Gene Information
|
Gene Name
|
PTRH2 |
|
Gene ID
|
51651
|
|
Gene Full Name
|
peptidyl-tRNA hydrolase 2 |
|
Gene Alias
|
BIT1|CFAP37|CGI-147|IMNEPD|PTH|PTH 2|PTH2 |
|
Transcripts
|
ENSG00000141378
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Mitochondria; 
|
|
Membrane Info
|
Disease related genes, Enzymes, Human disease related genes, Metabolic proteins, Potential drug targets, Predicted membrane proteins |
|
Uniport_ID
|
Q9Y3E5
|
|
HGNC ID
|
HGNC:24265
|
|
OMIM ID
|
608625 |
|
Summary
|
The protein encoded by this gene is a mitochondrial protein with two putative domains, an N-terminal mitochondrial localization sequence, and a UPF0099 domain. In vitro assays suggest that this protein possesses peptidyl-tRNA hydrolase activity, to release the peptidyl moiety from tRNA, thereby preventing the accumulation of dissociated peptidyl-tRNA that could reduce the efficiency of translation. This protein also plays a role regulating cell survival and death. It promotes survival as part of an integrin-signaling pathway for cells attached to the extracellular matrix (ECM), but also promotes apoptosis in cells that have lost their attachment to the ECM, a process called anoikos. After loss of cell attachment to the ECM, this protein is phosphorylated, is released from the mitochondria into the cytosol, and promotes caspase-independent apoptosis through interactions with transcriptional regulators. This gene has been implicated in the development and progression of tumors, and mutations in this gene have been associated with an infantile multisystem neurologic, endocrine, and pancreatic disease (INMEPD) characterized by intellectual disability, postnatal microcephaly, progressive cerebellar atrophy, hearing impairment, polyneuropathy, failure to thrive, and organ fibrosis with exocrine pancreas insufficiency (PMID: 25574476). Alternative splicing results in multiple transcript variants encoding different isoforms. [provided by RefSeq, Mar 2015] |
Target gene [PTRH2] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5015246 |
chr17 |
25 |
6 |
0 |
31 |
View |
| 5015247 |
chr17 |
7 |
0 |
1 |
8 |
View |
Target gene [PTRH2] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE40774
|
Expression array |
134 |
Agilent-026652 Whole Human Genome Microarray 4x44K v2 (Probe Name version) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE65858
|
Expression array |
270 |
Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE138080
|
Expression array |
35 |
Agilent-014850 Whole Human Genome Microarray 4x44K G4112F (Feature Number version) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
When the gene can detect a peak in the dataset, a peak plot will be displayed.
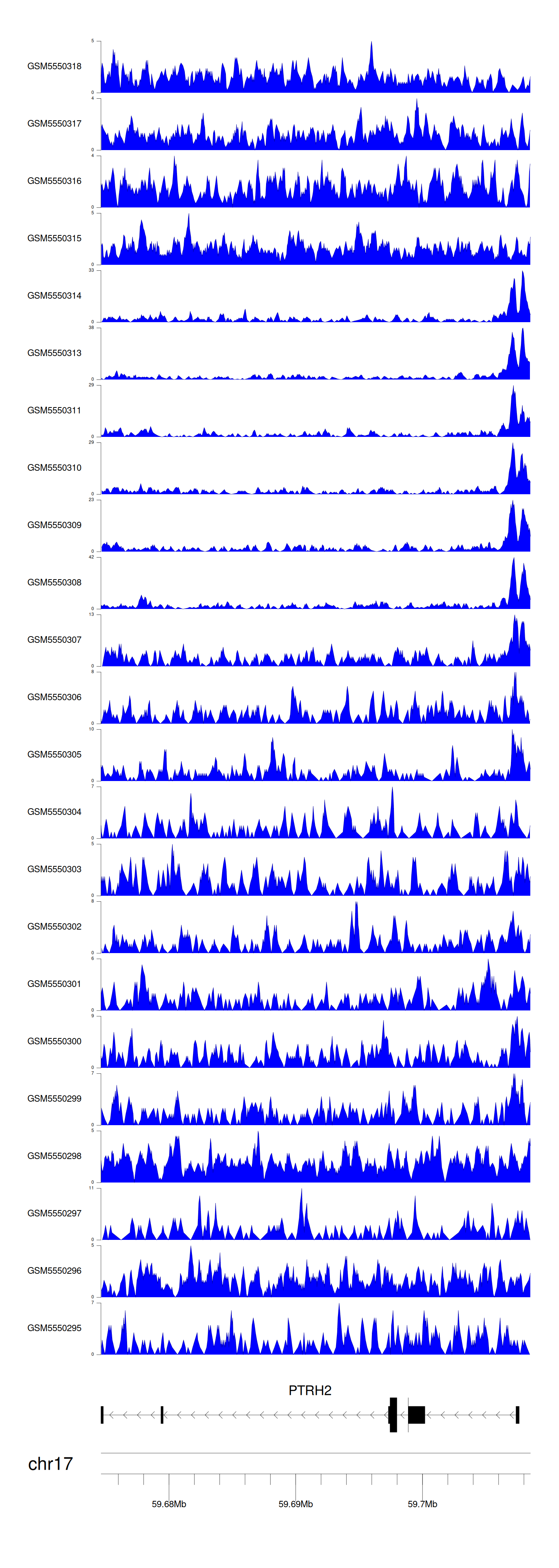
> Dataset: GSE183048 - PTRH2 peak across samples
|
Peak Plot
|
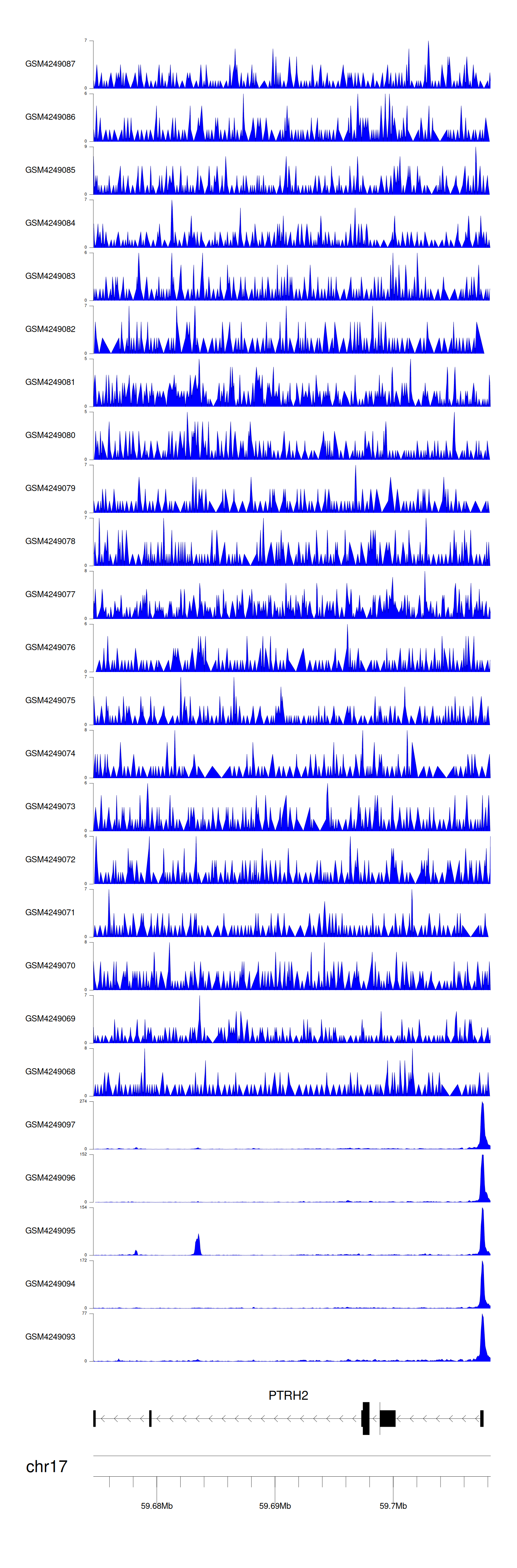
> Dataset: GSE143026 - PTRH2 peak across samples
|
Peak Plot
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information