GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0001562 | Response to protozoan |
| Process | GO:0001775 | Cell activation |
| Process | GO:0001817 | Regulation of cytokine production |
| Process | GO:0001818 | Negative regulation of cytokine production |
| Process | GO:0001819 | Positive regulation of cytokine production |
| Process | GO:0001910 | Regulation of leukocyte mediated cytotoxicity |
| Process | GO:0001912 | Positive regulation of leukocyte mediated cytotoxicity |
| Process | GO:0001914 | Regulation of T cell mediated cytotoxicity |
| Process | GO:0001916 | Positive regulation of T cell mediated cytotoxicity |
| Process | GO:0001932 | Regulation of protein phosphorylation |
| Process | GO:0001934 | Positive regulation of protein phosphorylation |
| Process | GO:0001936 | Regulation of endothelial cell proliferation |
| Process | GO:0001937 | Negative regulation of endothelial cell proliferation |
| Process | GO:0002230 | Positive regulation of defense response to virus by host |
| Process | GO:0002237 | Response to molecule of bacterial origin |
| Process | GO:0002252 | Immune effector process |
| Process | GO:0002263 | Cell activation involved in immune response |
| Process | GO:0002285 | Lymphocyte activation involved in immune response |
| Process | GO:0002286 | T cell activation involved in immune response |
| Process | GO:0002287 | Alpha-beta T cell activation involved in immune response |
| Process | GO:0002292 | T cell differentiation involved in immune response |
| Process | GO:0002293 | Alpha-beta T cell differentiation involved in immune response |
| Process | GO:0002294 | CD4-positive, alpha-beta T cell differentiation involved in immune response |
| Process | GO:0002323 | Natural killer cell activation involved in immune response |
| Process | GO:0002366 | Leukocyte activation involved in immune response |
| Process | GO:0002376 | Immune system process |
| Process | GO:0002520 | Immune system development |
| Process | GO:0002521 | Leukocyte differentiation |
| Process | GO:0002682 | Regulation of immune system process |
| Process | GO:0002683 | Negative regulation of immune system process |
| Process | GO:0002684 | Positive regulation of immune system process |
| Process | GO:0002694 | Regulation of leukocyte activation |
| Process | GO:0002696 | Positive regulation of leukocyte activation |
| Process | GO:0002697 | Regulation of immune effector process |
| Process | GO:0002699 | Positive regulation of immune effector process |
| Process | GO:0002703 | Regulation of leukocyte mediated immunity |
| Process | GO:0002705 | Positive regulation of leukocyte mediated immunity |
| Process | GO:0002706 | Regulation of lymphocyte mediated immunity |
| Process | GO:0002708 | Positive regulation of lymphocyte mediated immunity |
| Process | GO:0002709 | Regulation of T cell mediated immunity |
| Process | GO:0002711 | Positive regulation of T cell mediated immunity |
| Process | GO:0002715 | Regulation of natural killer cell mediated immunity |
| Process | GO:0002717 | Positive regulation of natural killer cell mediated immunity |
| Process | GO:0002761 | Regulation of myeloid leukocyte differentiation |
| Process | GO:0002763 | Positive regulation of myeloid leukocyte differentiation |
| Process | GO:0002819 | Regulation of adaptive immune response |
| Process | GO:0002821 | Positive regulation of adaptive immune response |
| Process | GO:0002822 | Regulation of adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains |
| Process | GO:0002824 | Positive regulation of adaptive immune response based on somatic recombination of immune receptors built from immunoglobulin superfamily domains |
| Process | GO:0002825 | Regulation of T-helper 1 type immune response |
| Process | GO:0002827 | Positive regulation of T-helper 1 type immune response |
| Process | GO:0002831 | Regulation of response to biotic stimulus |
| Process | GO:0002833 | Positive regulation of response to biotic stimulus |
| Process | GO:0002834 | Regulation of response to tumor cell |
| Process | GO:0002836 | Positive regulation of response to tumor cell |
| Process | GO:0002837 | Regulation of immune response to tumor cell |
| Process | GO:0002839 | Positive regulation of immune response to tumor cell |
| Process | GO:0002855 | Regulation of natural killer cell mediated immune response to tumor cell |
| Process | GO:0002857 | Positive regulation of natural killer cell mediated immune response to tumor cell |
| Process | GO:0002858 | Regulation of natural killer cell mediated cytotoxicity directed against tumor cell target |
| Process | GO:0002860 | Positive regulation of natural killer cell mediated cytotoxicity directed against tumor cell target |
| Process | GO:0002861 | Regulation of inflammatory response to antigenic stimulus |
| Process | GO:0002862 | Negative regulation of inflammatory response to antigenic stimulus |
| Process | GO:0003008 | System process |
| Process | GO:0006950 | Response to stress |
| Process | GO:0006952 | Defense response |
| Process | GO:0006955 | Immune response |
| Process | GO:0007154 | Cell communication |
| Process | GO:0007165 | Signal transduction |
| Process | GO:0007166 | Cell surface receptor signaling pathway |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0007600 | Sensory perception |
| Process | GO:0008283 | Cell population proliferation |
| Process | GO:0008284 | Positive regulation of cell population proliferation |
| Process | GO:0008285 | Negative regulation of cell population proliferation |
| Process | GO:0009314 | Response to radiation |
| Process | GO:0009411 | Response to UV |
| Process | GO:0009416 | Response to light stimulus |
| Process | GO:0009605 | Response to external stimulus |
| Process | GO:0009607 | Response to biotic stimulus |
| Process | GO:0009615 | Response to virus |
| Process | GO:0009617 | Response to bacterium |
| Process | GO:0009628 | Response to abiotic stimulus |
| Process | GO:0009892 | Negative regulation of metabolic process |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009966 | Regulation of signal transduction |
| Process | GO:0009967 | Positive regulation of signal transduction |
| Process | GO:0009968 | Negative regulation of signal transduction |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010033 | Response to organic substance |
| Process | GO:0010224 | Response to UV-B |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010533 | Regulation of activation of Janus kinase activity |
| Process | GO:0010536 | Positive regulation of activation of Janus kinase activity |
| Process | GO:0010562 | Positive regulation of phosphorus metabolic process |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0010605 | Negative regulation of macromolecule metabolic process |
| Process | GO:0010628 | Positive regulation of gene expression |
| Process | GO:0010629 | Negative regulation of gene expression |
| Process | GO:0010646 | Regulation of cell communication |
| Process | GO:0010647 | Positive regulation of cell communication |
| Process | GO:0010648 | Negative regulation of cell communication |
| Process | GO:0010660 | Regulation of muscle cell apoptotic process |
| Process | GO:0010661 | Positive regulation of muscle cell apoptotic process |
| Process | GO:0010941 | Regulation of cell death |
| Process | GO:0010942 | Positive regulation of cell death |
| Process | GO:0016477 | Cell migration |
| Process | GO:0019220 | Regulation of phosphate metabolic process |
| Process | GO:0019221 | Cytokine-mediated signaling pathway |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0019233 | Sensory perception of pain |
| Process | GO:0022407 | Regulation of cell-cell adhesion |
| Process | GO:0022409 | Positive regulation of cell-cell adhesion |
| Process | GO:0023051 | Regulation of signaling |
| Process | GO:0023052 | Signaling |
| Process | GO:0023056 | Positive regulation of signaling |
| Process | GO:0023057 | Negative regulation of signaling |
| Process | GO:0030097 | Hemopoiesis |
| Process | GO:0030098 | Lymphocyte differentiation |
| Process | GO:0030101 | Natural killer cell activation |
| Process | GO:0030154 | Cell differentiation |
| Process | GO:0030155 | Regulation of cell adhesion |
| Process | GO:0030217 | T cell differentiation |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0031325 | Positive regulation of cellular metabolic process |
| Process | GO:0031341 | Regulation of cell killing |
| Process | GO:0031343 | Positive regulation of cell killing |
| Process | GO:0031347 | Regulation of defense response |
| Process | GO:0031348 | Negative regulation of defense response |
| Process | GO:0031349 | Positive regulation of defense response |
| Process | GO:0031399 | Regulation of protein modification process |
| Process | GO:0031401 | Positive regulation of protein modification process |
| Process | GO:0032101 | Regulation of response to external stimulus |
| Process | GO:0032102 | Negative regulation of response to external stimulus |
| Process | GO:0032103 | Positive regulation of response to external stimulus |
| Process | GO:0032496 | Response to lipopolysaccharide |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032645 | Regulation of granulocyte macrophage colony-stimulating factor production |
| Process | GO:0032649 | Regulation of interferon-gamma production |
| Process | GO:0032653 | Regulation of interleukin-10 production |
| Process | GO:0032655 | Regulation of interleukin-12 production |
| Process | GO:0032660 | Regulation of interleukin-17 production |
| Process | GO:0032680 | Regulation of tumor necrosis factor production |
| Process | GO:0032693 | Negative regulation of interleukin-10 production |
| Process | GO:0032700 | Negative regulation of interleukin-17 production |
| Process | GO:0032725 | Positive regulation of granulocyte macrophage colony-stimulating factor production |
| Process | GO:0032729 | Positive regulation of interferon-gamma production |
| Process | GO:0032733 | Positive regulation of interleukin-10 production |
| Process | GO:0032735 | Positive regulation of interleukin-12 production |
| Process | GO:0032740 | Positive regulation of interleukin-17 production |
| Process | GO:0032760 | Positive regulation of tumor necrosis factor production |
| Process | GO:0032814 | Regulation of natural killer cell activation |
| Process | GO:0032816 | Positive regulation of natural killer cell activation |
| Process | GO:0032817 | Regulation of natural killer cell proliferation |
| Process | GO:0032819 | Positive regulation of natural killer cell proliferation |
| Process | GO:0032879 | Regulation of localization |
| Process | GO:0032880 | Regulation of protein localization |
| Process | GO:0032943 | Mononuclear cell proliferation |
| Process | GO:0032944 | Regulation of mononuclear cell proliferation |
| Process | GO:0032946 | Positive regulation of mononuclear cell proliferation |
| Process | GO:0033674 | Positive regulation of kinase activity |
| Process | GO:0033993 | Response to lipid |
| Process | GO:0034097 | Response to cytokine |
| Process | GO:0034341 | Response to interferon-gamma |
| Process | GO:0034391 | Regulation of smooth muscle cell apoptotic process |
| Process | GO:0034393 | Positive regulation of smooth muscle cell apoptotic process |
| Process | GO:0035710 | CD4-positive, alpha-beta T cell activation |
| Process | GO:0042093 | T-helper cell differentiation |
| Process | GO:0042098 | T cell proliferation |
| Process | GO:0042102 | Positive regulation of T cell proliferation |
| Process | GO:0042104 | Positive regulation of activated T cell proliferation |
| Process | GO:0042110 | T cell activation |
| Process | GO:0042127 | Regulation of cell population proliferation |
| Process | GO:0042129 | Regulation of T cell proliferation |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0042269 | Regulation of natural killer cell mediated cytotoxicity |
| Process | GO:0042325 | Regulation of phosphorylation |
| Process | GO:0042327 | Positive regulation of phosphorylation |
| Process | GO:0042509 | Regulation of tyrosine phosphorylation of STAT protein |
| Process | GO:0042531 | Positive regulation of tyrosine phosphorylation of STAT protein |
| Process | GO:0042742 | Defense response to bacterium |
| Process | GO:0042832 | Defense response to protozoan |
| Process | GO:0042981 | Regulation of apoptotic process |
| Process | GO:0043065 | Positive regulation of apoptotic process |
| Process | GO:0043067 | Regulation of programmed cell death |
| Process | GO:0043068 | Positive regulation of programmed cell death |
| Process | GO:0043085 | Positive regulation of catalytic activity |
| Process | GO:0043207 | Response to external biotic stimulus |
| Process | GO:0043367 | CD4-positive, alpha-beta T cell differentiation |
| Process | GO:0043549 | Regulation of kinase activity |
| Process | GO:0044093 | Positive regulation of molecular function |
| Process | GO:0044419 | Biological process involved in interspecies interaction between organisms |
| Process | GO:0045087 | Innate immune response |
| Process | GO:0045088 | Regulation of innate immune response |
| Process | GO:0045089 | Positive regulation of innate immune response |
| Process | GO:0045321 | Leukocyte activation |
| Process | GO:0045595 | Regulation of cell differentiation |
| Process | GO:0045597 | Positive regulation of cell differentiation |
| Process | GO:0045637 | Regulation of myeloid cell differentiation |
| Process | GO:0045639 | Positive regulation of myeloid cell differentiation |
| Process | GO:0045670 | Regulation of osteoclast differentiation |
| Process | GO:0045672 | Positive regulation of osteoclast differentiation |
| Process | GO:0045785 | Positive regulation of cell adhesion |
| Process | GO:0045859 | Regulation of protein kinase activity |
| Process | GO:0045860 | Positive regulation of protein kinase activity |
| Process | GO:0045937 | Positive regulation of phosphate metabolic process |
| Process | GO:0045954 | Positive regulation of natural killer cell mediated cytotoxicity |
| Process | GO:0046006 | Regulation of activated T cell proliferation |
| Process | GO:0046425 | Regulation of receptor signaling pathway via JAK-STAT |
| Process | GO:0046427 | Positive regulation of receptor signaling pathway via JAK-STAT |
| Process | GO:0046631 | Alpha-beta T cell activation |
| Process | GO:0046632 | Alpha-beta T cell differentiation |
| Process | GO:0046634 | Regulation of alpha-beta T cell activation |
| Process | GO:0046635 | Positive regulation of alpha-beta T cell activation |
| Process | GO:0046640 | Regulation of alpha-beta T cell proliferation |
| Process | GO:0046641 | Positive regulation of alpha-beta T cell proliferation |
| Process | GO:0046649 | Lymphocyte activation |
| Process | GO:0046651 | Lymphocyte proliferation |
| Process | GO:0048513 | Animal organ development |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048519 | Negative regulation of biological process |
| Process | GO:0048522 | Positive regulation of cellular process |
| Process | GO:0048523 | Negative regulation of cellular process |
| Process | GO:0048534 | Hematopoietic or lymphoid organ development |
| Process | GO:0048583 | Regulation of response to stimulus |
| Process | GO:0048584 | Positive regulation of response to stimulus |
| Process | GO:0048585 | Negative regulation of response to stimulus |
| Process | GO:0048660 | Regulation of smooth muscle cell proliferation |
| Process | GO:0048662 | Negative regulation of smooth muscle cell proliferation |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0048869 | Cellular developmental process |
| Process | GO:0048870 | Cell motility |
| Process | GO:0050670 | Regulation of lymphocyte proliferation |
| Process | GO:0050671 | Positive regulation of lymphocyte proliferation |
| Process | GO:0050678 | Regulation of epithelial cell proliferation |
| Process | GO:0050680 | Negative regulation of epithelial cell proliferation |
| Process | GO:0050688 | Regulation of defense response to virus |
| Process | GO:0050691 | Regulation of defense response to virus by host |
| Process | GO:0050708 | Regulation of protein secretion |
| Process | GO:0050709 | Negative regulation of protein secretion |
| Process | GO:0050727 | Regulation of inflammatory response |
| Process | GO:0050728 | Negative regulation of inflammatory response |
| Process | GO:0050730 | Regulation of peptidyl-tyrosine phosphorylation |
| Process | GO:0050731 | Positive regulation of peptidyl-tyrosine phosphorylation |
| Process | GO:0050776 | Regulation of immune response |
| Process | GO:0050777 | Negative regulation of immune response |
| Process | GO:0050778 | Positive regulation of immune response |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050790 | Regulation of catalytic activity |
| Process | GO:0050793 | Regulation of developmental process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050829 | Defense response to Gram-negative bacterium |
| Process | GO:0050863 | Regulation of T cell activation |
| Process | GO:0050865 | Regulation of cell activation |
| Process | GO:0050867 | Positive regulation of cell activation |
| Process | GO:0050870 | Positive regulation of T cell activation |
| Process | GO:0050877 | Nervous system process |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0051046 | Regulation of secretion |
| Process | GO:0051048 | Negative regulation of secretion |
| Process | GO:0051049 | Regulation of transport |
| Process | GO:0051051 | Negative regulation of transport |
| Process | GO:0051094 | Positive regulation of developmental process |
| Process | GO:0051133 | Regulation of NK T cell activation |
| Process | GO:0051135 | Positive regulation of NK T cell activation |
| Process | GO:0051140 | Regulation of NK T cell proliferation |
| Process | GO:0051142 | Positive regulation of NK T cell proliferation |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051173 | Positive regulation of nitrogen compound metabolic process |
| Process | GO:0051174 | Regulation of phosphorus metabolic process |
| Process | GO:0051223 | Regulation of protein transport |
| Process | GO:0051224 | Negative regulation of protein transport |
| Process | GO:0051239 | Regulation of multicellular organismal process |
| Process | GO:0051240 | Positive regulation of multicellular organismal process |
| Process | GO:0051241 | Negative regulation of multicellular organismal process |
| Process | GO:0051246 | Regulation of protein metabolic process |
| Process | GO:0051247 | Positive regulation of protein metabolic process |
| Process | GO:0051249 | Regulation of lymphocyte activation |
| Process | GO:0051251 | Positive regulation of lymphocyte activation |
| Process | GO:0051338 | Regulation of transferase activity |
| Process | GO:0051347 | Positive regulation of transferase activity |
| Process | GO:0051607 | Defense response to virus |
| Process | GO:0051707 | Response to other organism |
| Process | GO:0051716 | Cellular response to stimulus |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0060341 | Regulation of cellular localization |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065009 | Regulation of molecular function |
| Process | GO:0070201 | Regulation of establishment of protein localization |
| Process | GO:0070661 | Leukocyte proliferation |
| Process | GO:0070663 | Regulation of leukocyte proliferation |
| Process | GO:0070665 | Positive regulation of leukocyte proliferation |
| Process | GO:0070887 | Cellular response to chemical stimulus |
| Process | GO:0071216 | Cellular response to biotic stimulus |
| Process | GO:0071219 | Cellular response to molecule of bacterial origin |
| Process | GO:0071222 | Cellular response to lipopolysaccharide |
| Process | GO:0071310 | Cellular response to organic substance |
| Process | GO:0071345 | Cellular response to cytokine stimulus |
| Process | GO:0071346 | Cellular response to interferon-gamma |
| Process | GO:0071396 | Cellular response to lipid |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0080134 | Regulation of response to stress |
| Process | GO:0090287 | Regulation of cellular response to growth factor stimulus |
| Process | GO:0090288 | Negative regulation of cellular response to growth factor stimulus |
| Process | GO:0098542 | Defense response to other organism |
| Process | GO:0140546 | Defense response to symbiont |
| Process | GO:1900746 | Regulation of vascular endothelial growth factor signaling pathway |
| Process | GO:1900747 | Negative regulation of vascular endothelial growth factor signaling pathway |
| Process | GO:1901700 | Response to oxygen-containing compound |
| Process | GO:1901701 | Cellular response to oxygen-containing compound |
| Process | GO:1902105 | Regulation of leukocyte differentiation |
| Process | GO:1902107 | Positive regulation of leukocyte differentiation |
| Process | GO:1902547 | Regulation of cellular response to vascular endothelial growth factor stimulus |
| Process | GO:1902548 | Negative regulation of cellular response to vascular endothelial growth factor stimulus |
| Process | GO:1903037 | Regulation of leukocyte cell-cell adhesion |
| Process | GO:1903039 | Positive regulation of leukocyte cell-cell adhesion |
| Process | GO:1903131 | Mononuclear cell differentiation |
| Process | GO:1903530 | Regulation of secretion by cell |
| Process | GO:1903531 | Negative regulation of secretion by cell |
| Process | GO:1903555 | Regulation of tumor necrosis factor superfamily cytokine production |
| Process | GO:1903557 | Positive regulation of tumor necrosis factor superfamily cytokine production |
| Process | GO:1903587 | Regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis |
| Process | GO:1903588 | Negative regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis |
| Process | GO:1903706 | Regulation of hemopoiesis |
| Process | GO:1903708 | Positive regulation of hemopoiesis |
| Process | GO:1903828 | Negative regulation of protein localization |
| Process | GO:1904892 | Regulation of receptor signaling pathway via STAT |
| Process | GO:1904894 | Positive regulation of receptor signaling pathway via STAT |
| Process | GO:1904950 | Negative regulation of establishment of protein localization |
| Process | GO:2000026 | Regulation of multicellular organismal development |
| Component | GO:0005576 | Extracellular region |
| Component | GO:0005615 | Extracellular space |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005737 | Cytoplasm |
| Component | GO:0005886 | Plasma membrane |
| Component | GO:0009897 | External side of plasma membrane |
| Component | GO:0009986 | Cell surface |
| Component | GO:0016020 | Membrane |
| Component | GO:0032991 | Protein-containing complex |
| Component | GO:0043235 | Receptor complex |
| Component | GO:0043514 | interleukin-12 complex |
| Component | GO:0070743 | interleukin-23 complex |
| Component | GO:0071944 | Cell periphery |
| Component | GO:0098552 | Side of membrane |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0004888 | Transmembrane signaling receptor activity |
| Function | GO:0004896 | Cytokine receptor activity |
| Function | GO:0005102 | Signaling receptor binding |
| Function | GO:0005125 | Cytokine activity |
| Function | GO:0005126 | Cytokine receptor binding |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0008083 | Growth factor activity |
| Function | GO:0019955 | Cytokine binding |
| Function | GO:0019972 | interleukin-12 binding |
| Function | GO:0030545 | Signaling receptor regulator activity |
| Function | GO:0030546 | Signaling receptor activator activity |
| Function | GO:0038023 | Signaling receptor activity |
| Function | GO:0042802 | Identical protein binding |
| Function | GO:0044877 | Protein-containing complex binding |
| Function | GO:0045519 | interleukin-23 receptor binding |
| Function | GO:0046982 | Protein heterodimerization activity |
| Function | GO:0046983 | Protein dimerization activity |
| Function | GO:0048018 | Receptor ligand activity |
| Function | GO:0060089 | Molecular transducer activity |
| Function | GO:0098772 | Molecular function regulator activity |
| Function | GO:0140375 | Immune receptor activity |
| Function | GO:0042164 | interleukin-12 alpha subunit binding |
Pathway annotation
Expression profile of Il12b in omics data
| ID | GSE100499 |
| Title | Exploring acupuncture mechanism in preconditioning of IRI |
| Organism | Rattus norvegicus |
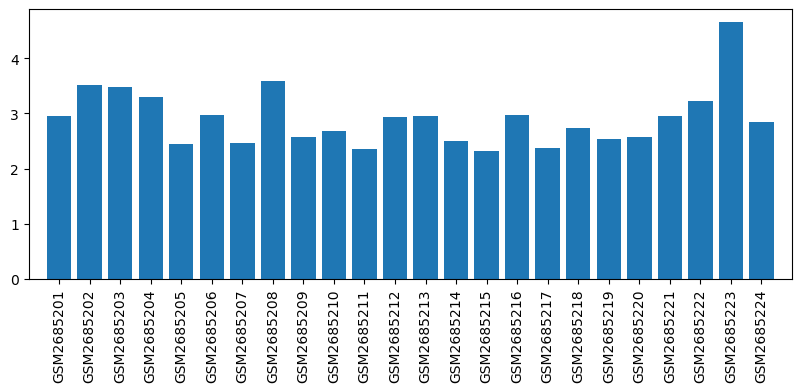
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2685201 | MX1 | tissue: Myocardium gender: male | 2.9610837 |
| GSM2685202 | MX2 | tissue: Myocardium gender: female | 3.5082302 |
| GSM2685203 | MX3 | tissue: Myocardium gender: female | 3.4824057 |
| GSM2685204 | MX4 | tissue: Myocardium gender: male | 3.2944362 |
| GSM2685205 | DZ1 | tissue: Myocardium gender: male | 2.452026 |
| GSM2685206 | DZ2 | tissue: Myocardium gender: male | 2.9656067 |
| GSM2685207 | DZ3 | tissue: Myocardium gender: female | 2.457881 |
| GSM2685208 | DZ4 | tissue: Myocardium gender: female | 3.5847838 |
| GSM2685209 | JJ1 | tissue: Myocardium gender: male | 2.570754 |
| GSM2685210 | JJ2 | tissue: Myocardium gender: female | 2.6776316 |
| GSM2685211 | JJ3 | tissue: Myocardium gender: female | 2.351032 |
| GSM2685212 | JJ4 | tissue: Myocardium gender: male | 2.9299695 |
| GSM2685213 | NG1 | tissue: Myocardium gender: male | 2.955559 |
| GSM2685214 | NG2 | tissue: Myocardium gender: female | 2.5001323 |
| GSM2685215 | NG3 | tissue: Myocardium gender: male | 2.3262157 |
| GSM2685216 | NG4 | tissue: Myocardium gender: female | 2.9670217 |
| GSM2685217 | YL1 | tissue: Myocardium gender: male | 2.3701637 |
| GSM2685218 | YL2 | tissue: Myocardium gender: male | 2.740888 |
| GSM2685219 | YL3 | tissue: Myocardium gender: female | 2.5313566 |
| GSM2685220 | YL4 | tissue: Myocardium gender: female | 2.5710938 |
| GSM2685221 | QC1 | tissue: Myocardium gender: male | 2.9538667 |
| GSM2685222 | QC2 | tissue: Myocardium gender: male | 3.2197704 |
| GSM2685223 | QC3 | tissue: Myocardium gender: female | 4.6611238 |
| GSM2685224 | QC4 | tissue: Myocardium gender: female | 2.849765 |
| ID | GSE136833 |
| Title | Effects of lidocaine on the expression of rat spinal cord |
| Organism | Rattus norvegicus |
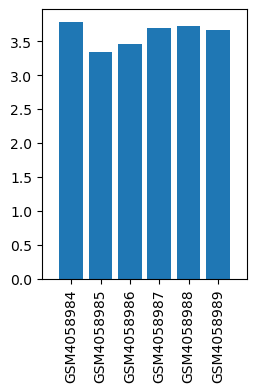
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4058984 | Control_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 3.784717173 |
| GSM4058985 | Control_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 3.347299089 |
| GSM4058986 | Control_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 3.464961624 |
| GSM4058987 | Lidocaine_1 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 3.701028421 |
| GSM4058988 | Lidocaine_2 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 3.73270197 |
| GSM4058989 | Lidocaine_3 | strain: Sprague-Dawley tissue: Lumbar segment of spinal cord | 3.673901752 |
| ID | GSE138454 |
| Title | Impact of inflammation on hepatic subcellular energetics in anesthetized rats |
| Organism | Rattus norvegicus |
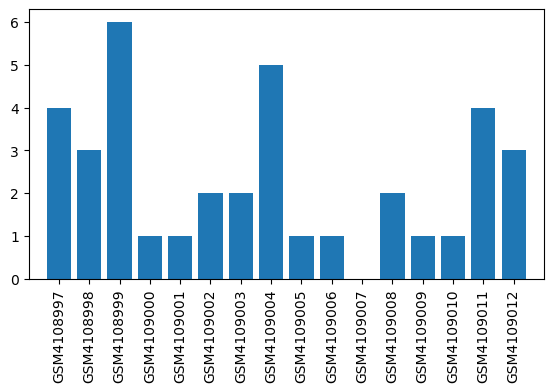
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4108997 | Sample 1 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 4.0 |
| GSM4108998 | Sample 2 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 3.0 |
| GSM4108999 | Sample 3 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 6.0 |
| GSM4109000 | Sample 4 Isoflurane sepsis | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 1.0 |
| GSM4109001 | Sample 5 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 1.0 |
| GSM4109002 | Sample 6 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 2.0 |
| GSM4109003 | Sample 7 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 2.0 |
| GSM4109004 | Sample 8 Isoflurane control | anesthetic background: Isoflurane strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 5.0 |
| GSM4109005 | Sample9 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 1.0 |
| GSM4109006 | Sample10 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 1.0 |
| GSM4109007 | Sample11 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 0.0 |
| GSM4109008 | Sample12 Propofol sepsis | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: sepsis gender: male | 2.0 |
| GSM4109009 | Sample13 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 1.0 |
| GSM4109010 | Sample14 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 1.0 |
| GSM4109011 | Sample 15 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 4.0 |
| GSM4109012 | Sample16 Propofol control | anesthetic background: Propofol strain: Sprague-Dawley age: 10 weeks old sepsis/control: control gender: male | 3.0 |
| ID | GSE139220 |
| Title | Expression data from sevoflurane anesthesia treated rat |
| Organism | Rattus norvegicus |
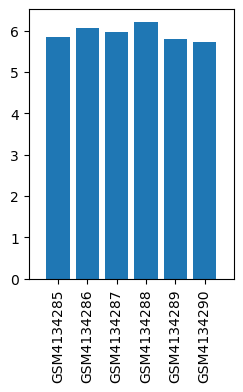
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4134285 | Brain_Control_rep1 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 5.856385 |
| GSM4134286 | Brain_Control_rep2 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 6.064647 |
| GSM4134287 | Brain_Control_rep3 | strain: Sprague-Dawley age: 18-month old treatment: control tissue: hippocampus | 5.970334 |
| GSM4134288 | Brain_Sevoflurane_rep1 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 6.212765 |
| GSM4134289 | Brain_Sevoflurane_rep2 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 5.808792 |
| GSM4134290 | Brain_Sevoflurane_rep3 | strain: Sprague-Dawley age: 18-month old treatment: sevoflurane anesthesia tissue: hippocampus | 5.729115 |
| ID | GSE143895 |
| Title | GPR160 de-orphanization reveals critical roles in neuropathic pain in rodents |
| Organism | Rattus norvegicus |
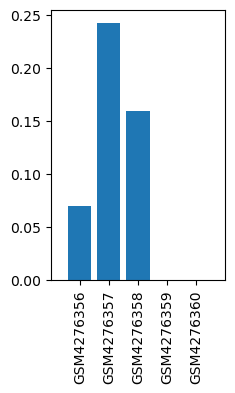
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4276356 | dhsc01 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.069 |
| GSM4276357 | dhsc03 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.242 |
| GSM4276358 | dhsc05 | strain: Sprague Dawley tissue: DHSC treatment: CCI | 0.159 |
| GSM4276359 | dhsc13 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 0.0 |
| GSM4276360 | dhsc15 | strain: Sprague Dawley tissue: DHSC treatment: SHAM | 0.0 |
| ID | GSE157924 |
| Title | Regulation of acupuncture on genes in brain stem |
| Organism | Rattus norvegicus |
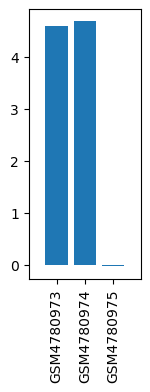
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4780973 | Rattus_JSSZ(Sham) | tissue: Rattus brainstem of modeling treatment: Sham | 4.595996282 |
| GSM4780974 | Rattus_JXZ(Hand Acu) | tissue: Rattus brainstem of modeling treatment: Hand Acu | 4.685510175 |
| GSM4780975 | Rattus_MXZ(TBI) | tissue: Rattus brainstem of modeling treatment: TBI | -0.027363416 |
| ID | GSE1616 |
| Title | Genomics of preconditioning |
| Organism | Rattus norvegicus |
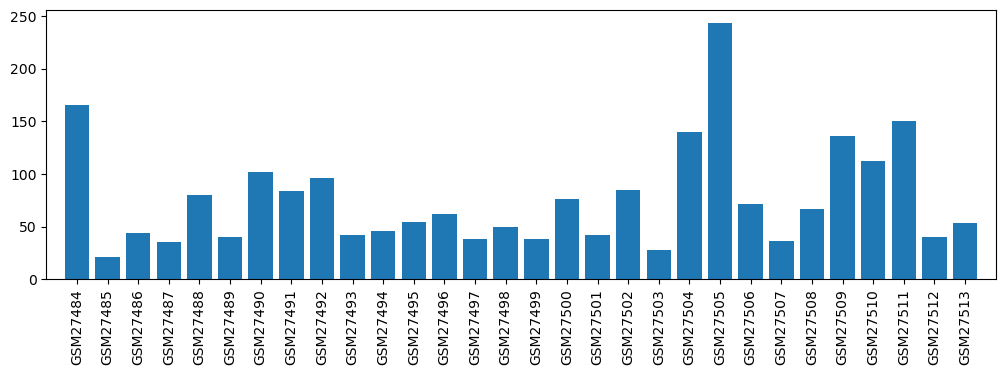
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM27484 | APC_a (anesthetic-preconditioning, 1st replicate) | 165.45 | |
| GSM27485 | APC_b (anesthetic-preconditioning, 2nd replicate) | 21.55 | |
| GSM27486 | APC_c (anesthetic-preconditioning, 3rd replicate) | 44.3 | |
| GSM27487 | APC_d (anesthetic-preconditioning, 4th replicate) | 35.35 | |
| GSM27488 | APC_e (anesthetic-preconditioning, 5th replicate) | 80.15 | |
| GSM27489 | APC_TRI_a (trigger of anesthetic-preconditioning, 1st replicate) | 39.9 | |
| GSM27490 | APC_TRI_b (trigger of anesthetic-preconditioning, 2nd replicate) | 101.55 | |
| GSM27491 | APC_TRI_c (trigger of anesthetic-preconditioning, 3rd replicate) | 84.0 | |
| GSM27492 | APC_TRI_d (trigger of anesthetic-preconditioning, 4th replicate) | 96.1 | |
| GSM27493 | APC_TRI_e (trigger of anesthetic-preconditioning, 5th replicate) | 41.7 | |
| GSM27494 | CTL_a (control, 1st replicate) | 46.15 | |
| GSM27495 | CTL_b (control, 2nd replicate) | 54.849999999999994 | |
| GSM27496 | CTL_c (control, 3rd replicate) | 62.45 | |
| GSM27497 | CTL_d (control, 4th replicate) | 38.5 | |
| GSM27498 | CTL_e (control, 5th replicate) | 50.05 | |
| GSM27499 | IPC_a (ischemic preconditioning, 1st replicate) | 38.1 | |
| GSM27500 | IPC_b (ischemic preconditioning, 2nd replicate) | 76.7 | |
| GSM27501 | IPC_c (ischemic preconditioning, 3rd replicate) | 42.400000000000006 | |
| GSM27502 | IPC_d (ischemic preconditioning, 4th replicate) | 84.45 | |
| GSM27503 | IPC_e (ischemic preconditioning, 5th replicate) | 27.75 | |
| GSM27504 | IPC_TRI_a (trigger of ischemic preconditioning, 1st replicate) | 139.45 | |
| GSM27505 | IPC_TRI_b (trigger of ischemic preconditioning, 2nd replicate) | 243.6 | |
| GSM27506 | IPC_TRI_c (trigger of ischemic preconditioning, 3rd replicate) | 71.65 | |
| GSM27507 | IPC_TRI_d (trigger of ischemic preconditioning, 4th replicate) | 36.3 | |
| GSM27508 | IPC_TRI_e (trigger of ischemic preconditioning, 5th replicate) | 67.05 | |
| GSM27509 | ISCH_a (ischemia, 1st replicate) | 136.2 | |
| GSM27510 | ISCH_b (ischemia, 2nd replicate) | 112.75 | |
| GSM27511 | ISCH_c (ischemia, 3rd replicate) | 150.25 | |
| GSM27512 | ISCH_d (ischemia, 4th replicate) | 40.5 | |
| GSM27513 | ISCH_e (ischemia, 5th replicate) | 53.2 |
| ID | GSE1779 |
| Title | Effect of Isoflurane and LTM on basolateral amygdala |
| Organism | Rattus norvegicus |
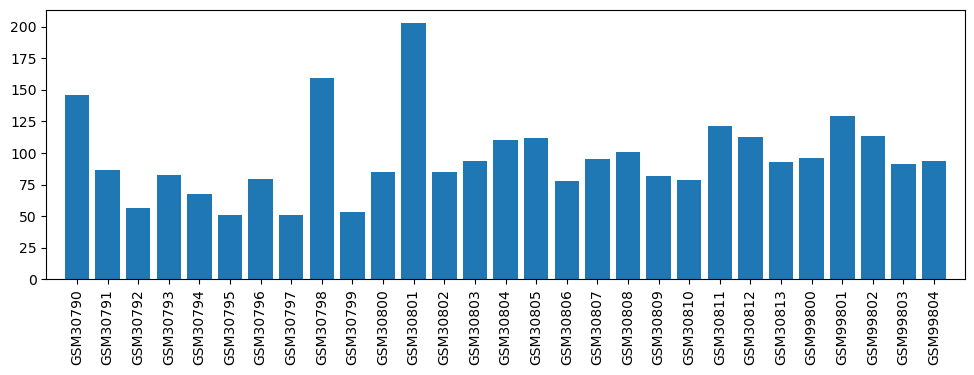
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM30790 | baseline control 01 | male, 275-300g | 145.9 |
| GSM30791 | baseline control 02 | male, 275-300g | 86.5 |
| GSM30792 | baseline control 03 | male, 275-300g | 56.6 |
| GSM30793 | baseline control 04 | male, 275-300g | 82.8 |
| GSM30794 | baseline control 05 | male, 275-300g | 67.5 |
| GSM30795 | baseline control 06 | male, 275-300g | 51.3 |
| GSM30796 | baseline control 07 | male, 275-300g | 79.3 |
| GSM30797 | baseline control 08 | male, 275-300g | 50.7 |
| GSM30798 | baseline control 09 | male, 275-300g | 159.2 |
| GSM30799 | pain 01 | 53.4 | |
| GSM30800 | pain 02 | 85.2 | |
| GSM30801 | pain 03 | 203.1 | |
| GSM30802 | pain 04 | 84.9 | |
| GSM30803 | pain 05 | 94.0 | |
| GSM30804 | FC 01 | 110.6 | |
| GSM30805 | FC 02 | 111.9 | |
| GSM30806 | FC 03 | 77.8 | |
| GSM30807 | FC 04 | 95.6 | |
| GSM30808 | FC 05 | 100.6 | |
| GSM30809 | IFC 01 | 81.9 | |
| GSM30810 | IFC 02 | 78.5 | |
| GSM30811 | IFC 03 | 121.7 | |
| GSM30812 | IFC 04 | 112.8 | |
| GSM30813 | IFC 05 | 92.9 | |
| GSM99800 | isoflurane 01 | male, 275-300g | 96.1 |
| GSM99801 | isoflurane 02 | male, 275-300g | 129.1 |
| GSM99802 | isoflurane 03 | male, 275-300g | 113.3 |
| GSM99803 | isoflurane 04 | male, 275-300g | 91.7 |
| GSM99804 | isoflurane 05 | male, 275-300g | 93.8 |
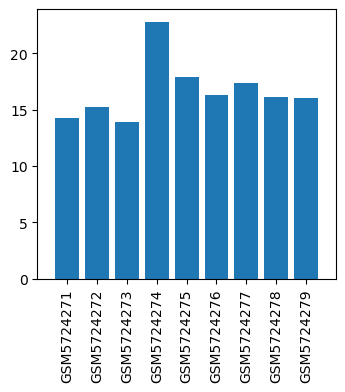
| ID | GSE190472 |
| Title | Study on VEGF regulation mechanism of acupuncture improving endometrial receptivity in IVF-ET |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5724271 | model 21 | Sex: female tissue: Uterine tissue condition: model | 14.2815 |
| GSM5724272 | model 22 | Sex: female tissue: Uterine tissue condition: model | 15.2629 |
| GSM5724273 | model 23 | Sex: female tissue: Uterine tissue condition: model | 13.9152 |
| GSM5724274 | treatment31 | Sex: female tissue: Uterine tissue condition: treatment | 22.7903 |
| GSM5724275 | treatment32 | Sex: female tissue: Uterine tissue condition: treatment | 17.9364 |
| GSM5724276 | treatment33 | Sex: female tissue: Uterine tissue condition: treatment | 16.2731 |
| GSM5724277 | control 41 | Sex: female tissue: Uterine tissue condition: control | 17.3631 |
| GSM5724278 | control 42 | Sex: female tissue: Uterine tissue condition: control | 16.1034 |
| GSM5724279 | control 43 | Sex: female tissue: Uterine tissue condition: control | 16.0768 |
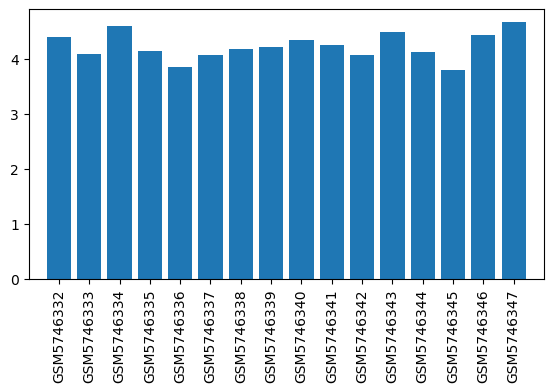
| ID | GSE192399 |
| Title | Gene expression of brain regions activated by sevoflurane and propofol |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5746332 | LHb, Sevoflurane, 60mins | brain region: the lateral habenula (LHb) treatment: Sevoflurane | 4.39875 |
| GSM5746333 | LHb, O2, 60mins | brain region: the lateral habenula (LHb) treatment: O2 | 4.07708 |
| GSM5746334 | MHb, Sevoflurane, 60mins | brain region: the medial habenula (MHb) treatment: Sevoflurane | 4.59536 |
| GSM5746335 | MHb, O2, 60mins | brain region: the medial habenula (MHb) treatment: O2 | 4.14179 |
| GSM5746336 | Sol, Sevoflurane, 60mins | brain region: the solitary nucleus (Sol) treatment: Sevoflurane | 3.85123 |
| GSM5746337 | Sol, O2, 60mins | brain region: the solitary nucleus (Sol) treatment: O2 | 4.06102 |
| GSM5746338 | Mve, Sevoflurane, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Sevoflurane | 4.16838 |
| GSM5746339 | Mve, O2, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: O2 | 4.20325 |
| GSM5746340 | LHb, Propofol, 60mins | brain region: the lateral habenula (LHb) treatment: Propofol | 4.33654 |
| GSM5746341 | LHb, Intralipos, 60mins | brain region: the lateral habenula (LHb) treatment: Intralipos | 4.23912 |
| GSM5746342 | MHb, Propofol, 60mins | brain region: the medial habenula (MHb) treatment: Propofol | 4.06027 |
| GSM5746343 | MHb, Intralipos, 60mins | brain region: the medial habenula (MHb) treatment: Intralipos | 4.48517 |
| GSM5746344 | Sol, Propofol, 60mins | brain region: the solitary nucleus (Sol) treatment: Propofol | 4.12257 |
| GSM5746345 | Sol, Intralipos, 60mins | brain region: the solitary nucleus (Sol) treatment: Intralipos | 3.78783 |
| GSM5746346 | MVe, Propofol, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Propofol | 4.43397 |
| GSM5746347 | MVe, Intralipos, 60mins | brain region: the medial vestibular nucleus (MVe) treatment: Intralipos | 4.66659 |
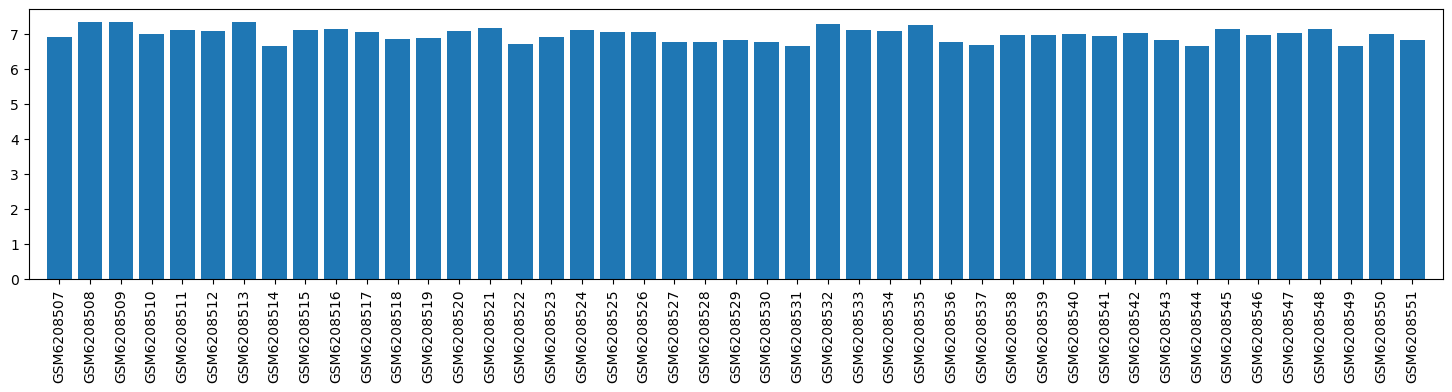
| ID | GSE205202 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [rat] |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208507 | Rat_Baseline_0h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 6.891523 |
| GSM6208508 | Rat_Baseline_0h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 7.316336 |
| GSM6208509 | Rat_Baseline_0h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 7.330974 |
| GSM6208510 | Rat_Baseline_0h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 6.983605 |
| GSM6208511 | Rat_Baseline_0h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Untreated hours after treatment: 0 | 7.097271 |
| GSM6208512 | Rat_Vehicle_03h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.059467 |
| GSM6208513 | Rat_Vehicle_03h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.31948 |
| GSM6208514 | Rat_Vehicle_03h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 6.653553 |
| GSM6208515 | Rat_Vehicle_03h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.107881 |
| GSM6208516 | Rat_Vehicle_03h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 7.137746 |
| GSM6208517 | Rat_Vehicle_06h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.051125 |
| GSM6208518 | Rat_Vehicle_06h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 6.85042 |
| GSM6208519 | Rat_Vehicle_06h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 6.876999 |
| GSM6208520 | Rat_Vehicle_06h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.065683 |
| GSM6208521 | Rat_Vehicle_06h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 7.152061 |
| GSM6208522 | Rat_Vehicle_09h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 6.700373 |
| GSM6208523 | Rat_Vehicle_09h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 6.899775 |
| GSM6208524 | Rat_Vehicle_09h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.092198 |
| GSM6208525 | Rat_Vehicle_09h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.042413 |
| GSM6208526 | Rat_Vehicle_09h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 7.055605 |
| GSM6208527 | Rat_Vehicle_24h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 6.745739 |
| GSM6208528 | Rat_Vehicle_24h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 6.759711 |
| GSM6208529 | Rat_Vehicle_24h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 6.830468 |
| GSM6208530 | Rat_Vehicle_24h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 6.758411 |
| GSM6208531 | Rat_Vehicle_24h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 6.635457 |
| GSM6208532 | Rat_APAP_03h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.278534 |
| GSM6208533 | Rat_APAP_03h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.097861 |
| GSM6208534 | Rat_APAP_03h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.072164 |
| GSM6208535 | Rat_APAP_03h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 7.237181 |
| GSM6208536 | Rat_APAP_03h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 6.770045 |
| GSM6208537 | Rat_APAP_06h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 6.682379 |
| GSM6208538 | Rat_APAP_06h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 6.955787 |
| GSM6208539 | Rat_APAP_06h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 6.948583 |
| GSM6208540 | Rat_APAP_06h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 6.979591 |
| GSM6208541 | Rat_APAP_06h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 6.929037 |
| GSM6208542 | Rat_APAP_09h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.006586 |
| GSM6208543 | Rat_APAP_09h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 6.826284 |
| GSM6208544 | Rat_APAP_09h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 6.6354 |
| GSM6208545 | Rat_APAP_09h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 7.142825 |
| GSM6208546 | Rat_APAP_09h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 6.964997 |
| GSM6208547 | Rat_APAP_24h_1 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.022771 |
| GSM6208548 | Rat_APAP_24h_2 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 7.136844 |
| GSM6208549 | Rat_APAP_24h_3 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 6.65166 |
| GSM6208550 | Rat_APAP_24h_4 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 6.983605 |
| GSM6208551 | Rat_APAP_24h_5 | tissue: liver strain: Sprague-Dawley Sex: male treatment: 1000 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 6.803828 |
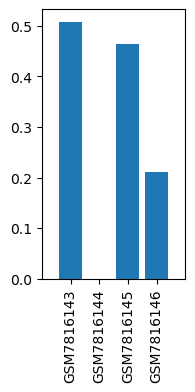
| ID | GSE244436 |
| Title | Similarity and dissimilarity in alteration of gene expression profile associated with inhalational anesthesia between sevoflurane and desflurane |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM7816143 | Liver, Control for SevoAnes | tissue: liver genotype: wild type treatment: Control for SevoAnes | 0.507718 |
| GSM7816144 | Liver, Anesthetized with sevoflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with sevoflurane for 6 h | 0.0 |
| GSM7816145 | Liver, Control for DesAnes | tissue: liver genotype: wild type treatment: Control for DesAnes | 0.463431 |
| GSM7816146 | Liver, Anesthetized with desflurane for 6 h | tissue: liver genotype: wild type treatment: Anesthetized with desflurane for 6 h | 0.211187 |
| ID | GSE2982 |
| Title | Effect of inhaled anesthetic to cultured cortical neurons |
| Organism | Rattus norvegicus |
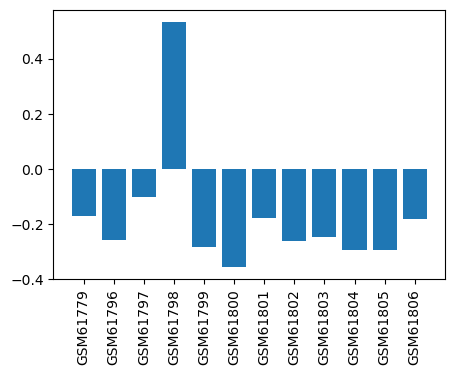
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM61779 | Cell Culture Exposure (control1) | Cy3 label;Cy5; | -0.169 |
| GSM61796 | Cell Culture Exposure (control2) | Cy3; Cy5 | -0.257 |
| GSM61797 | Cell Culture Exposure (control3) | Cy3; Cy5 | -0.103 |
| GSM61798 | Cell Culture Exposure (1MAC Halo1) | Cy3; Cy5 | 0.533 |
| GSM61799 | Cell Culture Exposure (1MAC Halo2) | Cy3; Cy5 | -0.282 |
| GSM61800 | Cell Culture Exposure (1MAC Halo3) | Cy3; Cy5 | -0.356 |
| GSM61801 | Cell Culture Exposure (3MAC Halo1) | Cy3; Cy5 | -0.177 |
| GSM61802 | Cell Culture Exposure (3MAC Halo2) | Cy3; Cy5 | -0.262 |
| GSM61803 | Cell Culture Exposure (3MAC Halo3) | Cy3; Cy5 | -0.245 |
| GSM61804 | Cell Culture Exposure (3MAC ISO1) | Cy3; Cy5 | -0.294 |
| GSM61805 | Cell Culture Exposure (3MAC ISO2) | Cy3; Cy5 | -0.295 |
| GSM61806 | Cell Culture Exposure (3MAC ISO3) | Cy3; Cy5 | -0.183 |
| ID | GSE359 |
| Title | Isoflurane Single Exposures |
| Organism | Rattus norvegicus |
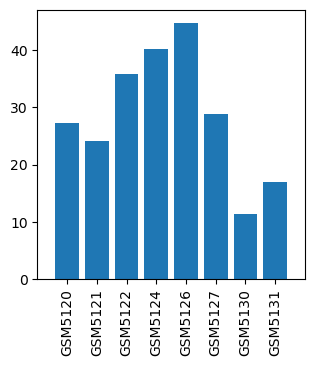
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5120 | Isoflurane single exposure control-1 | 27.2 | |
| GSM5121 | Isoflurane single exposure control-2 | 24.150000000000002 | |
| GSM5122 | Isoflurane single exposure control-3 | 35.9 | |
| GSM5124 | Isoflurane single exposure 1 hr-1 | 40.2 | |
| GSM5126 | Isoflurane single exposure 1 hr-2 | 44.8 | |
| GSM5127 | Isoflurane single exposure 2 hr-1 | 28.9 | |
| GSM5130 | Isoflurane single exposure 2 hr-2 | 11.45 | |
| GSM5131 | Isoflurane single exposure 2 hr-3 | 17.0 |
| ID | GSE48025 |
| Title | Affymetrix Chip Data of the Transcriptome of the Rheumatoid Arthritis Rat Model Treated with Acupuncture (Affymetrix, mRNA, batch1) |
| Organism | Rattus norvegicus |
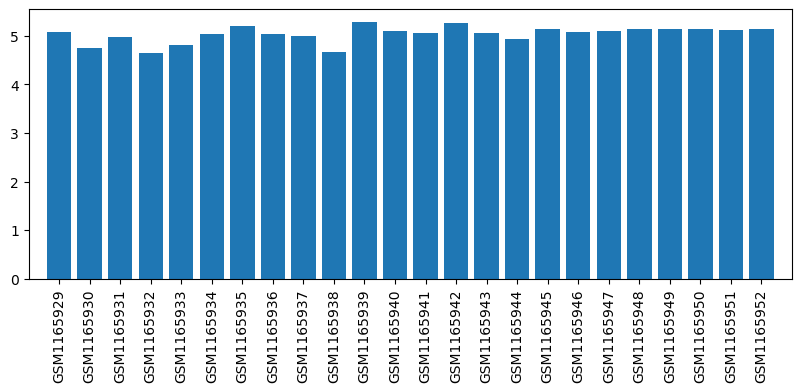
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1165929 | RAT_NORA_T0_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats before therapy | 5.083167908 |
| GSM1165930 | RAT_NORA_T0_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats before therapy | 4.748994804 |
| GSM1165931 | RAT_RA_T0_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats before therapy | 4.980205448 |
| GSM1165932 | RAT_RA_T0_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats before therapy | 4.642241839 |
| GSM1165933 | RAT_NORA_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 1 hour after therapy | 4.818003367 |
| GSM1165934 | RAT_NORA_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 1 hour after therapy | 5.028645759 |
| GSM1165935 | RAT_AP_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour acupuncture therapy | 5.206042332 |
| GSM1165936 | RAT_AP_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour acupuncture therapy | 5.037943766 |
| GSM1165937 | RAT_MTX_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour MTX therapy | 5.000758442 |
| GSM1165938 | RAT_MTX_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour MTX therapy | 4.670032483 |
| GSM1165939 | RAT_ANE_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour ANE therapy | 5.285504733 |
| GSM1165940 | RAT_ANE_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour ANE therapy | 5.089789062 |
| GSM1165941 | RAT_PLA_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour placebo therapy | 5.052449652 |
| GSM1165942 | RAT_PLA_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour placebo therapy | 5.270209587 |
| GSM1165943 | RAT_NORA_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 34 days after therapy | 5.066787097 |
| GSM1165944 | RAT_NORA_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 34 days after therapy | 4.926436266 |
| GSM1165945 | RAT_AP_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days acupuncture therapy | 5.144104846 |
| GSM1165946 | RAT_AP_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days acupuncture therapy | 5.082792657 |
| GSM1165947 | RAT_MTX_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days MTX therapy | 5.091195187 |
| GSM1165948 | RAT_MTX_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days MTX therapy | 5.145808764 |
| GSM1165949 | RAT_ANE_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days ANE therapy | 5.143255405 |
| GSM1165950 | RAT_ANE_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days ANE therapy | 5.151107657 |
| GSM1165951 | RAT_PLA_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days placebo therapy | 5.128288144 |
| GSM1165952 | RAT_PLA_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days placebo therapy | 5.143537823 |
| ID | GSE53897 |
| Title | Genome-wide analysis of dorsal root ganglion (DRG) gene expression after morphine or oxycodone treatment |
| Organism | Rattus norvegicus |
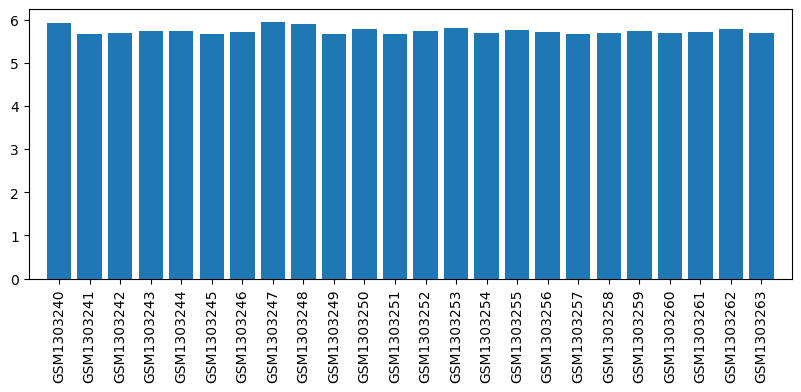
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1303240 | DRG RVM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.919677 |
| GSM1303241 | DRG RVM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.666903 |
| GSM1303242 | DRG RVM3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.690935 |
| GSM1303243 | DRG RVM7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.740078 |
| GSM1303244 | DRG RVOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.738307 |
| GSM1303245 | DRG RVOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.670802 |
| GSM1303246 | DRG RVOx5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.71441 |
| GSM1303247 | DRG RVOx7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.943155 |
| GSM1303248 | DRG RVS4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.898044 |
| GSM1303249 | DRG RVS5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.667056 |
| GSM1303250 | DRG RVS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.784102 |
| GSM1303251 | DRG RVS7 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.663336 |
| GSM1303252 | DRG RSM1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.735748 |
| GSM1303253 | DRG RSM2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.799071 |
| GSM1303254 | DRG RSM4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.690575 |
| GSM1303255 | DRG RSM5 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.768971 |
| GSM1303256 | DRG RSOx1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.720709 |
| GSM1303257 | DRG RSOx2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.654727 |
| GSM1303258 | DRG RSOx3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.688556 |
| GSM1303259 | DRG RSOx4 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.724449 |
| GSM1303260 | DRG RSS1 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.685169 |
| GSM1303261 | DRG RSS2 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.714156 |
| GSM1303262 | DRG RSS3 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.783576 |
| GSM1303263 | DRG RSS6 | strain: sprague-dawley age: 3 months tissue: dorsal root ganglion | 5.691554 |
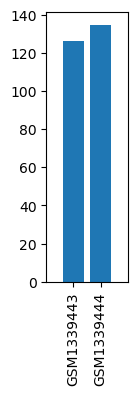
| ID | GSE55573 |
| Title | Using DNA microarray to screen intracellular signal pathways of hypoglycemic activity by electroacupuncture of Zusanli (ST36) acupoints in streptozotocin-induced rats with diabetes |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1339443 | Mock skeletal muscle | strain: Wistar age: 8-10 week-old gender: Male | 126.5003 |
| GSM1339444 | EA (electroacupuncture) skeletal muscle | strain: Wistar age: 8-10 week-old gender: Male | 134.6747 |
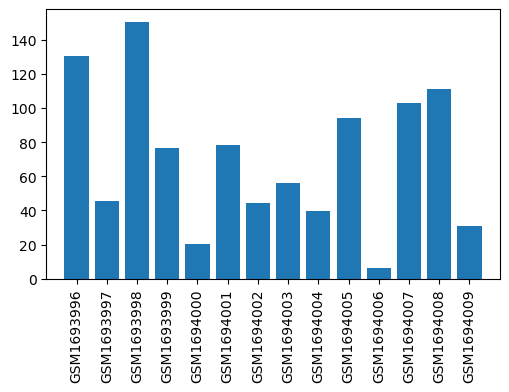
| ID | GSE58456 |
| Title | Affymetrix Chip Data of the Transcriptome of the Rheumatoid Arthritis Rat Model Treated with Acupuncture (Affymetrix, mRNA, batch 2) |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1693996 | Peripheral blood mononuclear cells from NORA before therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 130.7834581 |
| GSM1693997 | Peripheral blood mononuclear cells from NORA before therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 45.24435265 |
| GSM1693998 | Peripheral blood mononuclear cells from NORA before therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 150.4452888 |
| GSM1693999 | Peripheral blood mononuclear cells from RA before therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats before therapy | 76.5601194 |
| GSM1694000 | Peripheral blood mononuclear cells from RA before therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats before therapy | 20.08929709 |
| GSM1694001 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 78.52717001 |
| GSM1694002 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 44.32551353 |
| GSM1694003 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 55.89985048 |
| GSM1694004 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 39.73506158 |
| GSM1694005 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 94.20145667 |
| GSM1694006 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 5.98615891 |
| GSM1694007 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 103.1492002 |
| GSM1694008 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 110.9625494 |
| GSM1694009 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 30.82872697 |
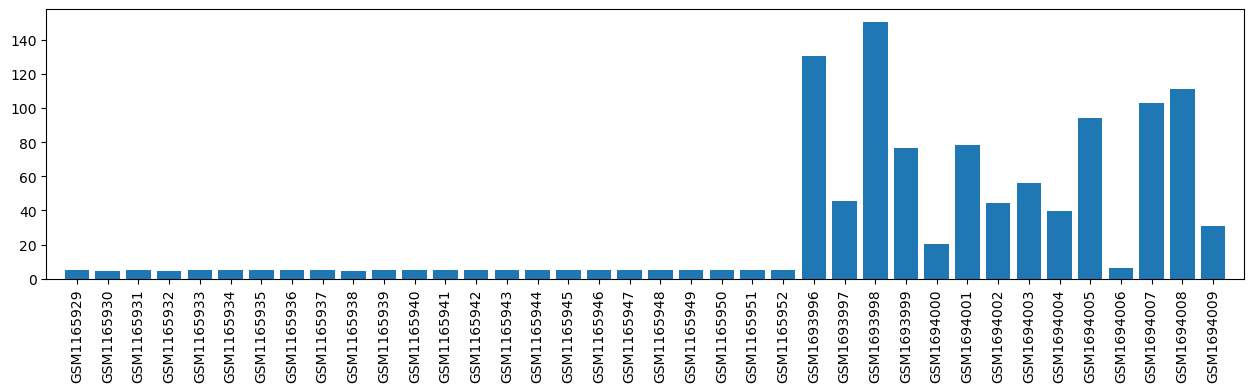
| ID | GSE58460 |
| Title | Rheumatoid Arthritis Rat Model Treated with Acupuncture |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1165929 | RAT_NORA_T0_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats before therapy | 5.083167908 |
| GSM1165930 | RAT_NORA_T0_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats before therapy | 4.748994804 |
| GSM1165931 | RAT_RA_T0_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats before therapy | 4.980205448 |
| GSM1165932 | RAT_RA_T0_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats before therapy | 4.642241839 |
| GSM1165933 | RAT_NORA_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 1 hour after therapy | 4.818003367 |
| GSM1165934 | RAT_NORA_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 1 hour after therapy | 5.028645759 |
| GSM1165935 | RAT_AP_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour acupuncture therapy | 5.206042332 |
| GSM1165936 | RAT_AP_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour acupuncture therapy | 5.037943766 |
| GSM1165937 | RAT_MTX_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour MTX therapy | 5.000758442 |
| GSM1165938 | RAT_MTX_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour MTX therapy | 4.670032483 |
| GSM1165939 | RAT_ANE_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour ANE therapy | 5.285504733 |
| GSM1165940 | RAT_ANE_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour ANE therapy | 5.089789062 |
| GSM1165941 | RAT_PLA_1h_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour placebo therapy | 5.052449652 |
| GSM1165942 | RAT_PLA_1h_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 1 hour placebo therapy | 5.270209587 |
| GSM1165943 | RAT_NORA_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 34 days after therapy | 5.066787097 |
| GSM1165944 | RAT_NORA_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: healthy rats at 34 days after therapy | 4.926436266 |
| GSM1165945 | RAT_AP_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days acupuncture therapy | 5.144104846 |
| GSM1165946 | RAT_AP_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days acupuncture therapy | 5.082792657 |
| GSM1165947 | RAT_MTX_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days MTX therapy | 5.091195187 |
| GSM1165948 | RAT_MTX_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days MTX therapy | 5.145808764 |
| GSM1165949 | RAT_ANE_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days ANE therapy | 5.143255405 |
| GSM1165950 | RAT_ANE_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days ANE therapy | 5.151107657 |
| GSM1165951 | RAT_PLA_34d_1 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days placebo therapy | 5.128288144 |
| GSM1165952 | RAT_PLA_34d_2 | gender: Female tissue: RAT Peripheral blood mononuclear cells strain: Wistar rat subject group: RA rats after 34 days placebo therapy | 5.143537823 |
| GSM1412003 | RAT_NORA_T0_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412004 | RAT_NORA_T0_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412005 | RAT_RA_T0_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412006 | RAT_RA_T0_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412007 | RAT_NORA_1h_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412008 | RAT_NORA_1h_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412009 | RAT_AP_1h_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412010 | RAT_AP_1h_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412011 | RAT_MTX_1h_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412012 | RAT_MTX_1h_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412013 | RAT_ANE_1h_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412014 | RAT_ANE_1h_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412015 | RAT_PLA_1h_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412016 | RAT_PLA_1h_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412017 | RAT_NORA_34d_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412018 | RAT_NORA_34d_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412019 | RAT_AP_34d_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412020 | RAT_AP_34d_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412021 | RAT_MTX_34d_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412022 | RAT_MTX_34d_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412023 | RAT_ANE_34d_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412024 | RAT_ANE_34d_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412025 | RAT_PLA_34d_1 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412026 | RAT_PLA_34d_2 | strain: Wistar gender: Female tissue: RAT Peripheral blood mononuclear cells | |
| GSM1412027 | RAT_RA_T0_1 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412028 | RAT_RA_T0_2 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412029 | RAT_AP_1h_1 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412030 | RAT_AP_1h_2 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412031 | RAT_PLA_1h_1 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412032 | RAT_PLA_1h_2 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412033 | RAT_AP_34d_1 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412034 | RAT_AP_34d_2 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412035 | RAT_PLA_34d_1 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1412036 | RAT_PLA_34d_2 | strain: Wistar gender: Female tissue: RAT Sub-epidermidis muscle tissue | |
| GSM1693996 | Peripheral blood mononuclear cells from NORA before therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therap | 130.7834581 |
| GSM1693997 | Peripheral blood mononuclear cells from NORA before therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 45.24435265 |
| GSM1693998 | Peripheral blood mononuclear cells from NORA before therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats before therapy | 150.4452888 |
| GSM1693999 | Peripheral blood mononuclear cells from RA before therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats before therapy | 76.5601194 |
| GSM1694000 | Peripheral blood mononuclear cells from RA before therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats before therapy | 20.08929709 |
| GSM1694001 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 78.52717001 |
| GSM1694002 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therap | 44.32551353 |
| GSM1694003 | Peripheral blood mononuclear cells from HAP after 34 days acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: healthy rats after 34 days acupuncture therapy | 55.89985048 |
| GSM1694004 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 39.73506158 |
| GSM1694005 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 94.20145667 |
| GSM1694006 | Peripheral blood mononuclear cells from AP after 34 days acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days acupuncture therapy | 5.98615891 |
| GSM1694007 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 1 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 103.1492002 |
| GSM1694008 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 2 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 110.9625494 |
| GSM1694009 | Peripheral blood mononuclear cells from MTXAP after 34 days MTX&acupuncture therapy, replicate 3 | strain/background: Wistar gender: female cell type: peripheral blood mononuclear cells subject group: rheumatoid arthritis rats after 34 days methotrexate & acupuncture therapy | 30.82872697 |
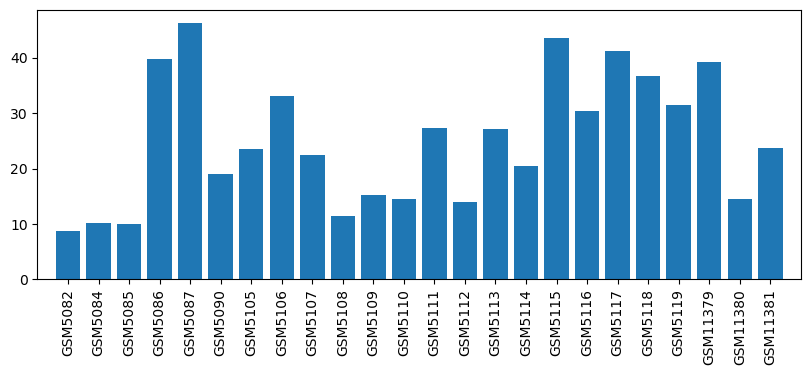
| ID | GSE752 |
| Title | Halothane/Isoflurane Repetitive Exposures |
| Organism | Rattus norvegicus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM5082 | Control 1 | 8.8 | |
| GSM5084 | Control 2 | 10.2 | |
| GSM5085 | Control 3 | 9.95 | |
| GSM5086 | Control 4 | 39.849999999999994 | |
| GSM5087 | Control 5 | 46.349999999999994 | |
| GSM5090 | Control 6 | 18.950000000000003 | |
| GSM5105 | Control 7 | 23.549999999999997 | |
| GSM5106 | Control 8 | 33.2 | |
| GSM5107 | Control 9 | 22.450000000000003 | |
| GSM5108 | Halothane 10 exposure-1 | 11.45 | |
| GSM5109 | Halothane 10 exposures-2 | 15.25 | |
| GSM5110 | Halothane 10 exposures-3 | 14.5 | |
| GSM5111 | Halothane 5 exposures-1 | 27.3 | |
| GSM5112 | Halothane 5 exposures-2 | 14.0 | |
| GSM5113 | Halothane 5 exposures-3 | 27.25 | |
| GSM5114 | Isoflurane 10 exposures-1 | 20.4 | |
| GSM5115 | Isoflurane 10 exposures-2 | 43.599999999999994 | |
| GSM5116 | Isoflurane 10 exposures-3 | 30.5 | |
| GSM5117 | Isoflurane 5 exposures-1 | 41.3 | |
| GSM5118 | Isoflurane 5 exposures-2 | 36.8 | |
| GSM5119 | Isoflurane 5 exposures-3 | 31.45 | |
| GSM11379 | Control 10 | 39.25 | |
| GSM11380 | Control 11 | 14.5 | |
| GSM11381 | Control 12 | 23.8 |
| ID | GSE8911 |
| Title | Skeletal muscle after fine needle stimulation |
| Organism | Rattus norvegicus |
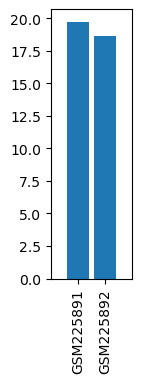
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM225891 | Skeletal muscle sample from fine needle-stimulated hindlimb | Strain: Lewis, Gender: Male, Body Weight: 275g, Tissue: Skeletal muscle sample from fine nnedle-stimulated hindlimb | 19.7109 |
| GSM225892 | Skeletal muscle sample from control hindlimb | Strain: Lewis, Gender: Male, Body weight: 275g, Tissue: skeletal muscle sample from non fine needle-stimulated hindlimb | 18.6075 |

