GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0000122 | Negative regulation of transcription by RNA polymerase II |
| Process | GO:0001568 | Blood vessel development |
| Process | GO:0001654 | Eye development |
| Process | GO:0001655 | Urogenital system development |
| Process | GO:0001656 | Metanephros development |
| Process | GO:0001822 | Kidney development |
| Process | GO:0001944 | Vasculature development |
| Process | GO:0002791 | Regulation of peptide secretion |
| Process | GO:0003008 | System process |
| Process | GO:0003014 | Renal system process |
| Process | GO:0003091 | Renal water homeostasis |
| Process | GO:0003407 | Neural retina development |
| Process | GO:0005975 | Carbohydrate metabolic process |
| Process | GO:0005996 | Monosaccharide metabolic process |
| Process | GO:0006006 | Glucose metabolic process |
| Process | GO:0006139 | Nucleobase-containing compound metabolic process |
| Process | GO:0006351 | Transcription, DNA-templated |
| Process | GO:0006355 | Regulation of transcription, DNA-templated |
| Process | GO:0006357 | Regulation of transcription by RNA polymerase II |
| Process | GO:0006366 | Transcription by RNA polymerase II |
| Process | GO:0006725 | Cellular aromatic compound metabolic process |
| Process | GO:0006807 | Nitrogen compound metabolic process |
| Process | GO:0006873 | Cellular ion homeostasis |
| Process | GO:0006915 | Apoptotic process |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0007399 | Nervous system development |
| Process | GO:0007423 | Sensory organ development |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0008219 | Cell death |
| Process | GO:0008284 | Positive regulation of cell population proliferation |
| Process | GO:0008285 | Negative regulation of cell population proliferation |
| Process | GO:0009058 | Biosynthetic process |
| Process | GO:0009059 | Macromolecule biosynthetic process |
| Process | GO:0009410 | Response to xenobiotic stimulus |
| Process | GO:0009653 | Anatomical structure morphogenesis |
| Process | GO:0009887 | Animal organ morphogenesis |
| Process | GO:0009888 | Tissue development |
| Process | GO:0009889 | Regulation of biosynthetic process |
| Process | GO:0009890 | Negative regulation of biosynthetic process |
| Process | GO:0009891 | Positive regulation of biosynthetic process |
| Process | GO:0009892 | Negative regulation of metabolic process |
| Process | GO:0009893 | Positive regulation of metabolic process |
| Process | GO:0009966 | Regulation of signal transduction |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010467 | Gene expression |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010556 | Regulation of macromolecule biosynthetic process |
| Process | GO:0010557 | Positive regulation of macromolecule biosynthetic process |
| Process | GO:0010558 | Negative regulation of macromolecule biosynthetic process |
| Process | GO:0010604 | Positive regulation of macromolecule metabolic process |
| Process | GO:0010605 | Negative regulation of macromolecule metabolic process |
| Process | GO:0010646 | Regulation of cell communication |
| Process | GO:0010817 | Regulation of hormone levels |
| Process | GO:0010842 | Retina layer formation |
| Process | GO:0010941 | Regulation of cell death |
| Process | GO:0010942 | Positive regulation of cell death |
| Process | GO:0010960 | Magnesium ion homeostasis |
| Process | GO:0012501 | Programmed cell death |
| Process | GO:0016070 | RNA metabolic process |
| Process | GO:0018130 | Heterocycle biosynthetic process |
| Process | GO:0019219 | Regulation of nucleobase-containing compound metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0019318 | Hexose metabolic process |
| Process | GO:0019438 | Aromatic compound biosynthetic process |
| Process | GO:0019725 | Cellular homeostasis |
| Process | GO:0023051 | Regulation of signaling |
| Process | GO:0030003 | Cellular cation homeostasis |
| Process | GO:0030004 | Cellular monovalent inorganic cation homeostasis |
| Process | GO:0030104 | Water homeostasis |
| Process | GO:0030154 | Cell differentiation |
| Process | GO:0030510 | Regulation of BMP signaling pathway |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0031324 | Negative regulation of cellular metabolic process |
| Process | GO:0031325 | Positive regulation of cellular metabolic process |
| Process | GO:0031326 | Regulation of cellular biosynthetic process |
| Process | GO:0031327 | Negative regulation of cellular biosynthetic process |
| Process | GO:0031328 | Positive regulation of cellular biosynthetic process |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032774 | RNA biosynthetic process |
| Process | GO:0032879 | Regulation of localization |
| Process | GO:0032880 | Regulation of protein localization |
| Process | GO:0033500 | Carbohydrate homeostasis |
| Process | GO:0034641 | Cellular nitrogen compound metabolic process |
| Process | GO:0034654 | Nucleobase-containing compound biosynthetic process |
| Process | GO:0035107 | Appendage morphogenesis |
| Process | GO:0035108 | Limb morphogenesis |
| Process | GO:0035136 | Forelimb morphogenesis |
| Process | GO:0035137 | Hindlimb morphogenesis |
| Process | GO:0035239 | Tube morphogenesis |
| Process | GO:0035295 | Tube development |
| Process | GO:0035809 | Regulation of urine volume |
| Process | GO:0035810 | Positive regulation of urine volume |
| Process | GO:0035904 | Aorta development |
| Process | GO:0035909 | Aorta morphogenesis |
| Process | GO:0042127 | Regulation of cell population proliferation |
| Process | GO:0042221 | Response to chemical |
| Process | GO:0042592 | Homeostatic process |
| Process | GO:0042593 | Glucose homeostasis |
| Process | GO:0042981 | Regulation of apoptotic process |
| Process | GO:0043010 | Camera-type eye development |
| Process | GO:0043065 | Positive regulation of apoptotic process |
| Process | GO:0043066 | Negative regulation of apoptotic process |
| Process | GO:0043067 | Regulation of programmed cell death |
| Process | GO:0043068 | Positive regulation of programmed cell death |
| Process | GO:0043069 | Negative regulation of programmed cell death |
| Process | GO:0043170 | Macromolecule metabolic process |
| Process | GO:0043523 | Regulation of neuron apoptotic process |
| Process | GO:0043524 | Negative regulation of neuron apoptotic process |
| Process | GO:0043525 | Positive regulation of neuron apoptotic process |
| Process | GO:0043588 | Skin development |
| Process | GO:0044237 | Cellular metabolic process |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0044249 | Cellular biosynthetic process |
| Process | GO:0044271 | Cellular nitrogen compound biosynthetic process |
| Process | GO:0044281 | Small molecule metabolic process |
| Process | GO:0045444 | Fat cell differentiation |
| Process | GO:0045595 | Regulation of cell differentiation |
| Process | GO:0045892 | Negative regulation of transcription, DNA-templated |
| Process | GO:0045893 | Positive regulation of transcription, DNA-templated |
| Process | GO:0045934 | Negative regulation of nucleobase-containing compound metabolic process |
| Process | GO:0045935 | Positive regulation of nucleobase-containing compound metabolic process |
| Process | GO:0045944 | Positive regulation of transcription by RNA polymerase II |
| Process | GO:0046483 | Heterocycle metabolic process |
| Process | GO:0046883 | Regulation of hormone secretion |
| Process | GO:0048483 | Autonomic nervous system development |
| Process | GO:0048485 | Sympathetic nervous system development |
| Process | GO:0048513 | Animal organ development |
| Process | GO:0048514 | Blood vessel morphogenesis |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048519 | Negative regulation of biological process |
| Process | GO:0048522 | Positive regulation of cellular process |
| Process | GO:0048523 | Negative regulation of cellular process |
| Process | GO:0048583 | Regulation of response to stimulus |
| Process | GO:0048592 | Eye morphogenesis |
| Process | GO:0048593 | Camera-type eye morphogenesis |
| Process | GO:0048646 | Anatomical structure formation involved in morphogenesis |
| Process | GO:0048731 | System development |
| Process | GO:0048736 | Appendage development |
| Process | GO:0048745 | Smooth muscle tissue development |
| Process | GO:0048844 | Artery morphogenesis |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0048869 | Cellular developmental process |
| Process | GO:0048871 | Multicellular organismal homeostasis |
| Process | GO:0048878 | Chemical homeostasis |
| Process | GO:0048880 | Sensory system development |
| Process | GO:0050708 | Regulation of protein secretion |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050793 | Regulation of developmental process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0050796 | Regulation of insulin secretion |
| Process | GO:0050801 | Ion homeostasis |
| Process | GO:0050878 | Regulation of body fluid levels |
| Process | GO:0050891 | Multicellular organismal water homeostasis |
| Process | GO:0050896 | Response to stimulus |
| Process | GO:0051046 | Regulation of secretion |
| Process | GO:0051049 | Regulation of transport |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051172 | Negative regulation of nitrogen compound metabolic process |
| Process | GO:0051173 | Positive regulation of nitrogen compound metabolic process |
| Process | GO:0051223 | Regulation of protein transport |
| Process | GO:0051252 | Regulation of RNA metabolic process |
| Process | GO:0051253 | Negative regulation of RNA metabolic process |
| Process | GO:0051254 | Positive regulation of RNA metabolic process |
| Process | GO:0051402 | Neuron apoptotic process |
| Process | GO:0055062 | Phosphate ion homeostasis |
| Process | GO:0055065 | Metal ion homeostasis |
| Process | GO:0055067 | Monovalent inorganic cation homeostasis |
| Process | GO:0055074 | Calcium ion homeostasis |
| Process | GO:0055075 | Potassium ion homeostasis |
| Process | GO:0055078 | Sodium ion homeostasis |
| Process | GO:0055080 | Cation homeostasis |
| Process | GO:0055081 | Anion homeostasis |
| Process | GO:0055082 | Cellular chemical homeostasis |
| Process | GO:0060041 | Retina development in camera-type eye |
| Process | GO:0060042 | Retina morphogenesis in camera-type eye |
| Process | GO:0060173 | Limb development |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0060341 | Regulation of cellular localization |
| Process | GO:0060429 | Epithelium development |
| Process | GO:0060537 | Muscle tissue development |
| Process | GO:0060548 | Negative regulation of cell death |
| Process | GO:0060840 | Artery development |
| Process | GO:0061326 | Renal tubule development |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065008 | Regulation of biological quality |
| Process | GO:0070201 | Regulation of establishment of protein localization |
| Process | GO:0070997 | Neuron death |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0072001 | Renal system development |
| Process | GO:0072006 | Nephron development |
| Process | GO:0072009 | Nephron epithelium development |
| Process | GO:0072017 | Distal tubule development |
| Process | GO:0072044 | Collecting duct development |
| Process | GO:0072073 | Kidney epithelium development |
| Process | GO:0072080 | Nephron tubule development |
| Process | GO:0072210 | Metanephric nephron development |
| Process | GO:0072359 | Circulatory system development |
| Process | GO:0072506 | Trivalent inorganic anion homeostasis |
| Process | GO:0072507 | Divalent inorganic cation homeostasis |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0090087 | Regulation of peptide transport |
| Process | GO:0090092 | Regulation of transmembrane receptor protein serine/threonine kinase signaling pathway |
| Process | GO:0090276 | Regulation of peptide hormone secretion |
| Process | GO:0090287 | Regulation of cellular response to growth factor stimulus |
| Process | GO:0090304 | Nucleic acid metabolic process |
| Process | GO:0090596 | Sensory organ morphogenesis |
| Process | GO:0097070 | Ductus arteriosus closure |
| Process | GO:0097272 | Ammonium homeostasis |
| Process | GO:0097273 | Creatinine homeostasis |
| Process | GO:0097274 | Urea homeostasis |
| Process | GO:0097275 | Cellular ammonium homeostasis |
| Process | GO:0097276 | Cellular creatinine homeostasis |
| Process | GO:0097277 | Cellular urea homeostasis |
| Process | GO:0097659 | Nucleic acid-templated transcription |
| Process | GO:0098771 | Inorganic ion homeostasis |
| Process | GO:0150063 | Visual system development |
| Process | GO:1901214 | Regulation of neuron death |
| Process | GO:1901215 | Negative regulation of neuron death |
| Process | GO:1901216 | Positive regulation of neuron death |
| Process | GO:1901360 | Organic cyclic compound metabolic process |
| Process | GO:1901362 | Organic cyclic compound biosynthetic process |
| Process | GO:1901576 | Organic substance biosynthetic process |
| Process | GO:1902679 | Negative regulation of RNA biosynthetic process |
| Process | GO:1902680 | Positive regulation of RNA biosynthetic process |
| Process | GO:1903506 | Regulation of nucleic acid-templated transcription |
| Process | GO:1903507 | Negative regulation of nucleic acid-templated transcription |
| Process | GO:1903508 | Positive regulation of nucleic acid-templated transcription |
| Process | GO:1903530 | Regulation of secretion by cell |
| Process | GO:2001141 | Regulation of RNA biosynthetic process |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005634 | Nucleus |
| Component | GO:0043226 | Organelle |
| Component | GO:0043227 | Membrane-bounded organelle |
| Component | GO:0043229 | Intracellular organelle |
| Component | GO:0043231 | Intracellular membrane-bounded organelle |
| Component | GO:0110165 | Cellular anatomical entity |
| Function | GO:0000976 | Transcription cis-regulatory region binding |
| Function | GO:0000977 | RNA polymerase II transcription regulatory region sequence-specific DNA binding |
| Function | GO:0000978 | RNA polymerase II cis-regulatory region sequence-specific DNA binding |
| Function | GO:0000981 | DNA-binding transcription factor activity, RNA polymerase II-specific |
| Function | GO:0000987 | Cis-regulatory region sequence-specific DNA binding |
| Function | GO:0001067 | Transcription regulatory region nucleic acid binding |
| Function | GO:0001216 | DNA-binding transcription activator activity |
| Function | GO:0001228 | DNA-binding transcription activator activity, RNA polymerase II-specific |
| Function | GO:0003676 | Nucleic acid binding |
| Function | GO:0003677 | DNA binding |
| Function | GO:0003682 | Chromatin binding |
| Function | GO:0003690 | Double-stranded DNA binding |
| Function | GO:0003700 | DNA-binding transcription factor activity |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0042802 | Identical protein binding |
| Function | GO:0042803 | Protein homodimerization activity |
| Function | GO:0043565 | Sequence-specific DNA binding |
| Function | GO:0046982 | Protein heterodimerization activity |
| Function | GO:0046983 | Protein dimerization activity |
| Function | GO:0097159 | Organic cyclic compound binding |
| Function | GO:0140110 | Transcription regulator activity |
| Function | GO:1901363 | Heterocyclic compound binding |
| Function | GO:1990837 | Sequence-specific double-stranded DNA binding |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| RCTM | MMU-212436 | Generic Transcription Pathway |
| RCTM | MMU-73857 | RNA Polymerase II Transcription |
| RCTM | MMU-74160 | Gene expression (Transcription) |
| RCTM | MMU-8864260 | Transcriptional regulation by the AP-2 (TFAP2) family of transcription factors |
| RCTM | MMU-8866904 | Negative regulation of activity of TFAP2 (AP-2) family transcription factors |
| RCTM | MMU-8866907 | Activation of the TFAP2 (AP-2) family of transcription factors |
| WikiPathways | WP2074 | Neural crest differentiation |
Expression profile of Tfap2b in omics data
| ID | GSE106799 |
| Title | Propofol alters the mRNA profile in the developing mouse hippocampus |
| Organism | Mus musculus |
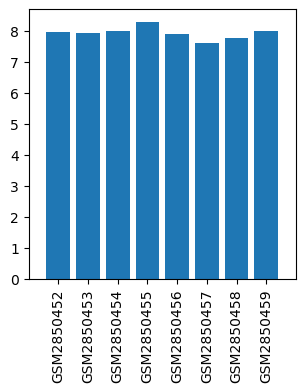
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2850452 | Control 1 | tissue: hippocampus age: 7 days-old | 7.950949 |
| GSM2850453 | Control 2 | tissue: hippocampus age: 7 days-old | 7.9353027 |
| GSM2850454 | Control 3 | tissue: hippocampus age: 7 days-old | 7.9934497 |
| GSM2850455 | Control 4 | tissue: hippocampus age: 7 days-old | 8.28029 |
| GSM2850456 | Propofol 1 | tissue: hippocampus age: 7 days-old | 7.8958755 |
| GSM2850457 | Propofol 2 | tissue: hippocampus age: 7 days-old | 7.5970526 |
| GSM2850458 | Propofol 3 | tissue: hippocampus age: 7 days-old | 7.7698474 |
| GSM2850459 | Propofol 4 | tissue: hippocampus age: 7 days-old | 7.9898467 |
| ID | GSE109165 |
| Title | Suppression of RGSz1 function optimizes the actions of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway |
| Organism | Mus musculus |
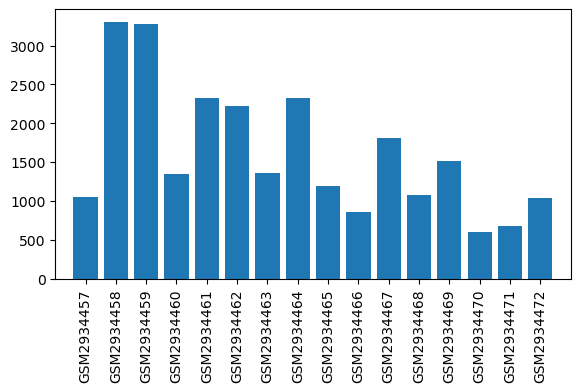
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2934457 | WT_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 1048.0 |
| GSM2934458 | WT_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 3304.0 |
| GSM2934459 | WT_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 3279.0 |
| GSM2934460 | WT_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 1344.0 |
| GSM2934461 | KO_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 2320.0 |
| GSM2934462 | KO_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 2219.0 |
| GSM2934463 | KO_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 1360.0 |
| GSM2934464 | KO_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 2326.0 |
| GSM2934465 | WT_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 1196.0 |
| GSM2934466 | WT_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 862.0 |
| GSM2934467 | WT_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 1813.0 |
| GSM2934468 | WT_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 1072.0 |
| GSM2934469 | KO_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 1512.0 |
| GSM2934470 | KO_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 599.0 |
| GSM2934471 | KO_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 683.0 |
| GSM2934472 | KO_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 1041.0 |
| ID | GSE125076 |
| Title | A mouse model of acute post-surgical pain |
| Organism | Mus musculus |
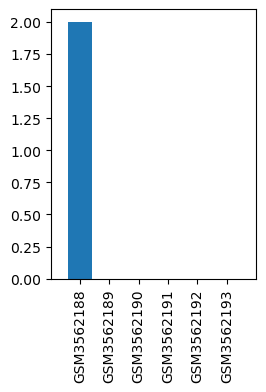
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM3562188 | DRG_WT_sham_1 | strain: C57BL/6 gender: male age: 8 weeks treatment: naïve | 2.0 |
| GSM3562189 | DRG_WT_sham_2 | strain: C57BL/6 gender: male age: 8 weeks treatment: naïve | 0.0 |
| GSM3562190 | DRG_WT_sham_3 | strain: C57BL/6 gender: male age: 8 weeks treatment: naïve | 0.0 |
| GSM3562191 | DRG_WT_PSP_1 | strain: C57BL/6 gender: male age: 8 weeks treatment: 24h post surgery | 0.0 |
| GSM3562192 | DRG_WT_PSP_2 | strain: C57BL/6 gender: male age: 8 weeks treatment: 24h post surgery | 0.0 |
| GSM3562193 | DRG_WT_PSP_3 | strain: C57BL/6 gender: male age: 8 weeks treatment: 24h post surgery | 0.0 |
| ID | GSE126662 |
| Title | Comparison between opioid (morphine) and vehicle treatments in mouse trigeminal ganglia and nucleus accumbens |
| Organism | Mus musculus |
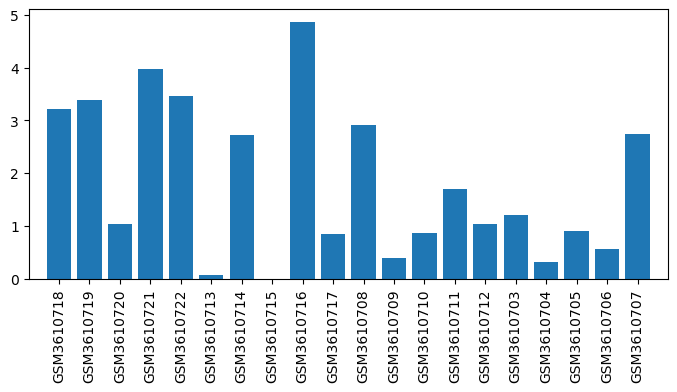
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM3610703 | TG_M1: OIH_M1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 1.1996383333333334 |
| GSM3610704 | TG_M2: OIH_M2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 0.3143533333333333 |
| GSM3610705 | TG_M3: OIH_M3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 0.90634 |
| GSM3610706 | TG_M4: OIH_M4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 0.5586816666666666 |
| GSM3610707 | TG_M5: OIH_M5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 2.7444395 |
| GSM3610708 | TG_V1: VehicleOIH_V1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 2.9059111666666664 |
| GSM3610709 | TG_V2: VehicleOIH_V2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 0.39179499999999995 |
| GSM3610710 | TG_V3: VehicleOIH_V3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 0.855875 |
| GSM3610711 | TG_V4: VehicleOIH_V4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 1.693876666666667 |
| GSM3610712 | TG_V5: VehicleOIH_V5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 1.0350466666666667 |
| GSM3610713 | NAc_M1: OIH_M1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 0.06993733333333334 |
| GSM3610714 | NAc_M2: OIH_M2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 2.7209333333333334 |
| GSM3610715 | NAc_M3: OIH_M3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 0.0 |
| GSM3610716 | NAc_M4: OIH_M4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 4.863131666666667 |
| GSM3610717 | NAc_M5: OIH_M5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 0.8415516666666667 |
| GSM3610718 | NAc_V1: VehicleOIH_V1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 3.2168166666666664 |
| GSM3610719 | NAc_V2: VehicleOIH_V2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 3.387408333333333 |
| GSM3610720 | NAc_V3: VehicleOIH_V3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 1.0332166666666667 |
| GSM3610721 | NAc_V4: VehicleOIH_V4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 3.967636666666667 |
| GSM3610722 | NAc_V5: VehicleOIH_V5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 3.456099 |
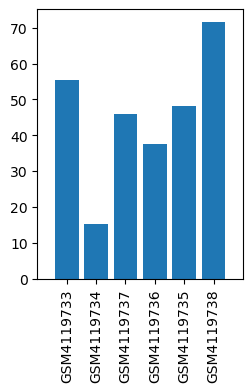
| ID | GSE138802 |
| Title | Mesocorticolimbic circuit mechanisms underlying the effects of ketamine on dopamine: a translational imaging study |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4119733 | PLc_1_Sal | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Saline | 55.56 |
| GSM4119734 | PLc_10_Sal | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Saline | 15.37 |
| GSM4119735 | PLc_6_Sal | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Saline | 48.26 |
| GSM4119736 | PLc_5_Ket | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Ketamine | 37.63 |
| GSM4119737 | PLc_3_Ket | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Ketamine | 46.01 |
| GSM4119738 | PLc_9_Ket | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Ketamine | 71.61 |
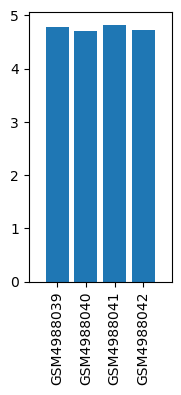
| ID | GSE163828 |
| Title | Regulation of electro-acupuncture(EA) and astragalus membranaceus (AM) on genes in bone marrow |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4988039 | BH190126-1_control | tissue: bone marrow | 4.79 |
| GSM4988040 | BH190126-1_treatment of astragalus membranaceus | tissue: bone marrow | 4.7 |
| GSM4988041 | BH190126-1_cisplatin | tissue: bone marrow | 4.82 |
| GSM4988042 | BH190126-1_treatment of electro-acupuncture | tissue: bone marrow | 4.72 |
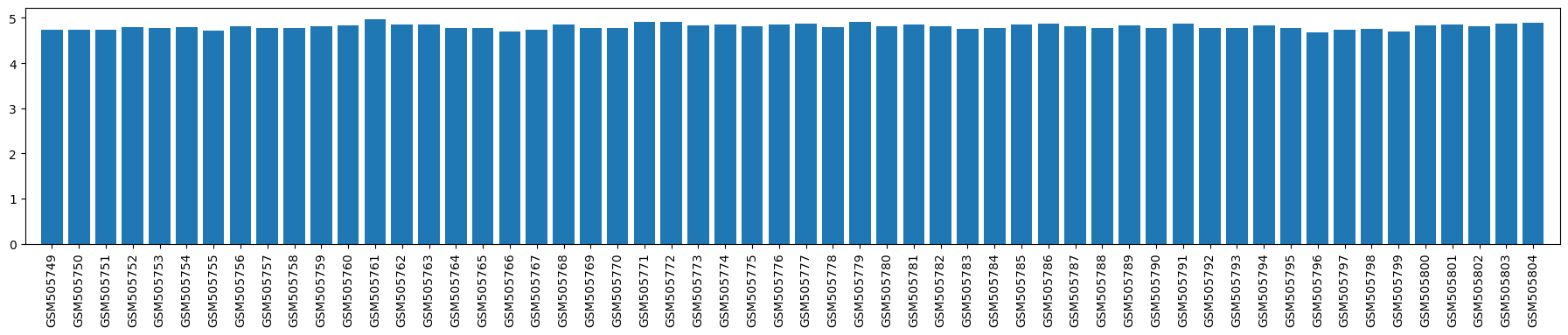
| ID | GSE20160 |
| Title | Gene Expression Data from Prefrontal Cortex and Nucleus Accumbens from Inbred Strains of Mice |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM505749 | 129S1/SvImJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: 129S1/SvImJ | 4.731999999999999 |
| GSM505750 | A/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: A/J | 4.7445 |
| GSM505751 | AKR/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: AKR/J | 4.744 |
| GSM505752 | BALB/cByJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: BALB/cByJ | 4.8045 |
| GSM505753 | BTBR_T+_tf/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: BTBR_T+_tf/J | 4.769500000000001 |
| GSM505754 | BUB/BnJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: BUB/BnJ | 4.803 |
| GSM505755 | C3H/HeJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C3H/HeJ | 4.71 |
| GSM505756 | C57BL/6J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C57BL/6J | 4.8115000000000006 |
| GSM505757 | C57BR/cdJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C57BR/cdJ | 4.779 |
| GSM505758 | C58/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C58/J | 4.7765 |
| GSM505759 | CBA/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: CBA/J | 4.8115 |
| GSM505760 | CE/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: CE/J | 4.8415 |
| GSM505761 | DBA/2J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: DBA/2J | 4.9735 |
| GSM505762 | FVB/NJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: FVB/NJ | 4.8625 |
| GSM505763 | I/LnJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: I/LnJ | 4.8575 |
| GSM505764 | KK/HlJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: KK/HlJ | 4.7844999999999995 |
| GSM505765 | MA/MyJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: MA/MyJ | 4.769 |
| GSM505766 | MRL_MpJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: MRL/MpJ | 4.702999999999999 |
| GSM505767 | NOD/LtJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NOD/LtJ | 4.734500000000001 |
| GSM505768 | NON/LtJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NON/LtJ | 4.854 |
| GSM505769 | NZO/HILtJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NZO/HILtJ | 4.784000000000001 |
| GSM505770 | NZW_LacJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NZW_LacJ | 4.7765 |
| GSM505771 | P/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: P/J | 4.92 |
| GSM505772 | PL/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: PL/J | 4.913 |
| GSM505773 | RIIIS/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: RIIIS/J | 4.8445 |
| GSM505774 | SJL/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: SJL/J | 4.85 |
| GSM505775 | SM/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: SM/J | 4.8245000000000005 |
| GSM505776 | SWR/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: SWR/J | 4.864 |
| GSM505777 | WSB/EiJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: WSB/EiJ | 4.8675 |
| GSM505778 | 129S1/SvImJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: 129S1/SvImJ | 4.7940000000000005 |
| GSM505779 | A/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: A/J | 4.9165 |
| GSM505780 | AKR/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: AKR/J | 4.821 |
| GSM505781 | BALB/cByJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: BALB/cByJ | 4.85 |
| GSM505782 | BTBR_T+_tf/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: BTBR_T+_tf/J | 4.8215 |
| GSM505783 | BUB/BnJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: BUB/BnJ | 4.7535 |
| GSM505784 | C3H/HeJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C3H/HeJ | 4.782500000000001 |
| GSM505785 | C57BL/6J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C57BL/6J | 4.846 |
| GSM505786 | C57BR/cdJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C57BR/cdJ | 4.872 |
| GSM505787 | C58/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C58/J | 4.824 |
| GSM505788 | CBA/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: CBA/J | 4.773 |
| GSM505789 | CE/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: CE/J | 4.8355 |
| GSM505790 | DBA/2J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: DBA/2J | 4.7835 |
| GSM505791 | FVB/NJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: FVB/NJ | 4.87 |
| GSM505792 | I/LnJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: I/LnJ | 4.768000000000001 |
| GSM505793 | KK/HlJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: KK/HlJ | 4.7675 |
| GSM505794 | MA/MyJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: MA/MyJ | 4.8309999999999995 |
| GSM505795 | NOD/LtJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: NOD/LtJ | 4.7705 |
| GSM505796 | NON/LtJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: NON/LtJ | 4.6845 |
| GSM505797 | NZW/LacJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: NZW/LacJ | 4.729 |
| GSM505798 | P/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: P/J | 4.7669999999999995 |
| GSM505799 | PL/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: PL/J | 4.708 |
| GSM505800 | RIIIS/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: RIIIS/J | 4.836499999999999 |
| GSM505801 | SJL/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: SJL/J | 4.861 |
| GSM505802 | SM/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: SM/J | 4.817 |
| GSM505803 | SWR/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: SWR/J | 4.8755 |
| GSM505804 | WSB/EiJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: WSB/EiJ | 4.9005 |
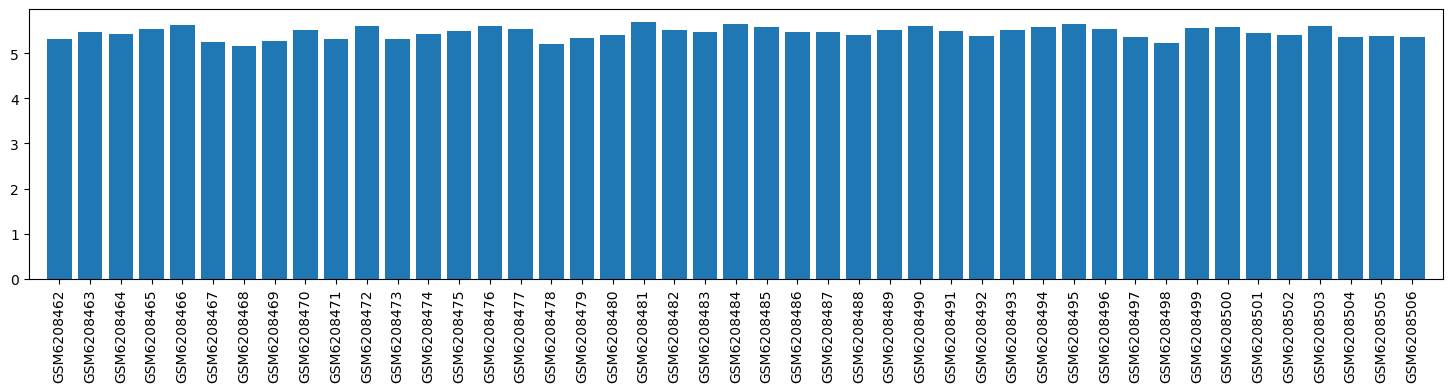
| ID | GSE205201 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [mouse] |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208462 | Mouse_Baseline_0h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 5.308687 |
| GSM6208463 | Mouse_Baseline_0h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 5.480176 |
| GSM6208464 | Mouse_Baseline_0h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 5.421494 |
| GSM6208465 | Mouse_Baseline_0h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 5.541683 |
| GSM6208466 | Mouse_Baseline_0h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 5.633369 |
| GSM6208467 | Mouse_Vehicle_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 5.246946 |
| GSM6208468 | Mouse_Vehicle_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 5.164423 |
| GSM6208469 | Mouse_Vehicle_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 5.276444 |
| GSM6208470 | Mouse_Vehicle_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 5.529023 |
| GSM6208471 | Mouse_Vehicle_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 5.313415 |
| GSM6208472 | Mouse_Vehicle_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 5.610015 |
| GSM6208473 | Mouse_Vehicle_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 5.308649 |
| GSM6208474 | Mouse_Vehicle_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 5.430967 |
| GSM6208475 | Mouse_Vehicle_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 5.489412 |
| GSM6208476 | Mouse_Vehicle_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 5.615752 |
| GSM6208477 | Mouse_Vehicle_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 5.549932 |
| GSM6208478 | Mouse_Vehicle_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 5.204894 |
| GSM6208479 | Mouse_Vehicle_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 5.341013 |
| GSM6208480 | Mouse_Vehicle_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 5.400503 |
| GSM6208481 | Mouse_Vehicle_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 5.69643 |
| GSM6208482 | Mouse_Vehicle_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 5.522622 |
| GSM6208483 | Mouse_Vehicle_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 5.469911 |
| GSM6208484 | Mouse_Vehicle_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 5.647684 |
| GSM6208485 | Mouse_Vehicle_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 5.582515 |
| GSM6208486 | Mouse_Vehicle_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 5.469911 |
| GSM6208487 | Mouse_APAP_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 5.467338 |
| GSM6208488 | Mouse_APAP_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 5.40351 |
| GSM6208489 | Mouse_APAP_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 5.516241 |
| GSM6208490 | Mouse_APAP_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 5.600036 |
| GSM6208491 | Mouse_APAP_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 5.495377 |
| GSM6208492 | Mouse_APAP_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 5.390646 |
| GSM6208493 | Mouse_APAP_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 5.508916 |
| GSM6208494 | Mouse_APAP_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 5.587298 |
| GSM6208495 | Mouse_APAP_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 5.640889 |
| GSM6208496 | Mouse_APAP_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 5.533849 |
| GSM6208497 | Mouse_APAP_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 5.372155 |
| GSM6208498 | Mouse_APAP_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 5.223806 |
| GSM6208499 | Mouse_APAP_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 5.567228 |
| GSM6208500 | Mouse_APAP_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 5.589476 |
| GSM6208501 | Mouse_APAP_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 5.448993 |
| GSM6208502 | Mouse_APAP_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 5.414197 |
| GSM6208503 | Mouse_APAP_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 5.615181 |
| GSM6208504 | Mouse_APAP_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 5.369863 |
| GSM6208505 | Mouse_APAP_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 5.389733 |
| GSM6208506 | Mouse_APAP_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 5.362457 |
| ID | GSE223541 |
| Title | Oxycodone withdrawal induces HDAC1/2-dependent transcriptional maladaptations in the reward pathway in a murine model of peripheral nerve injury |
| Organism | Mus musculus |
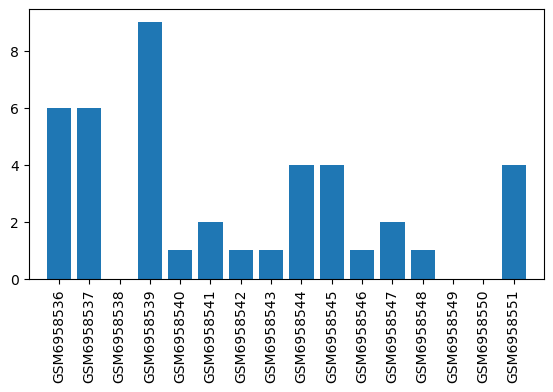
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6958536 | B1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sa | 6.0 |
| GSM6958537 | C1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 6.0 |
| GSM6958538 | F1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958539 | H1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 9.0 |
| GSM6958540 | B2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 1.0 |
| GSM6958541 | C2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 2.0 |
| GSM6958542 | F2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 1.0 |
| GSM6958543 | H2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 1.0 |
| GSM6958544 | B3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 4.0 |
| GSM6958545 | C3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 4.0 |
| GSM6958546 | F3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 1.0 |
| GSM6958547 | H3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 2.0 |
| GSM6958548 | B4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 1.0 |
| GSM6958549 | C4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 0.0 |
| GSM6958550 | F4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 0.0 |
| GSM6958551 | H4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 4.0 |
| ID | GSE24829 |
| Title | Gene expression data from striatal regions of MPTP-intoxicated mouse brain by acupuncture |
| Organism | Mus musculus |
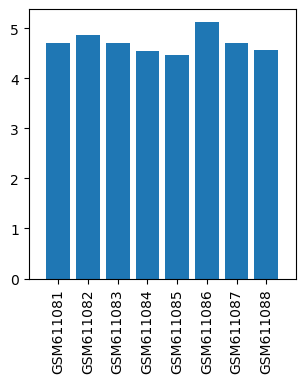
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM611081 | Control mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 4.69706 |
| GSM611082 | Control mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 4.86593 |
| GSM611083 | MPTP-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 4.71109 |
| GSM611084 | MPTP-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 4.54749 |
| GSM611085 | MPTP and acupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 4.46335 |
| GSM611086 | MPTP and acupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 5.12315 |
| GSM611087 | MPTP and nonacupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 4.70307 |
| GSM611088 | MPTP and nonacupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 4.56652 |
| ID | GSE24830 |
| Title | Gene expression data from cervical spinal cord regions of MPTP-intoxicated mouse by acupuncture |
| Organism | Mus musculus |
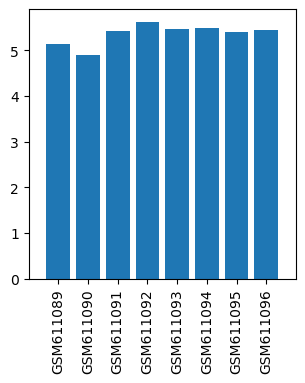
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM611089 | Control mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 5.12631 |
| GSM611090 | Control mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 4.88682 |
| GSM611091 | MPTP-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 5.41202 |
| GSM611092 | MPTP-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 5.62057 |
| GSM611093 | MPTP and acupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 5.45911 |
| GSM611094 | MPTP and acupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 5.47957 |
| GSM611095 | MPTP and nonacupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 5.39078 |
| GSM611096 | MPTP and nonacupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 5.44708 |
| ID | GSE24838 |
| Title | Gene expression data from brain striatal and spinal cord regions of MPTP-intoxicated mouse following acupuncture |
| Organism | Mus musculus |
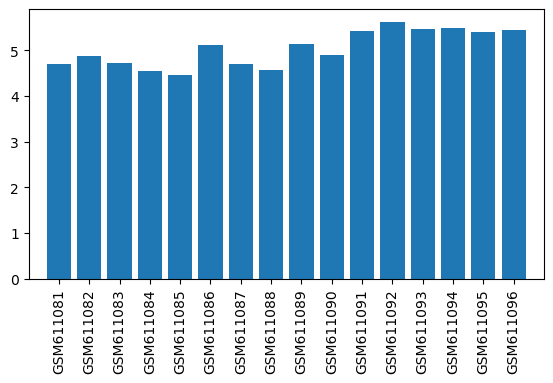
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM611081 | Control mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 4.69706 |
| GSM611082 | Control mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 4.86593 |
| GSM611083 | MPTP-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 4.71109 |
| GSM611084 | MPTP-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 4.54749 |
| GSM611085 | MPTP and acupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 4.46335 |
| GSM611086 | MPTP and acupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 5.12315 |
| GSM611087 | MPTP and nonacupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 4.70307 |
| GSM611088 | MPTP and nonacupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 4.56652 |
| GSM611089 | Control mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 5.12631 |
| GSM611090 | Control mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 4.88682 |
| GSM611091 | MPTP-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 5.41202 |
| GSM611092 | MPTP-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 5.62057 |
| GSM611093 | MPTP and acupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 5.45911 |
| GSM611094 | MPTP and acupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 5.47957 |
| GSM611095 | MPTP and nonacupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 5.39078 |
| GSM611096 | MPTP and nonacupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 5.44708 |
| ID | GSE26364 |
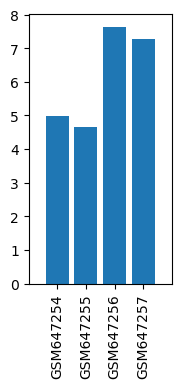
| Title | Gene expression changes in alpha GABA receptors in the mice brains after prolonged ketamine administration |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM647254 | Saline PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 4.99365 |
| GSM647255 | Saline PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 4.64165 |
| GSM647256 | Ketamine PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 7.63408 |
| GSM647257 | Ketamine PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 7.2826 |
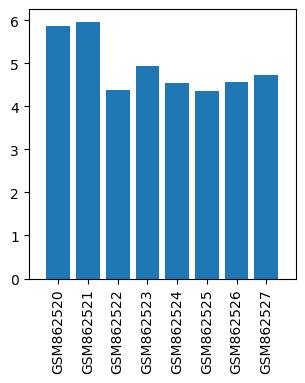
| ID | GSE35138 |
| Title | Gene expression data from thalamic regions of MPTP-intoxicated mouse brain by acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM862520 | Control mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 5.85861 |
| GSM862521 | Control mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 5.95936 |
| GSM862522 | MPTP-treated mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 4.3834 |
| GSM862523 | MPTP-treated mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 4.9311 |
| GSM862524 | MPTP and acupoints acupuncture-treated mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 4.53678 |
| GSM862525 | MPTP and acupoints acupuncture-treated mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 4.34374 |
| GSM862526 | MPTP and nonacupoints acupuncture-treated mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 4.56434 |
| GSM862527 | MPTP and nonacupoints acupuncture-treated mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 4.72705 |
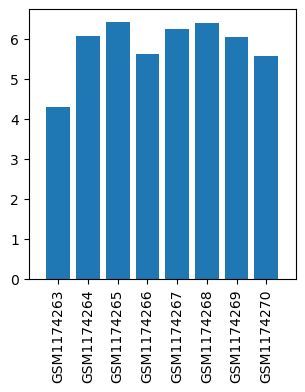
| ID | GSE48293 |
| Title | Gene expression data from substantia nigral regions of MPTP-intoxicated mouse brain by acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1174263 | Control mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: Control | 4.29669 |
| GSM1174264 | Control mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: Control | 6.06297 |
| GSM1174265 | MPTP-treated mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP-treated | 6.41897 |
| GSM1174266 | MPTP-treated mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP-treated | 5.61316 |
| GSM1174267 | MPTP and acupoints acupuncture-treated mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 6.24682 |
| GSM1174268 | MPTP and acupoints acupuncture-treated mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 6.38544 |
| GSM1174269 | MPTP and nonacupoints acupuncture-treated mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 6.04351 |
| GSM1174270 | MPTP and nonacupoints acupuncture-treated mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 5.56538 |

