Gene Information
|
Gene Name
|
NUP88 |
|
Gene ID
|
4927
|
|
Gene Full Name
|
nucleoporin 88 |
|
Gene Alias
|
FADS4 |
|
Transcripts
|
ENSG00000108559
|
|
Virus
|
HBV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Nucleoplasm; 
|
|
Membrane Info
|
Disease related genes, Human disease related genes, Metabolic proteins, Plasma proteins, Potential drug targets, Predicted intracellular proteins, Predicted membrane proteins, Transporters |
|
Uniport_ID
|
Q99567
|
|
HGNC ID
|
HGNC:8067
|
|
OMIM ID
|
602552 |
|
Summary
|
The nuclear pore complex is a massive structure that extends across the nuclear envelope, forming a gateway that regulates the flow of macromolecules between the nucleus and the cytoplasm. Nucleoporins, a family of 50 to 100 proteins, are the main components of the nuclear pore complex in eukaryotic cells. The protein encoded by this gene belongs to the nucleoporin family and is associated with the oncogenic nucleoporin CAN/Nup214 in a dynamic subcomplex. This protein is also overexpressed in a large number of malignant neoplasms and precancerous dysplasias. Alternative splicing results in multiple transcript variants encoding different isoforms. [provided by RefSeq, Mar 2016] |
Target gene [NUP88] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 1010056 |
chr17 |
13 |
0 |
0 |
13 |
View |
| 1042226 |
chr17 |
16 |
1 |
0 |
17 |
View |
Target gene [NUP88] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
GSE252863
|
scRNA-seq |
10 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE236281
|
RNA-seq |
12 |
Illumina MiSeq (Homo sapiens) |
|
C GSE35465
|
Chip-seq;RNA-seq |
6 |
Illumina HiSeq 2000 (Homo sapiens) |
|
C GSE68402
|
Chip-seq |
26 |
Illumina MiSeq (Homo sapiens);Illumina HiSeq 2500 (Homo sapiens) |
|
S GSE247322
|
scRNA-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
TCGA_LIHC_HBV
|
DNA methylation sequencing;RNA-seq |
97 |
TCGA |
|
C GSE270130
|
Chip-seq |
27 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE224901
|
RNA-seq |
21 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
C GSE100400
|
Chip-seq;RNA-seq;4C_cccDNA |
31 |
Illumina NextSeq 500 (Homo sapiens);Illumina NextSeq 500 (Mus musculus) |
|
GSE173897
|
RNA-seq |
95 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE262515
|
RNA-seq |
21 |
Illumina HiSeq 2500 (Homo sapiens);Illumina HiSeq 2500 (Mus musculus) |
|
GSE110345
|
RNA-seq |
4 |
Illumina HiSeq 2500 (Homo sapiens) |
|
C GSE131257
|
ATAC-seq;RNA-seq |
19 |
Illumina HiSeq 2500 (Homo sapiens) |
|
GSE94660
|
RNA-seq |
42 |
Illumina HiSeq 2500 (Homo sapiens) |
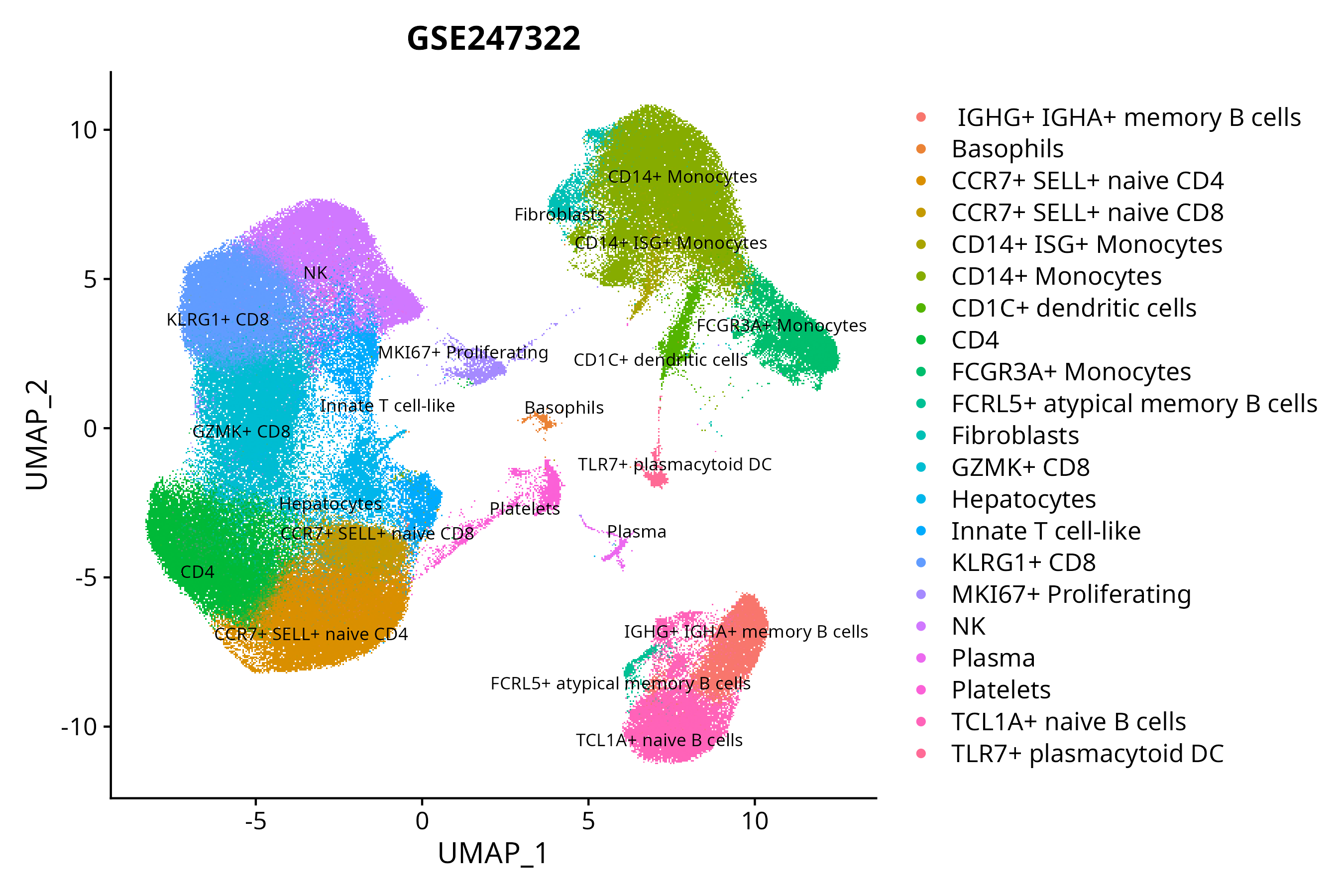
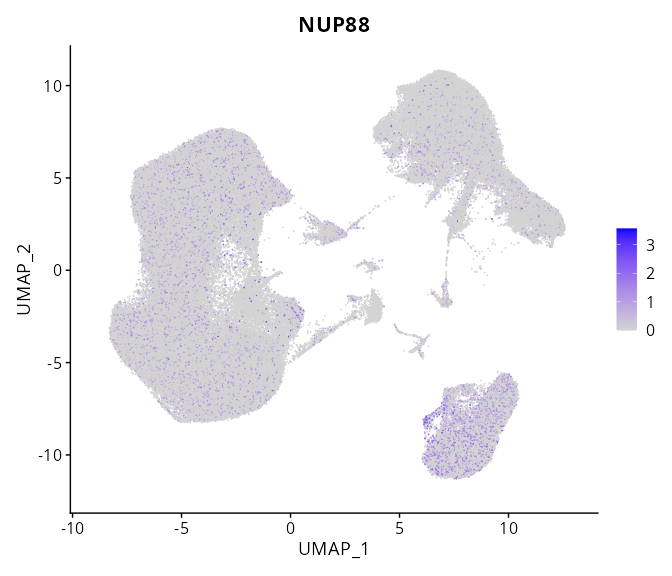
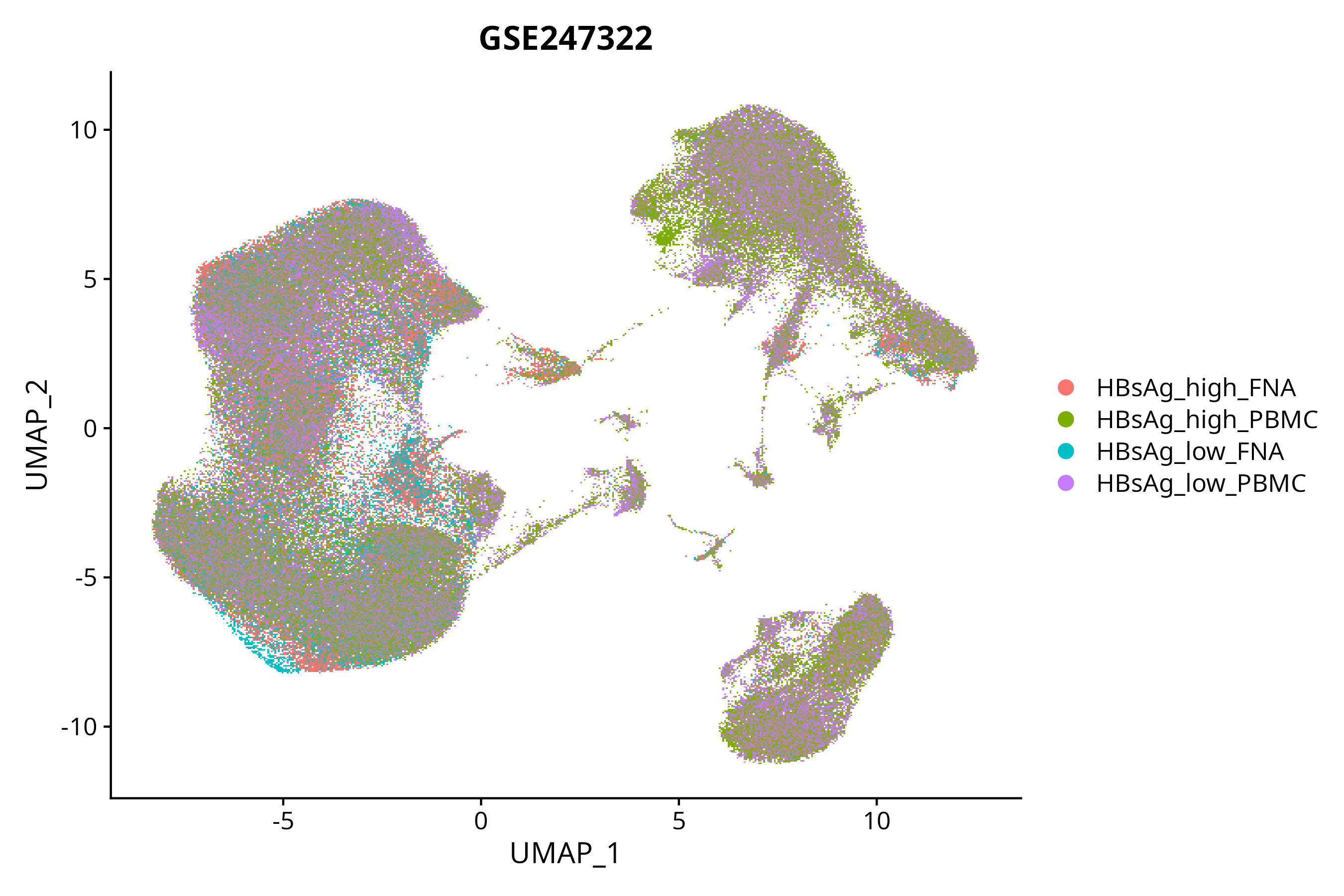
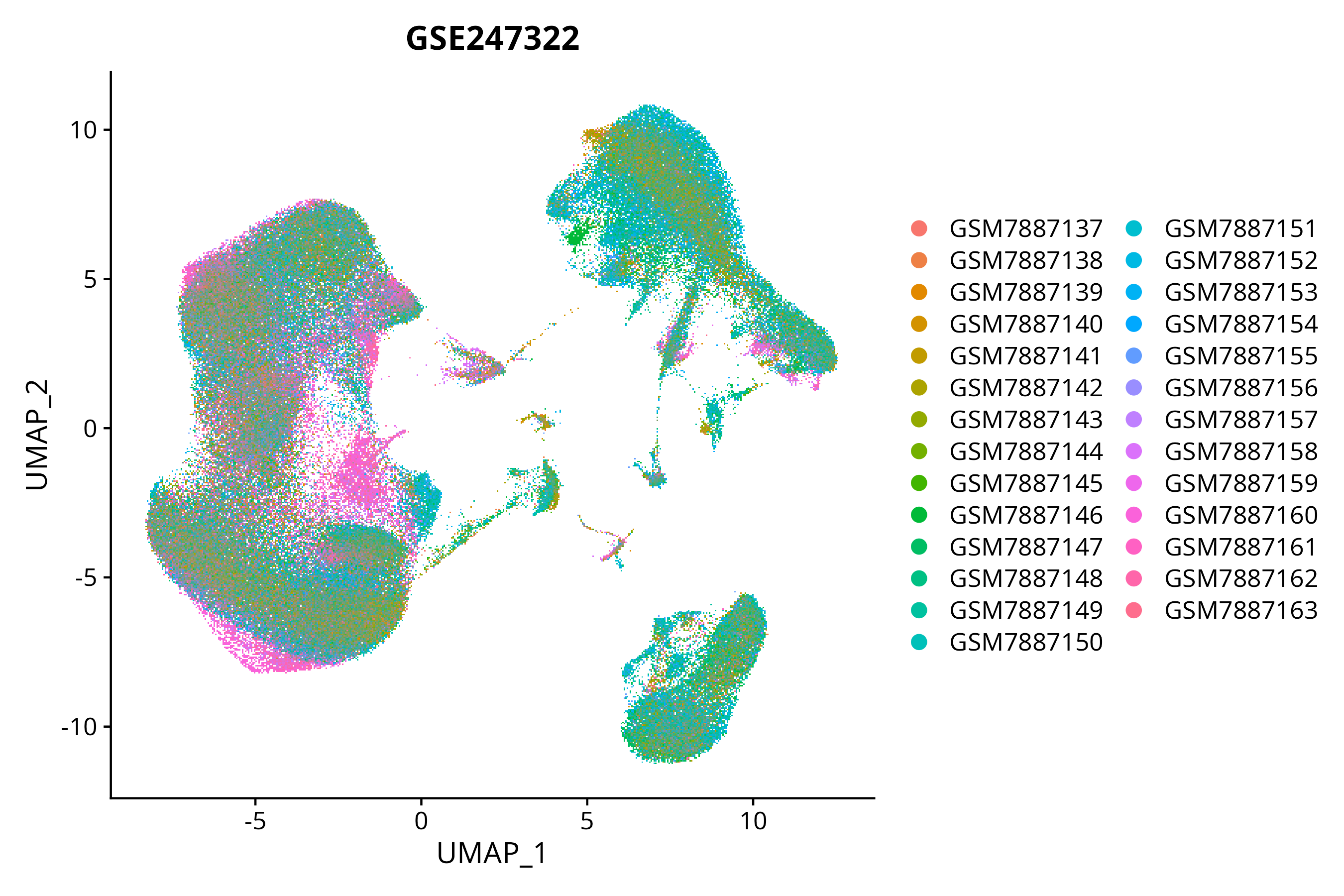
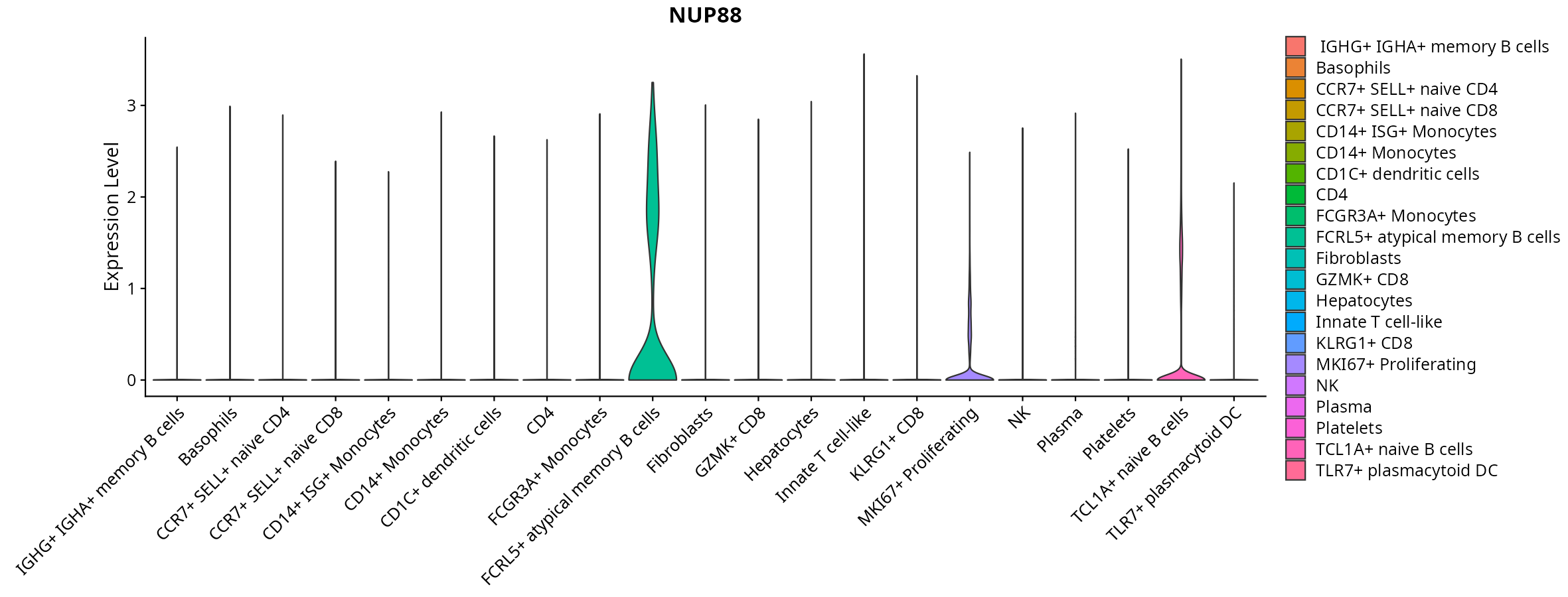
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE247322 - Gene expression in cell subsets
|

|
|
|
|
|
|
|
 HBV Target gene Detail Information
HBV Target gene Detail Information