Gene Information
|
Gene Name
|
SHH |
|
Gene ID
|
6469
|
|
Gene Full Name
|
sonic hedgehog signaling molecule |
|
Gene Alias
|
HHG1|HLP3|HPE3|MCOPCB5|SMMCI|ShhNC|TPT|TPTPS |
|
Transcripts
|
ENSG00000164690
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Intracellular and membrane; 
|
|
Membrane Info
|
Disease related genes, Human disease related genes, Predicted intracellular proteins |
|
Uniport_ID
|
Q15465
|
|
HGNC ID
|
HGNC:10848
|
|
OMIM ID
|
600725 |
|
Summary
|
This gene encodes a protein that is instrumental in patterning the early embryo. It has been implicated as the key inductive signal in patterning of the ventral neural tube, the anterior-posterior limb axis, and the ventral somites. Of three human proteins showing sequence and functional similarity to the sonic hedgehog protein of Drosophila, this protein is the most similar. The protein is made as a precursor that is autocatalytically cleaved; the N-terminal portion is soluble and contains the signalling activity while the C-terminal portion is involved in precursor processing. More importantly, the C-terminal product covalently attaches a cholesterol moiety to the N-terminal product, restricting the N-terminal product to the cell surface and preventing it from freely diffusing throughout the developing embryo. Defects in this protein or in its signalling pathway are a cause of holoprosencephaly (HPE), a disorder in which the developing forebrain fails to correctly separate into right and left hemispheres. HPE is manifested by facial deformities. It is also thought that mutations in this gene or in its signalling pathway may be responsible for VACTERL syndrome, which is characterized by vertebral defects, anal atresia, tracheoesophageal fistula with esophageal atresia, radial and renal dysplasia, cardiac anomalies, and limb abnormalities. Additionally, mutations in a long range enhancer located approximately 1 megabase upstream of this gene disrupt limb patterning and can result in preaxial polydactyly. [provided by RefSeq, Jul 2008] |
Target gene [SHH] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5010607 |
chr7 |
6 |
11 |
0 |
17 |
View |
| 5010608 |
chr7 |
6 |
10 |
0 |
16 |
View |
| 5010690 |
chr7 |
4 |
4 |
0 |
8 |
View |
Target gene [SHH] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
E TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
When the query gene is differentially changed in the dataset, a volcano/bar plot will be displayed.
> Dataset: TCGA_CESC - SHH expression across samples
|
Volcano Plot
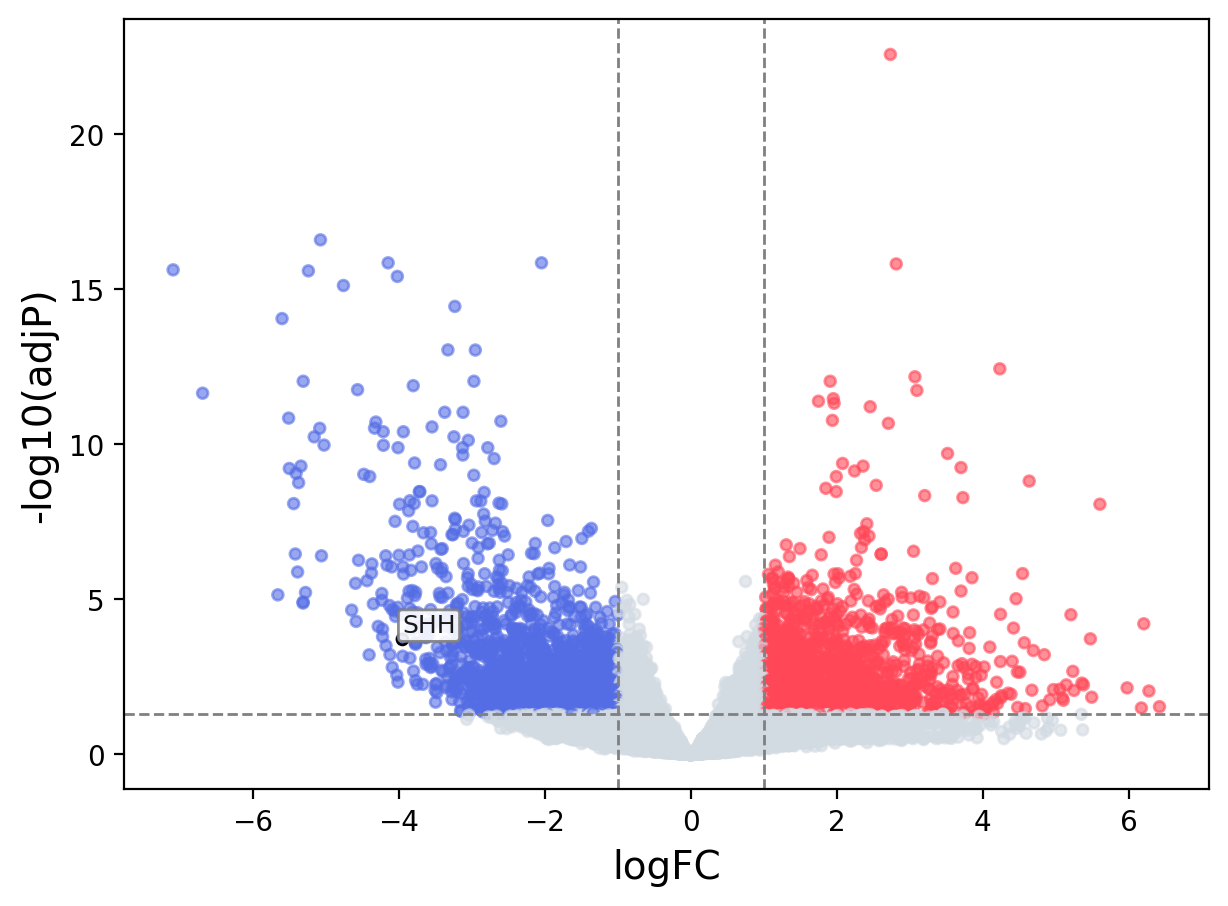
|
Bar Plot
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information