Gene Information
|
Gene Name
|
ATP6V0A4 |
|
Gene ID
|
50617
|
|
Gene Full Name
|
ATPase H+ transporting V0 subunit a4 |
|
Gene Alias
|
A4|ATP6N1B|ATP6N2|DRTA3|RDRTA2|RTA1C|RTADR|STV1|VPH1|VPP2 |
|
Transcripts
|
ENSG00000105929
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Cytosol; 
|
|
Membrane Info
|
Disease related genes, Human disease related genes, Metabolic proteins, Potential drug targets, Predicted intracellular proteins, Predicted membrane proteins, Transporters |
|
Uniport_ID
|
Q9HBG4
|
|
HGNC ID
|
HGNC:866
|
|
OMIM ID
|
605239 |
|
Summary
|
This gene encodes a component of vacuolar ATPase (V-ATPase), a multisubunit enzyme that mediates acidification of intracellular compartments of eukaryotic cells. V-ATPase dependent acidification is necessary for such intracellular processes as protein sorting, zymogen activation, receptor-mediated endocytosis, and synaptic vesicle proton gradient generation. V-ATPase is composed of a cytosolic V1 domain and a transmembrane V0 domain. The V1 domain consists of three A and three B subunits, two G subunits plus the C, D, E, F, and H subunits. The V1 domain contains the ATP catalytic site. The V0 domain consists of five different subunits: a, c, c', c'', and d. This gene is one of four genes in man and mouse that encode different isoforms of the a subunit. Alternatively spliced transcript variants encoding the same protein have been described. Mutations in this gene are associated with renal tubular acidosis associated with preserved hearing. [provided by RefSeq, Jul 2008] |
Target gene [ATP6V0A4] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5005068 |
chr7 |
2 |
4 |
14 |
20 |
View |
| 5008215 |
chr7 |
10 |
12 |
0 |
22 |
View |
Target gene [ATP6V0A4] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE40774
|
Expression array |
134 |
Agilent-026652 Whole Human Genome Microarray 4x44K v2 (Probe Name version) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE65858
|
Expression array |
270 |
Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE138080
|
Expression array |
35 |
Agilent-014850 Whole Human Genome Microarray 4x44K G4112F (Feature Number version) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
When the gene can detect a peak in the dataset, a peak plot will be displayed.
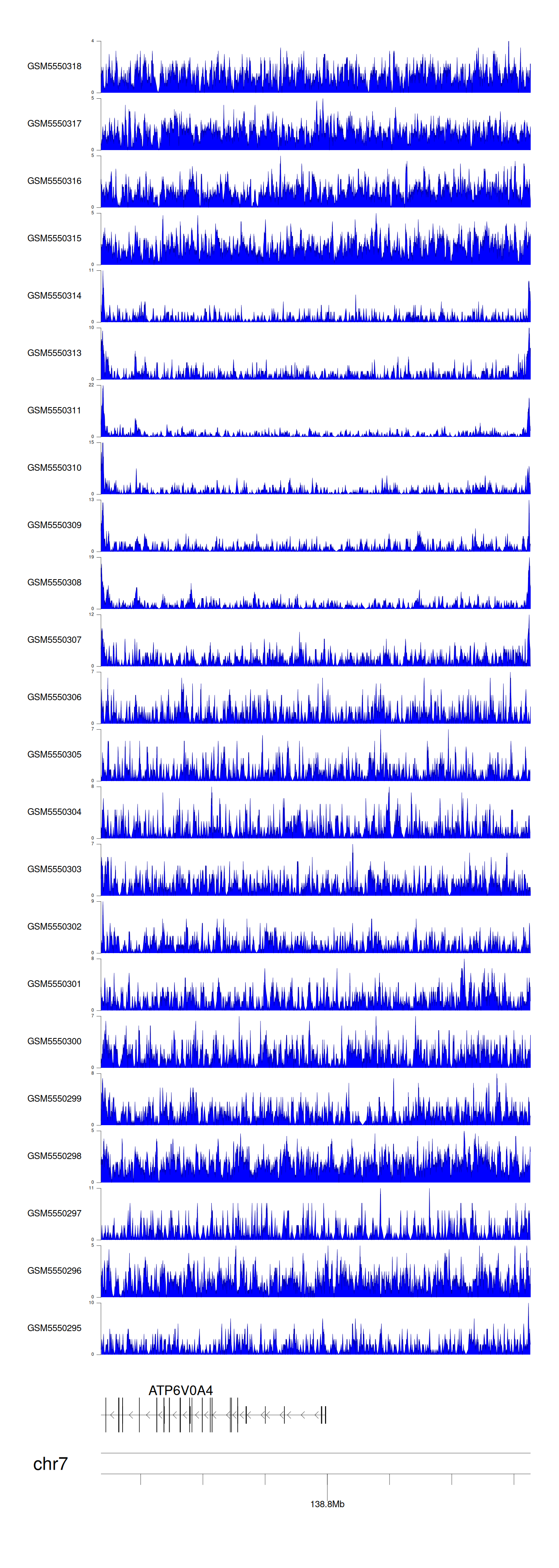
> Dataset: GSE183048 - ATP6V0A4 peak across samples
|
Peak Plot
|
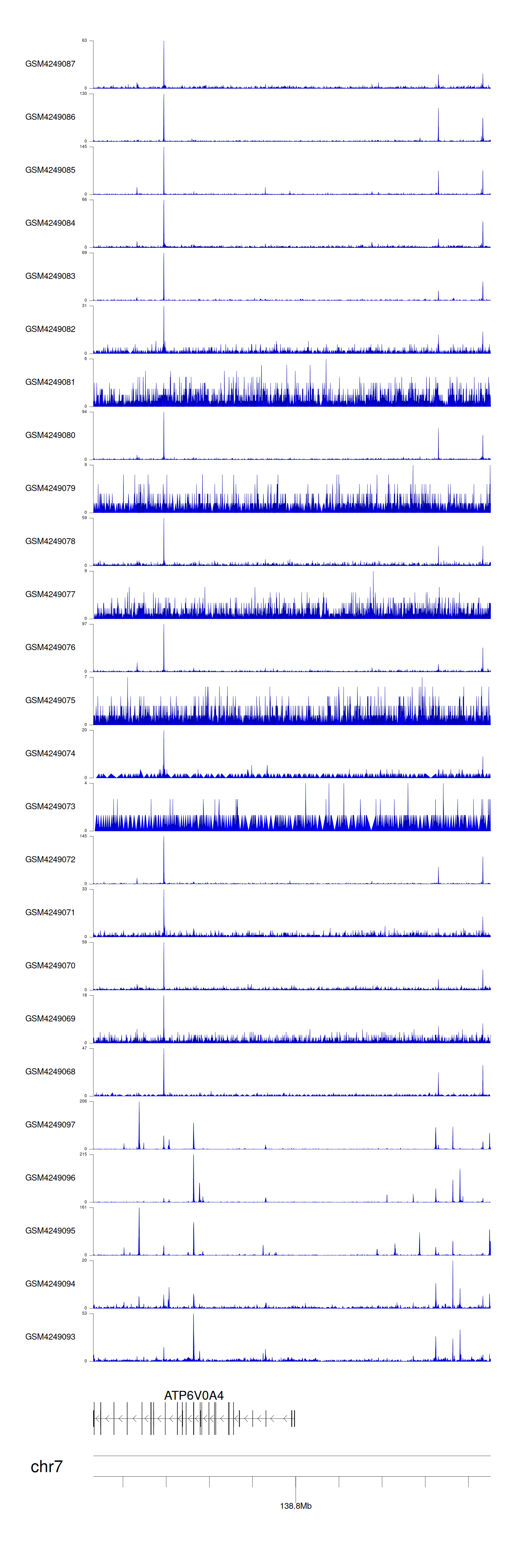
> Dataset: GSE143026 - ATP6V0A4 peak across samples
|
Peak Plot
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information