Gene Information
|
Gene Name
|
IL7 |
|
Gene ID
|
3574
|
|
Gene Full Name
|
interleukin 7 |
|
Gene Alias
|
IL-7|IMD130 |
|
Transcripts
|
ENSG00000104432
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Secreted to blood; 
|
|
Membrane Info
|
Cancer-related genes, Disease related genes, Human disease related genes, Plasma proteins, Predicted secreted proteins |
|
Uniport_ID
|
P13232
|
|
HGNC ID
|
HGNC:6023
|
|
OMIM ID
|
146660 |
|
Summary
|
The protein encoded by this gene is a cytokine important for B and T cell development. This cytokine and the hepatocyte growth factor (HGF) form a heterodimer that functions as a pre-pro-B cell growth-stimulating factor. IL7 is found to be a cofactor for V(D)J rearrangement of the T cell receptor beta (TCRB) during early T cell development. This cytokine can be produced locally by intestinal epithelial and epithelial goblet cells, and may serve as a regulatory factor for intestinal mucosal lymphocytes. IL7 plays an essential role in lymphoid cell survival, and in the maintenance of naive and memory T cells. Alternative splicing results in multiple transcript variants encoding distinct isoforms. Additional splice variants have been described but their presence in normal tissues has not been confirmed. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection can be a potent inducer of proinflammatory cytokines and chemokines which may defend against the infection, but may also mediate destructive lung injury. Elevated serum IL7 levels, together with several other circulating cytokines and chemokines, has been found to be associated with the severity of Coronavirus Disease 19 (COVID-19). [provided by RefSeq, Jul 2020] |
Target gene [IL7] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1041587 |
chr8 |
10 |
0 |
0 |
10 |
View |
| 1041588 |
chr8 |
2 |
9 |
0 |
11 |
View |
Target gene [IL7] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
GSE247322
|
scRNA-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
C GSE64877
|
Chip-seq |
2 |
Illumina Genome Analyzer (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
C GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
When the gene can detect a peak in the dataset, a peak plot will be displayed.
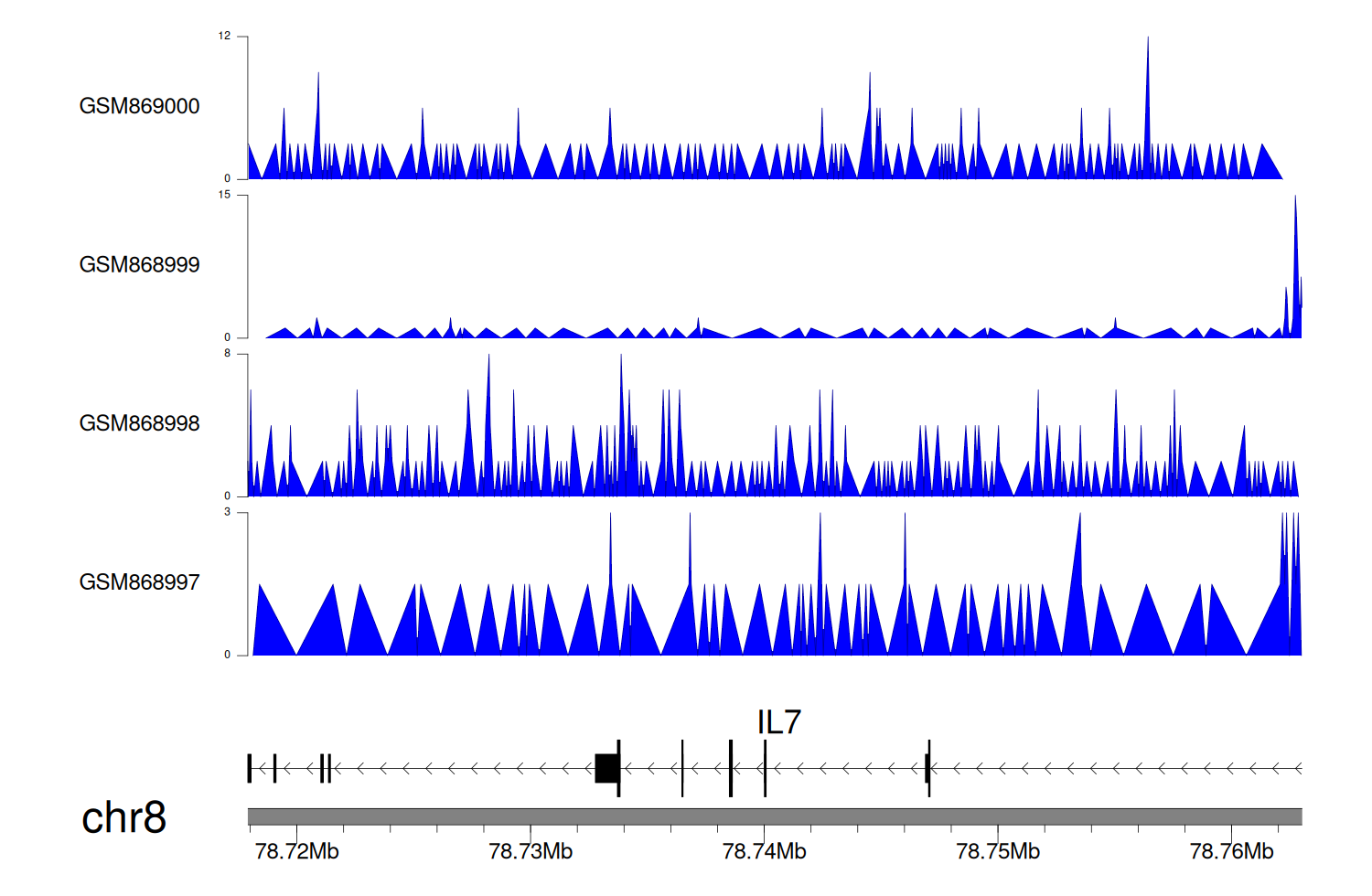
> Dataset: GSE35465 - IL7 peak across samples
|
Peak Plot
|
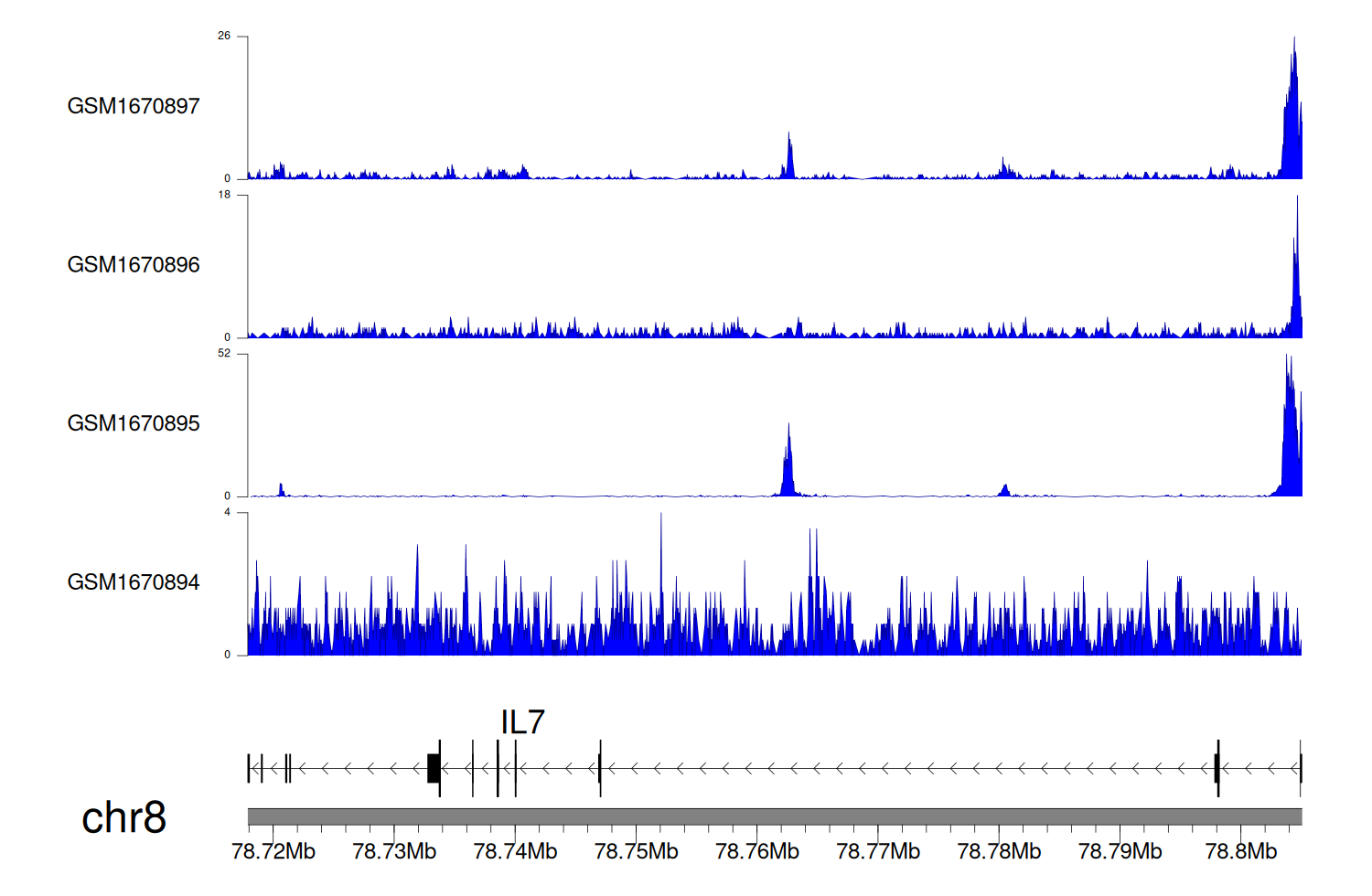
> Dataset: GSE68402 - IL7 peak across samples
|
Peak Plot
|
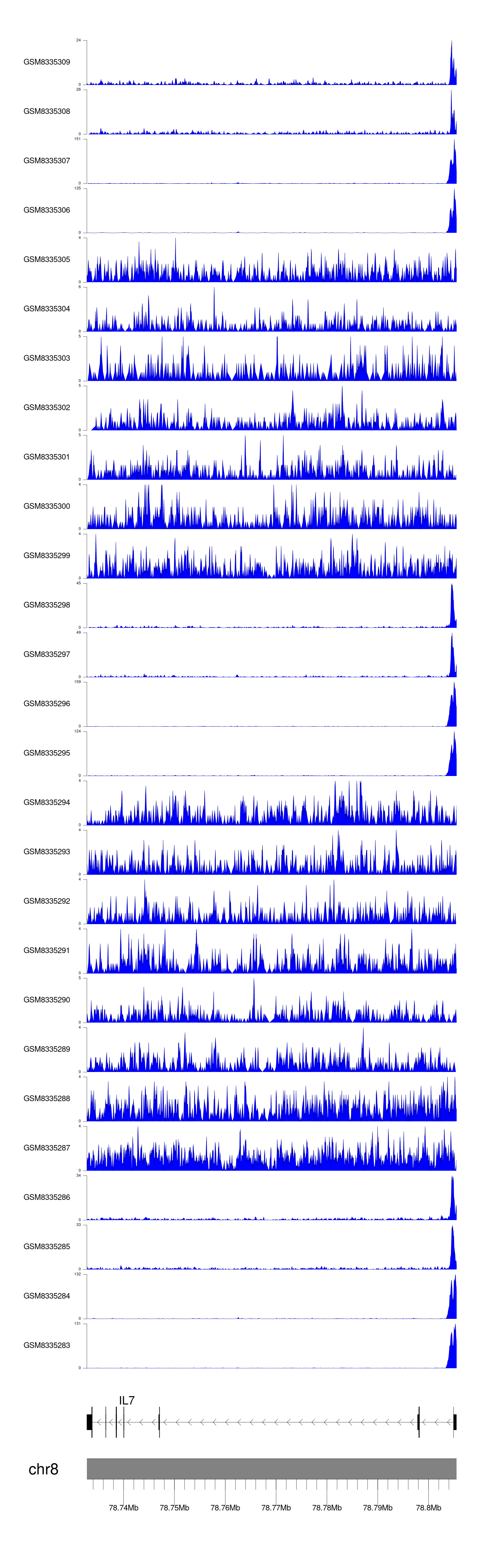
> Dataset: GSE270130 - IL7 peak across samples
|
Peak Plot
|
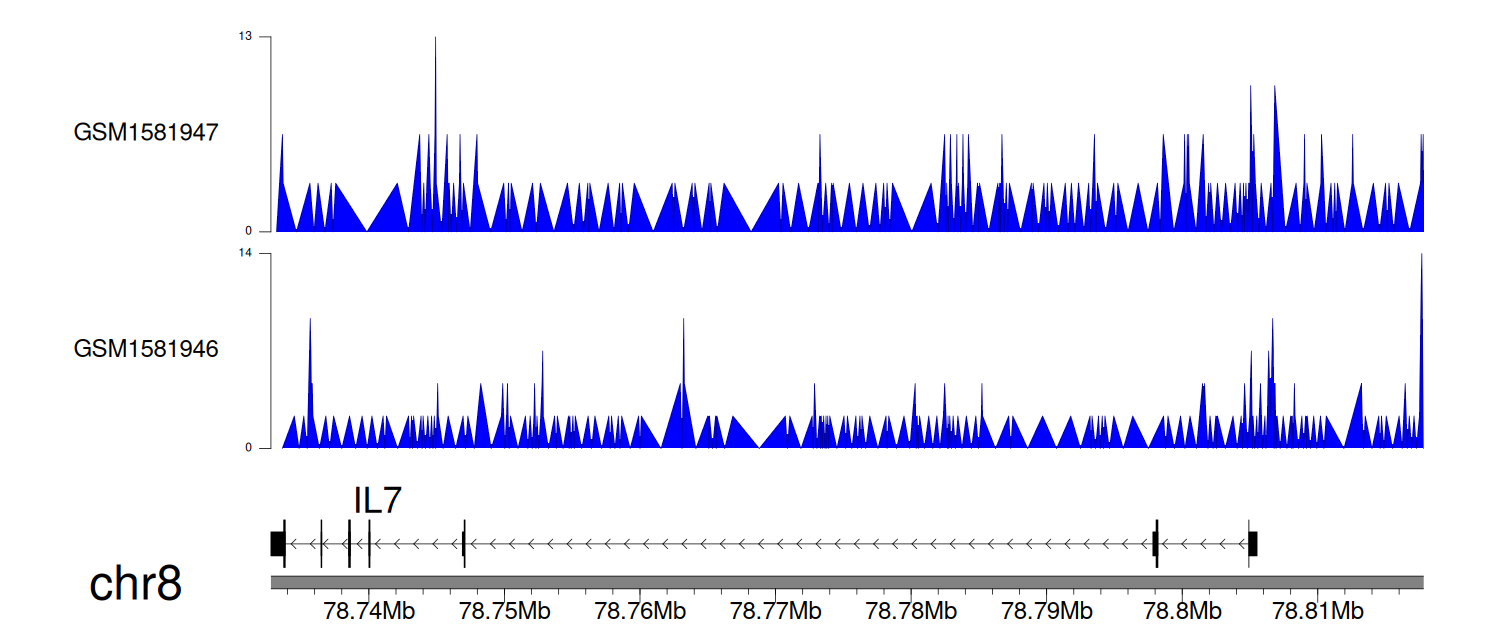
> Dataset: GSE64877 - IL7 peak across samples
|
Peak Plot
|
> Dataset: GSE100400 - IL7 peak across samples
|
Peak Plot
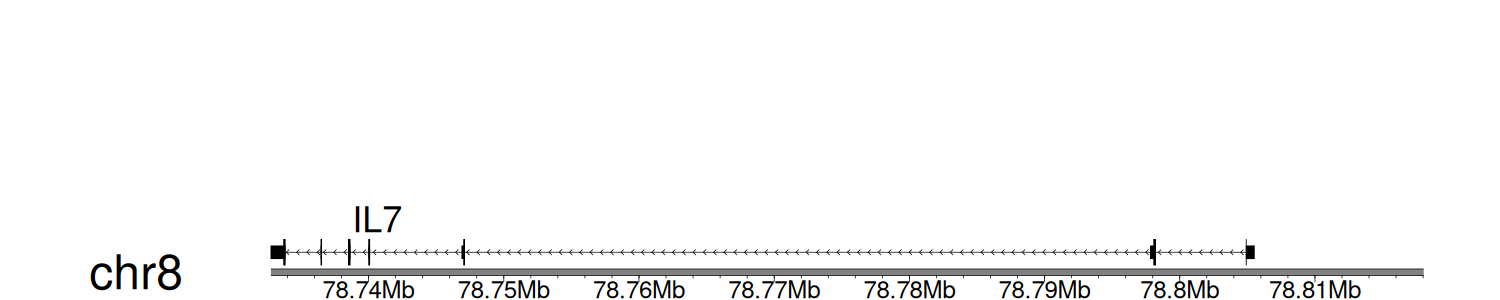
|
> Dataset: GSE131257 - IL7 peak across samples
|
Peak Plot
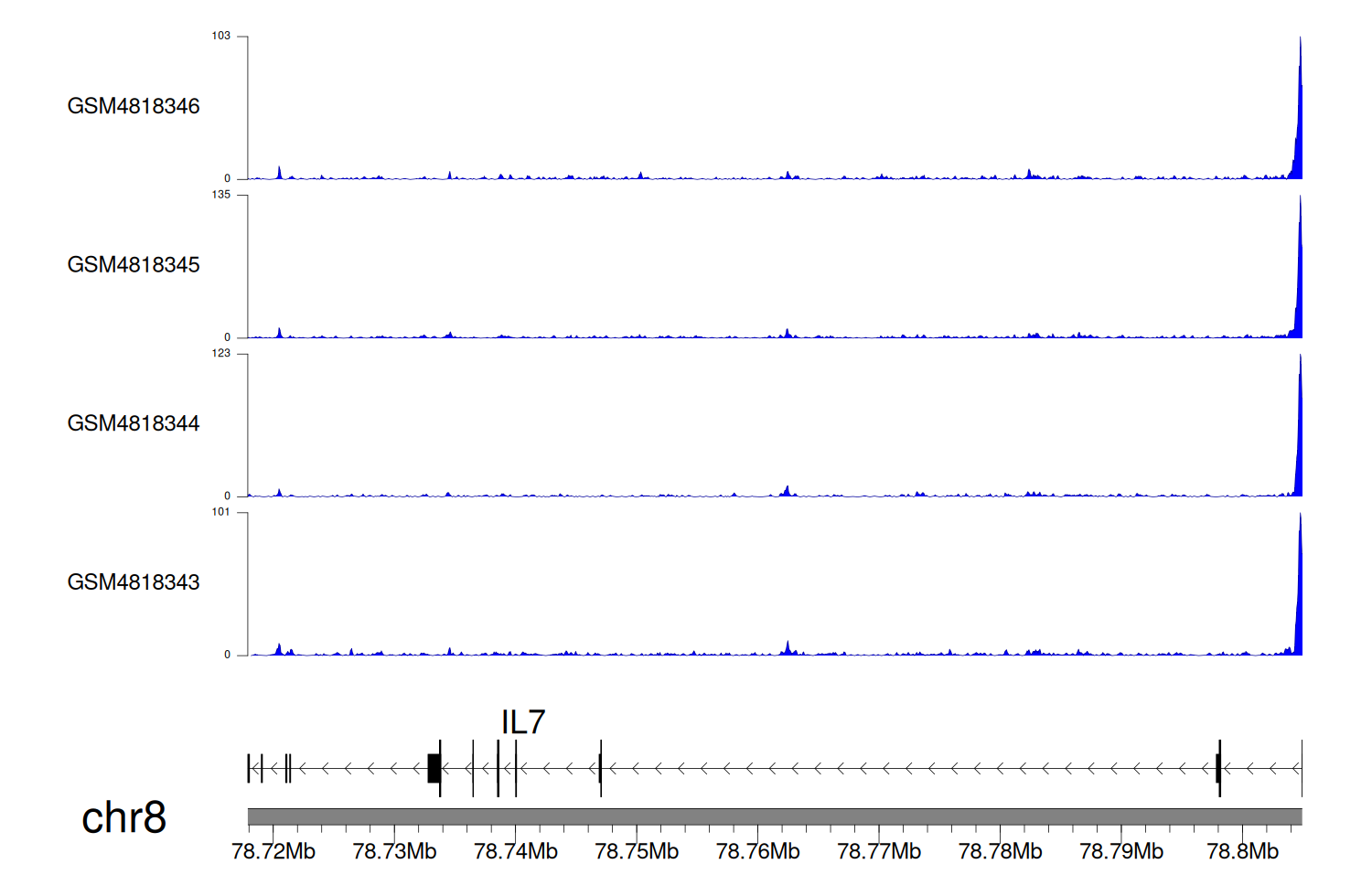
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information