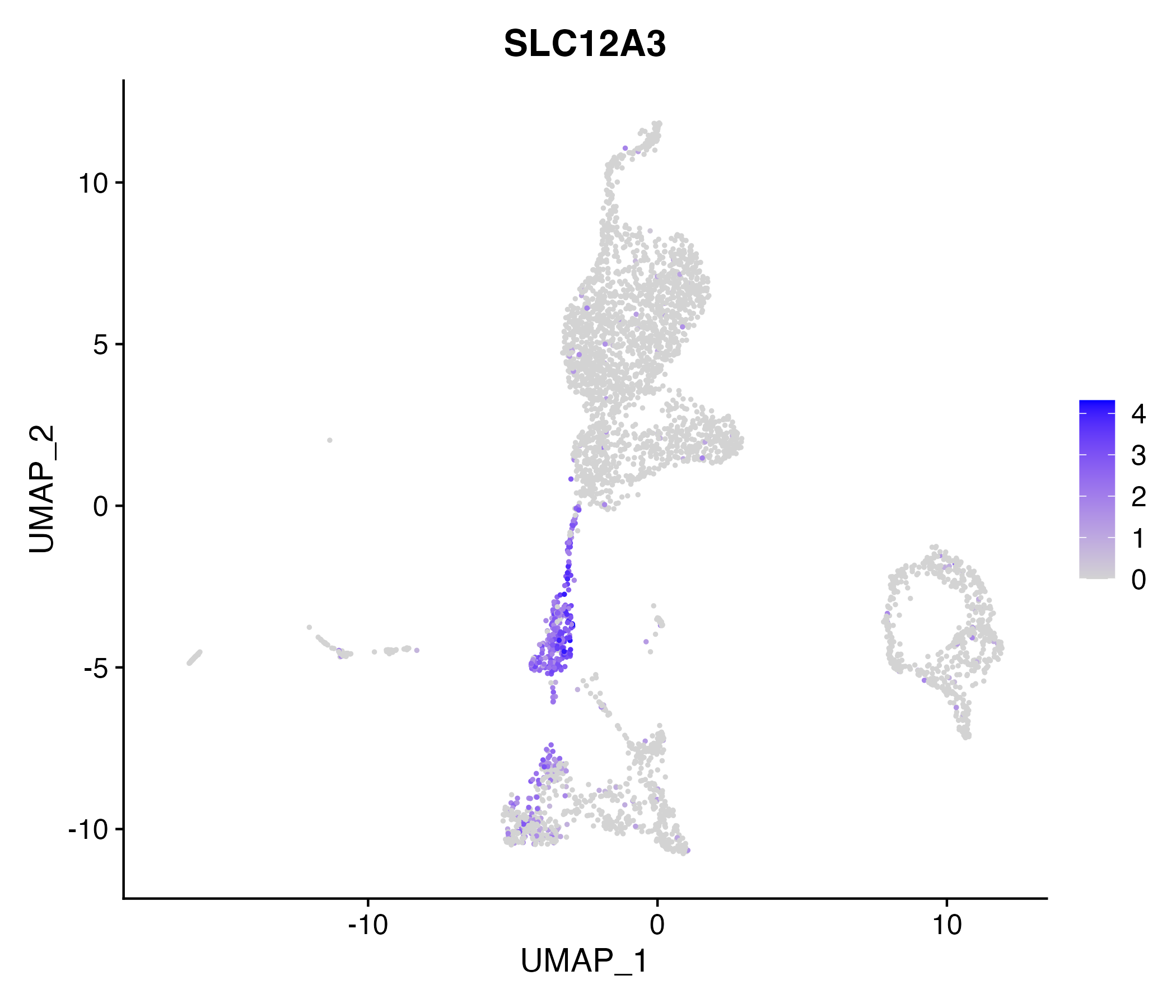
|
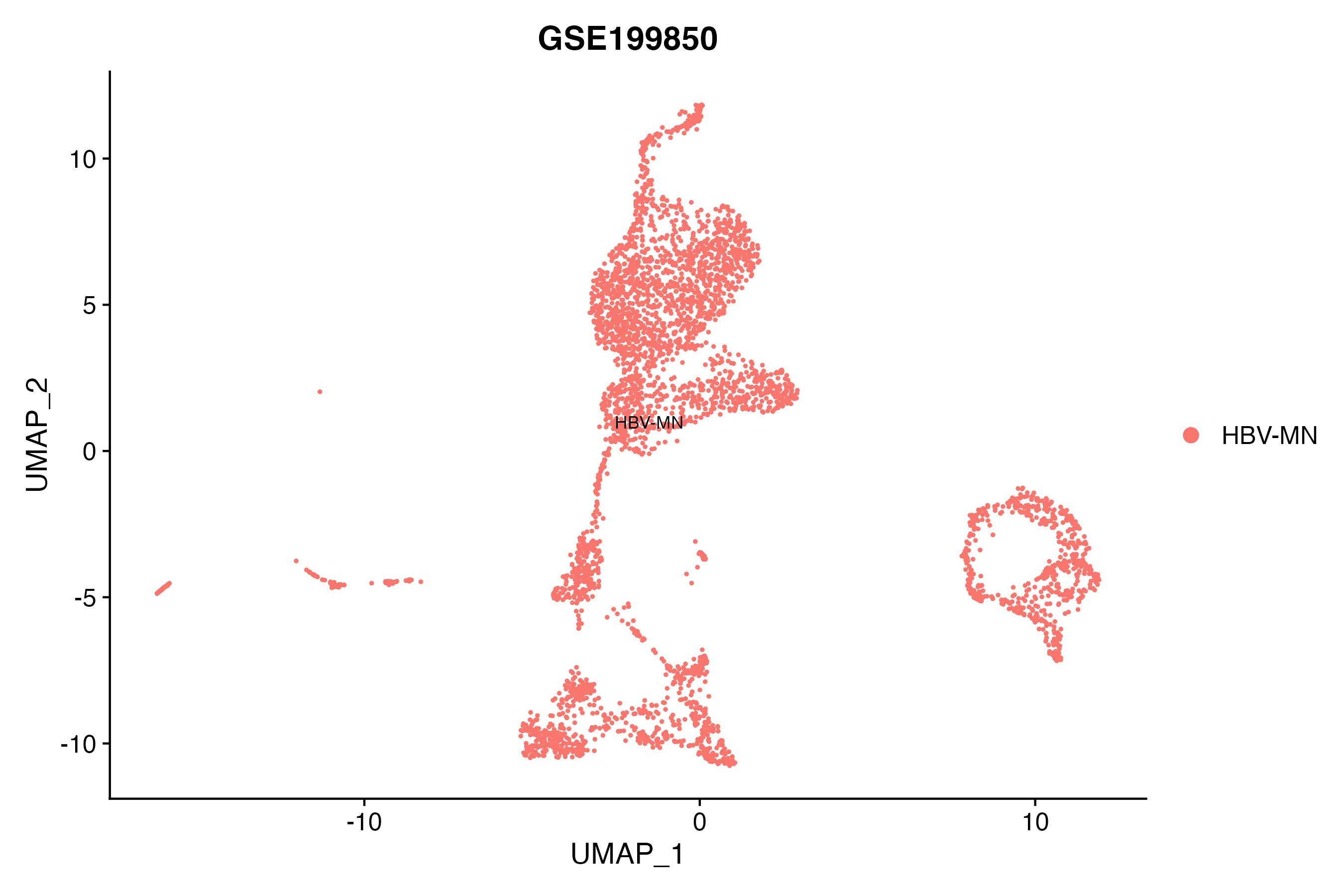
|
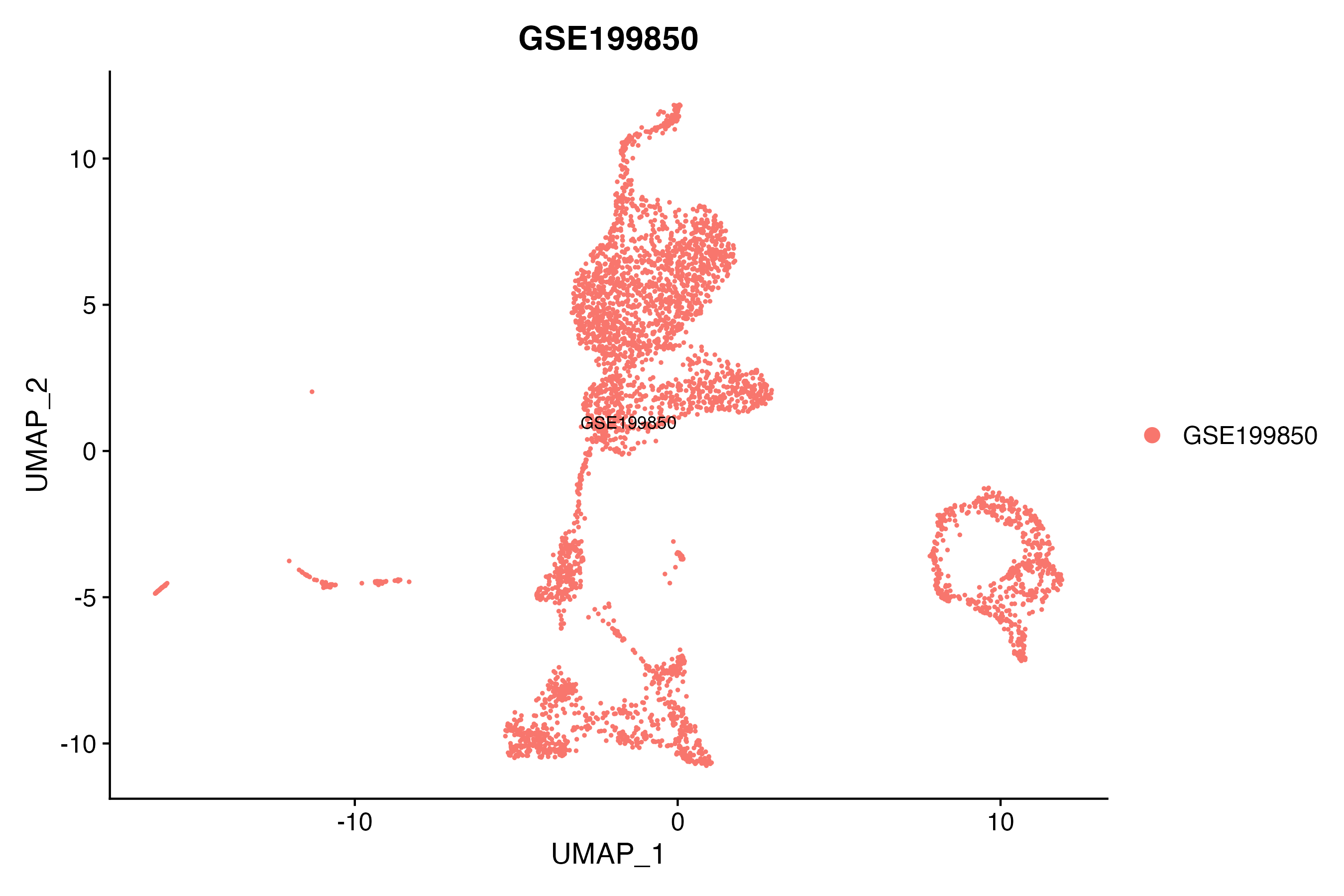
|
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information