Gene Information
|
Gene Name
|
HSPG2 |
|
Gene ID
|
3339
|
|
Gene Full Name
|
heparan sulfate proteoglycan 2 |
|
Gene Alias
|
HSPG|PLC|PRCAN|SJA|SJS|SJS1 |
|
Transcripts
|
ENSG00000142798
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Secreted to extracellular matrix; 
|
|
Membrane Info
|
Disease related genes, Human disease related genes, Plasma proteins, Predicted intracellular proteins, Predicted secreted proteins |
|
Uniport_ID
|
P98160
|
|
HGNC ID
|
HGNC:5273
|
|
OMIM ID
|
142461 |
|
Summary
|
This gene encodes the perlecan protein, which consists of a core protein to which three long chains of glycosaminoglycans (heparan sulfate or chondroitin sulfate) are attached. The perlecan protein is a large multidomain proteoglycan that binds to and cross-links many extracellular matrix components and cell-surface molecules. It has been shown that this protein interacts with laminin, prolargin, collagen type IV, FGFBP1, FBLN2, FGF7 and transthyretin, etc., and it plays essential roles in multiple biological activities. Perlecan is a key component of the vascular extracellular matrix, where it helps to maintain the endothelial barrier function. It is a potent inhibitor of smooth muscle cell proliferation and is thus thought to help maintain vascular homeostasis. It can also promote growth factor (e.g., FGF2) activity and thus stimulate endothelial growth and re-generation. It is a major component of basement membranes, where it is involved in the stabilization of other molecules as well as being involved with glomerular permeability to macromolecules and cell adhesion. Mutations in this gene cause Schwartz-Jampel syndrome type 1, Silverman-Handmaker type of dyssegmental dysplasia, and tardive dyskinesia. Alternative splicing of this gene results in multiple transcript variants. [provided by RefSeq, May 2014] |
Target gene [HSPG2] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5010014 |
chr1 |
31 |
4 |
0 |
35 |
View |
| 5016928 |
chr1 |
10 |
10 |
2 |
22 |
View |
Target gene [HSPG2] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
S GSE226620
|
scRNA-seq |
30 |
HiSeq X Ten (Homo sapiens) |
|
GSE40774
|
Expression array |
134 |
Agilent-026652 Whole Human Genome Microarray 4x44K v2 (Probe Name version) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE65858
|
Expression array |
270 |
Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
S GSE164690
|
scRNA-seq |
51 |
Illumina NextSeq 500 (Homo sapiens) |
|
S GSE197461
|
scRNA-seq |
16 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
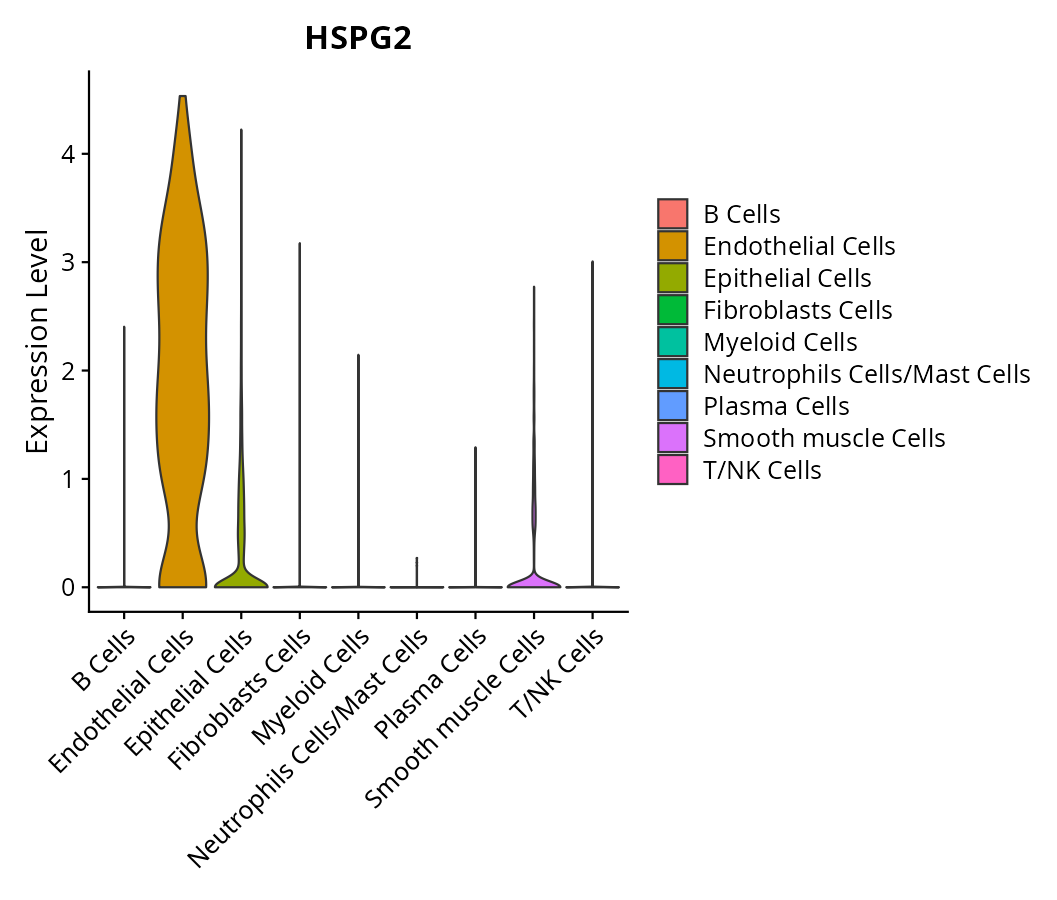
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE226620 - Gene expression in cell subsets
|
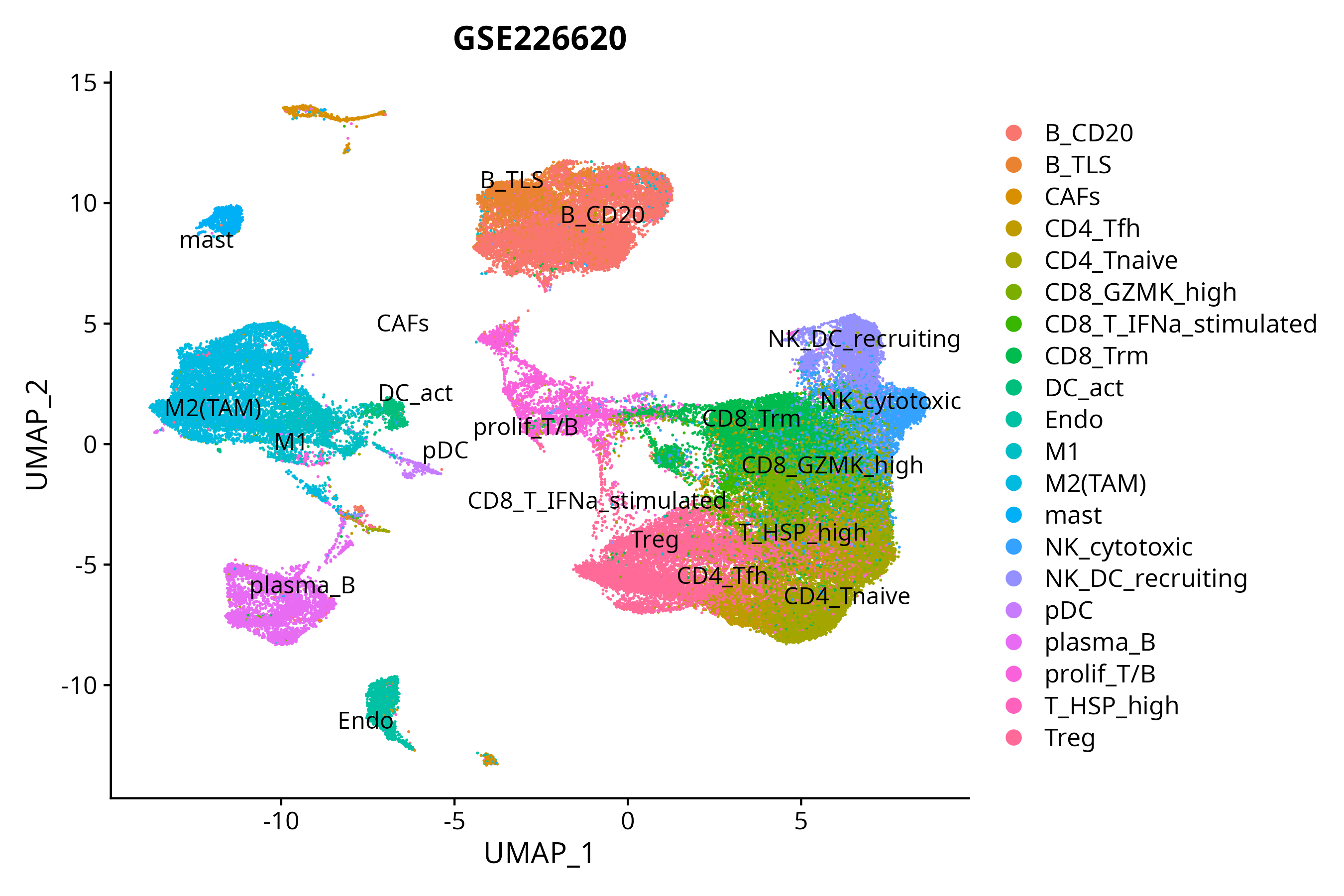
|
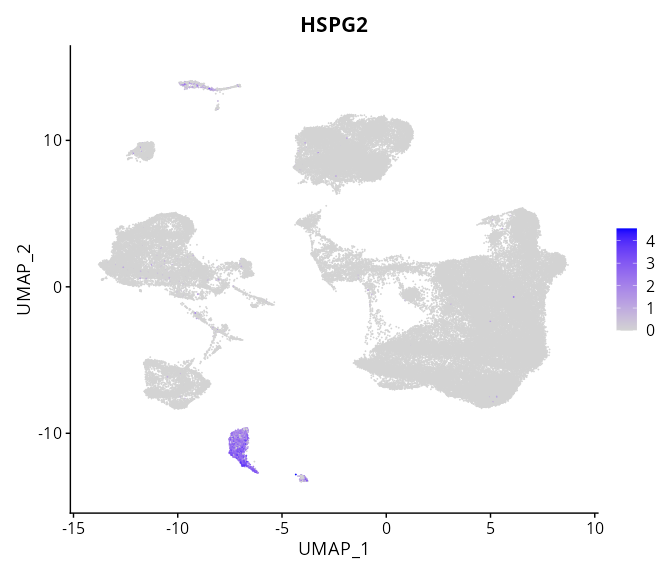
|

|
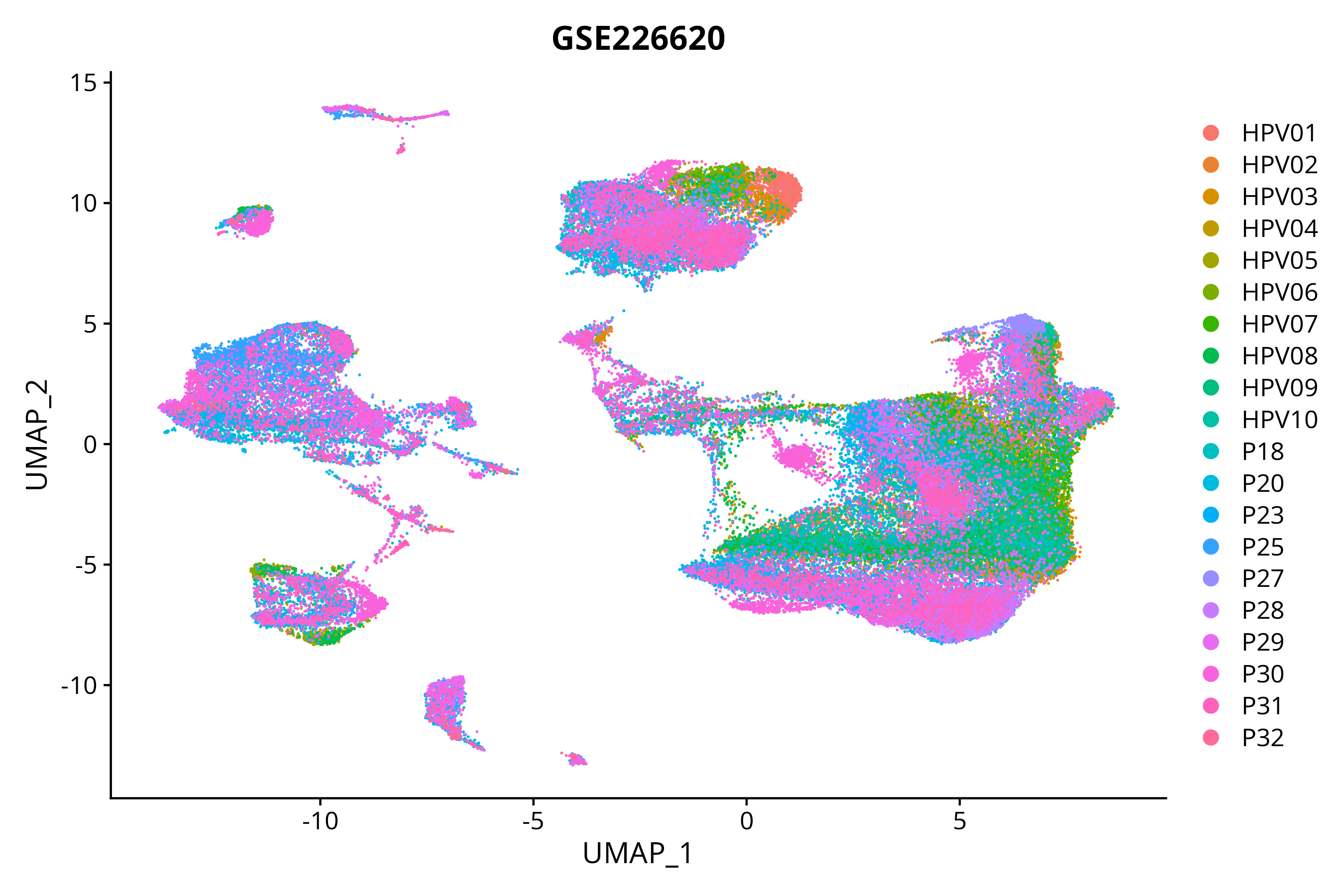
|
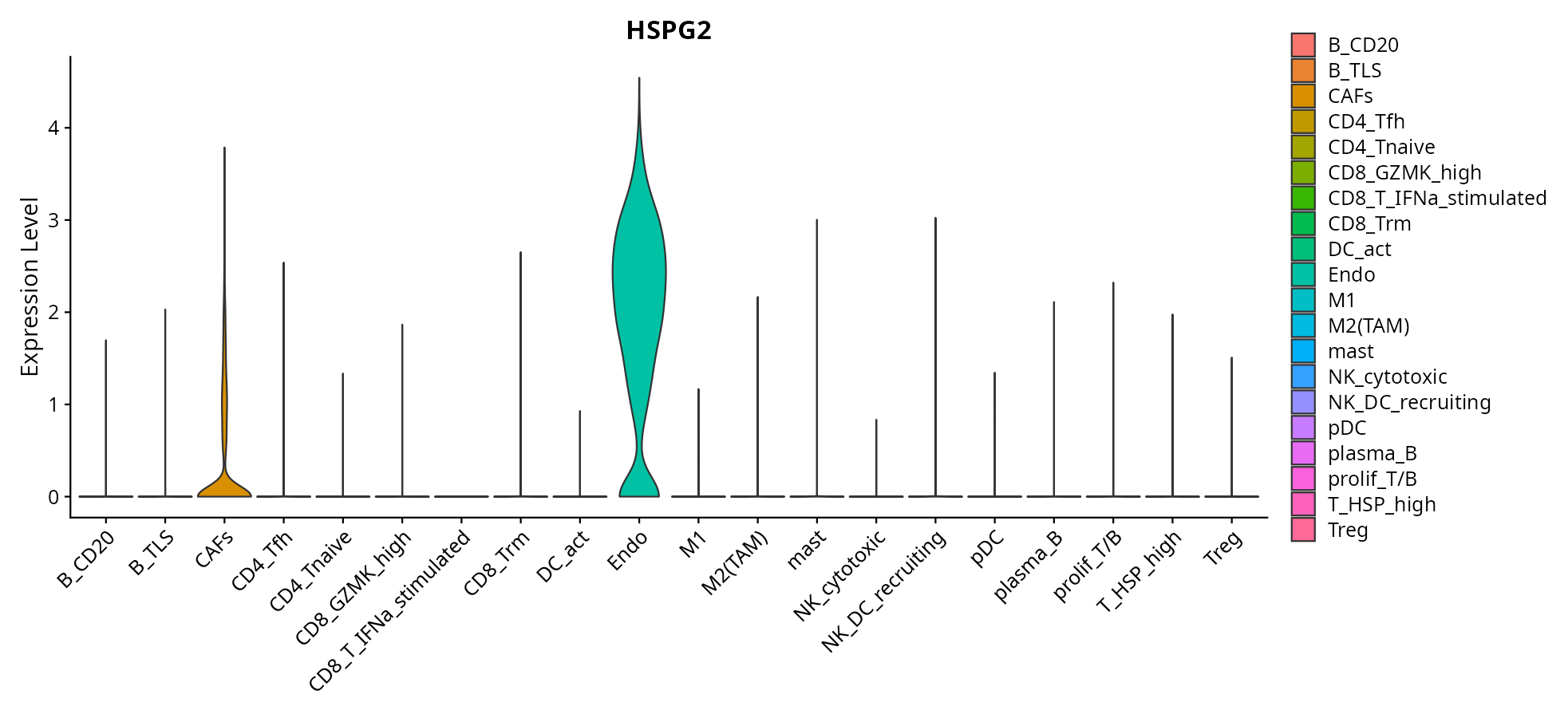
|
> Dataset: GSE164690 - Gene expression in cell subsets
|
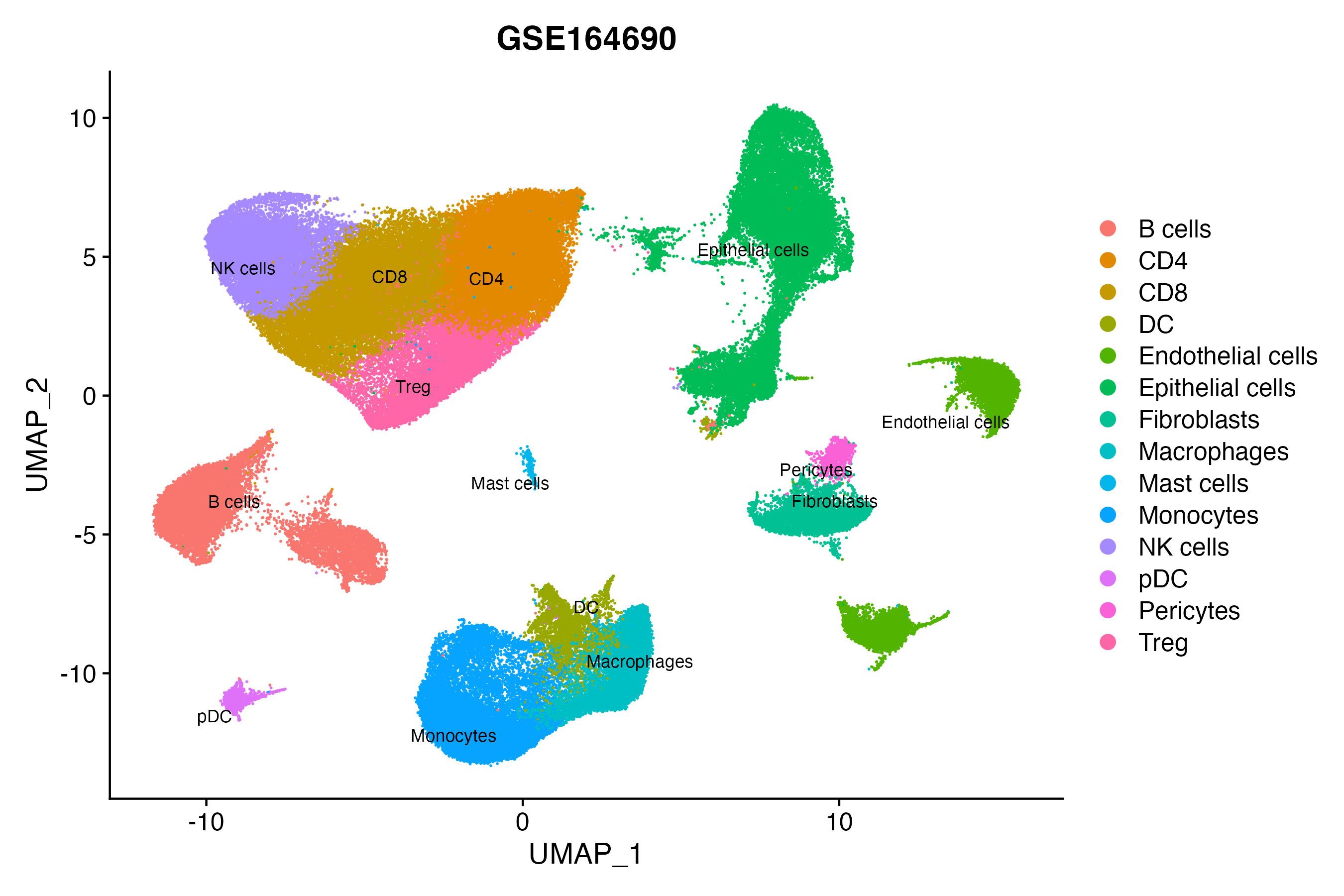
|
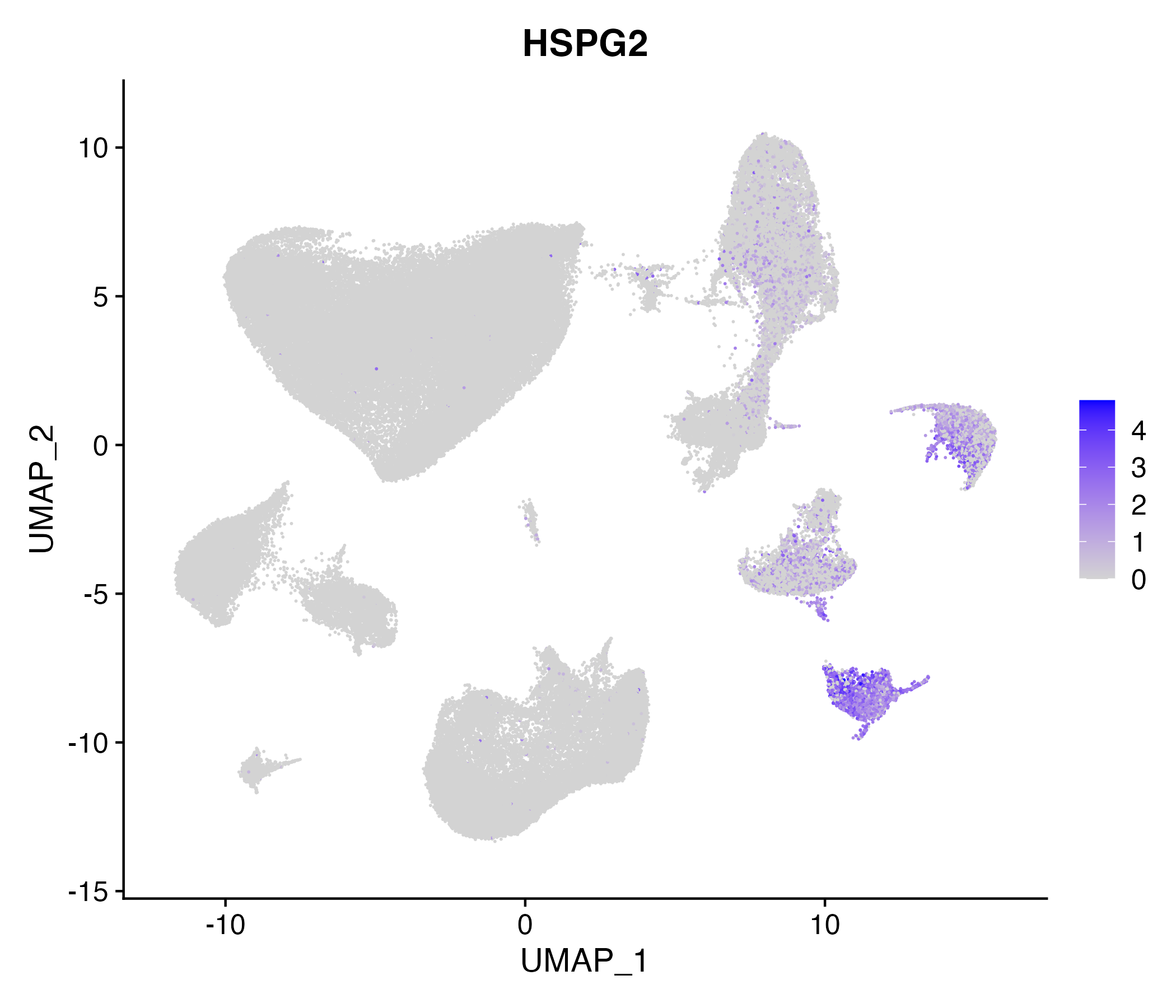
|

|
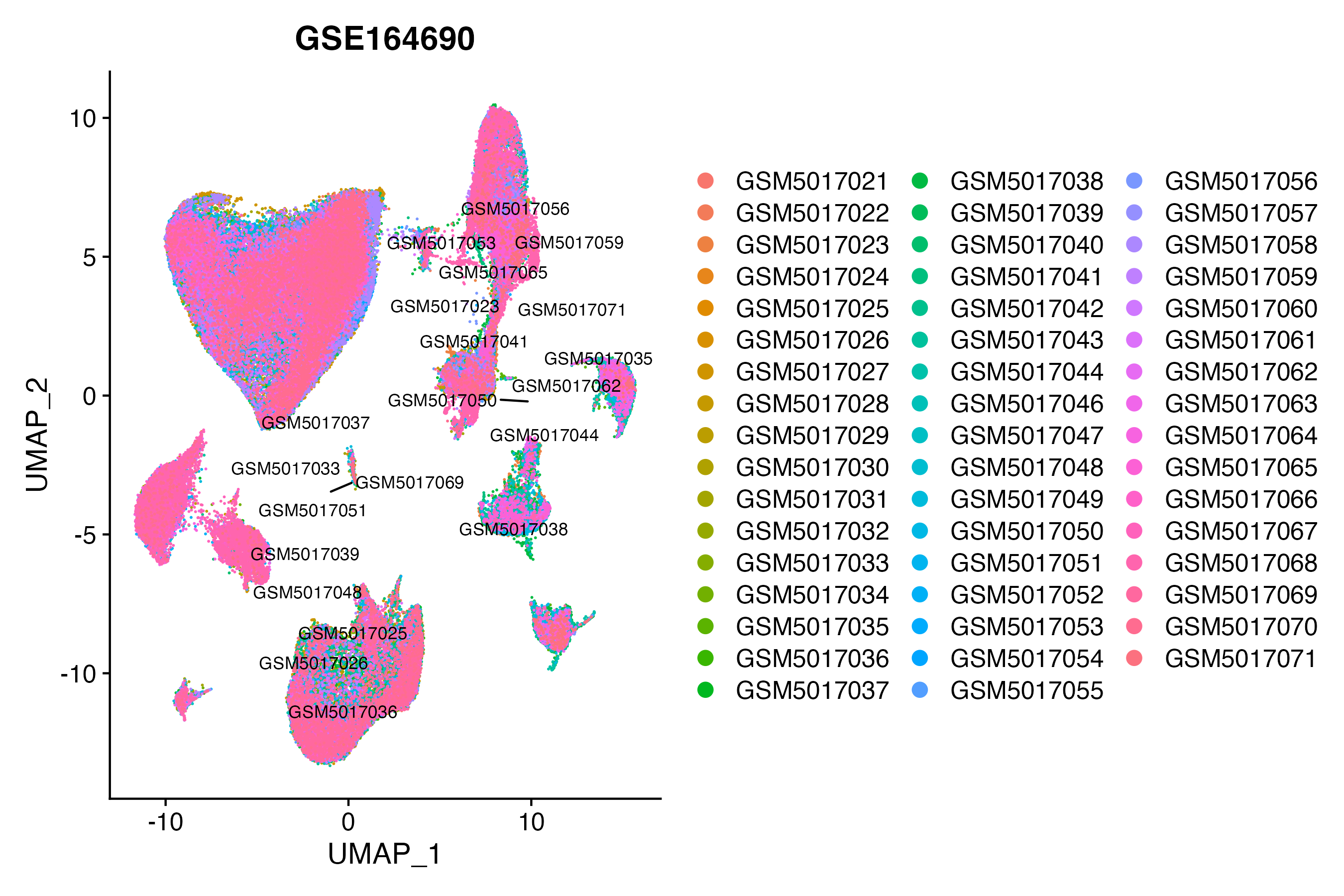
|
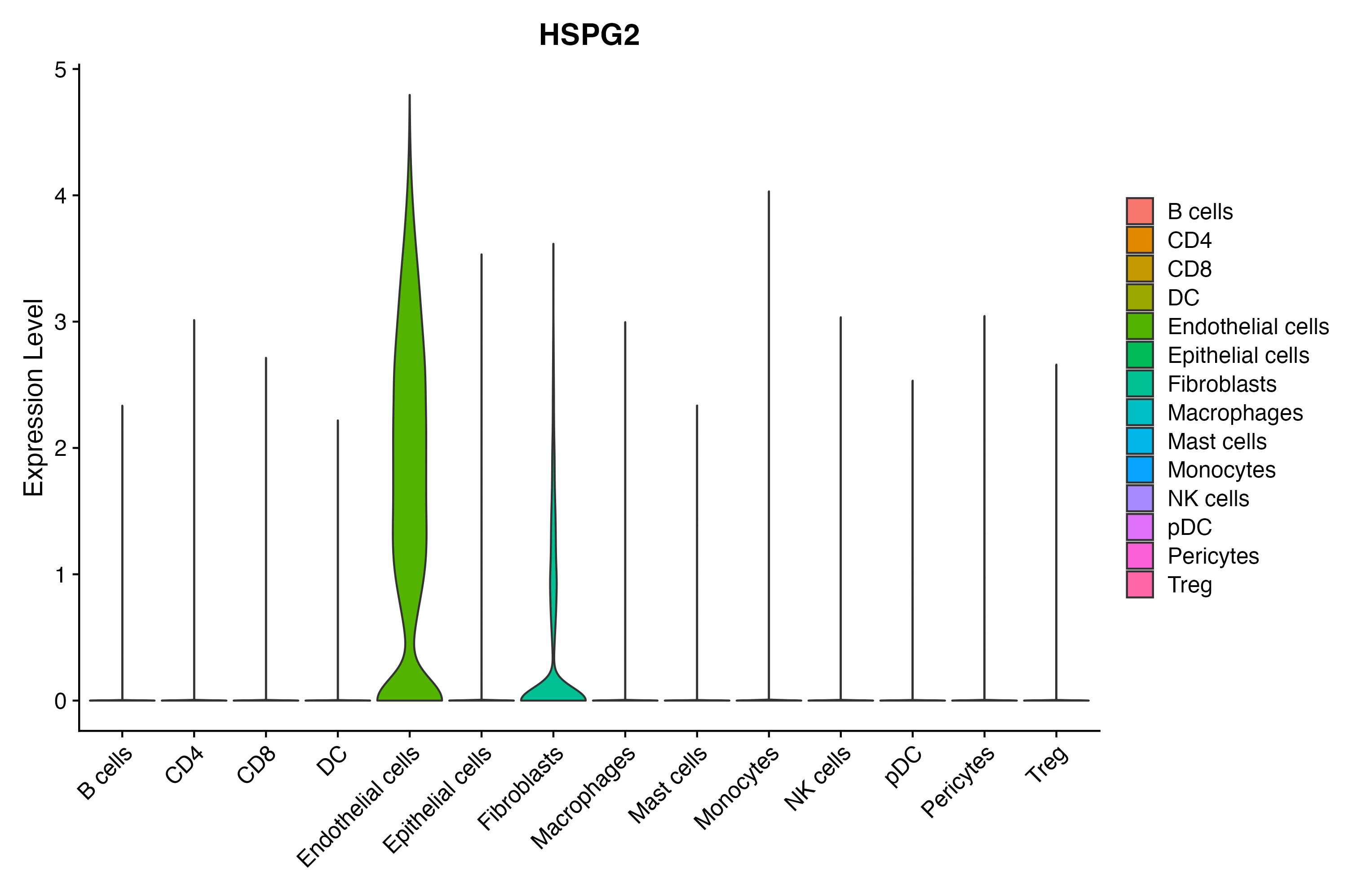
|
> Dataset: GSE197461 - Gene expression in cell subsets
|
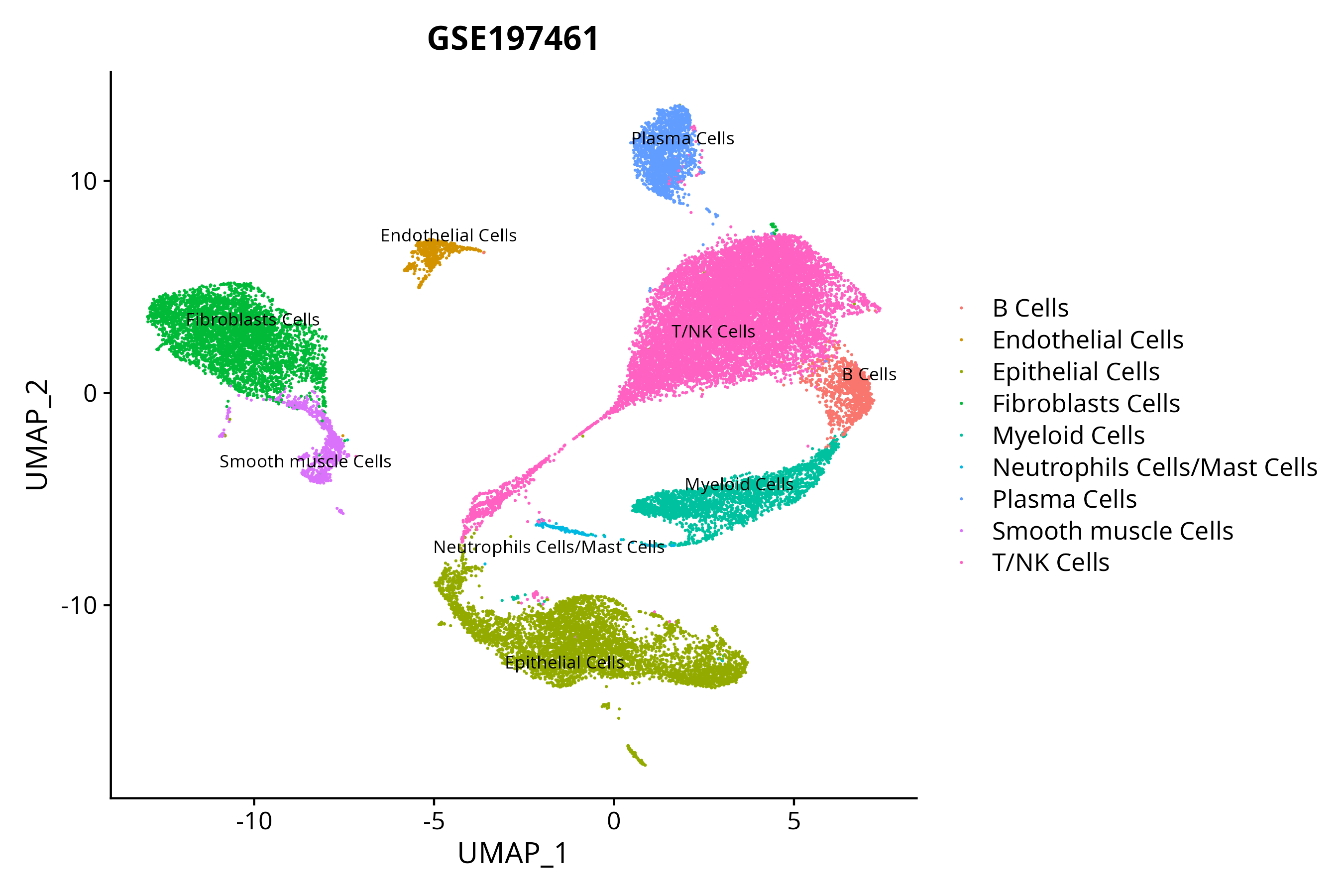
|
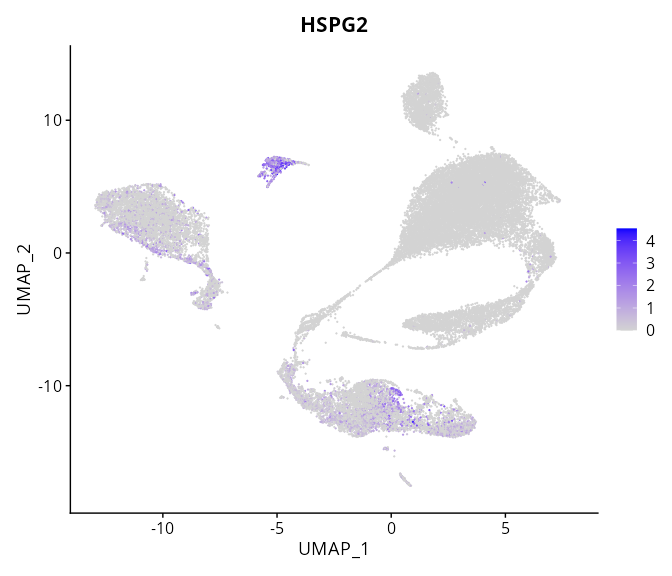
|
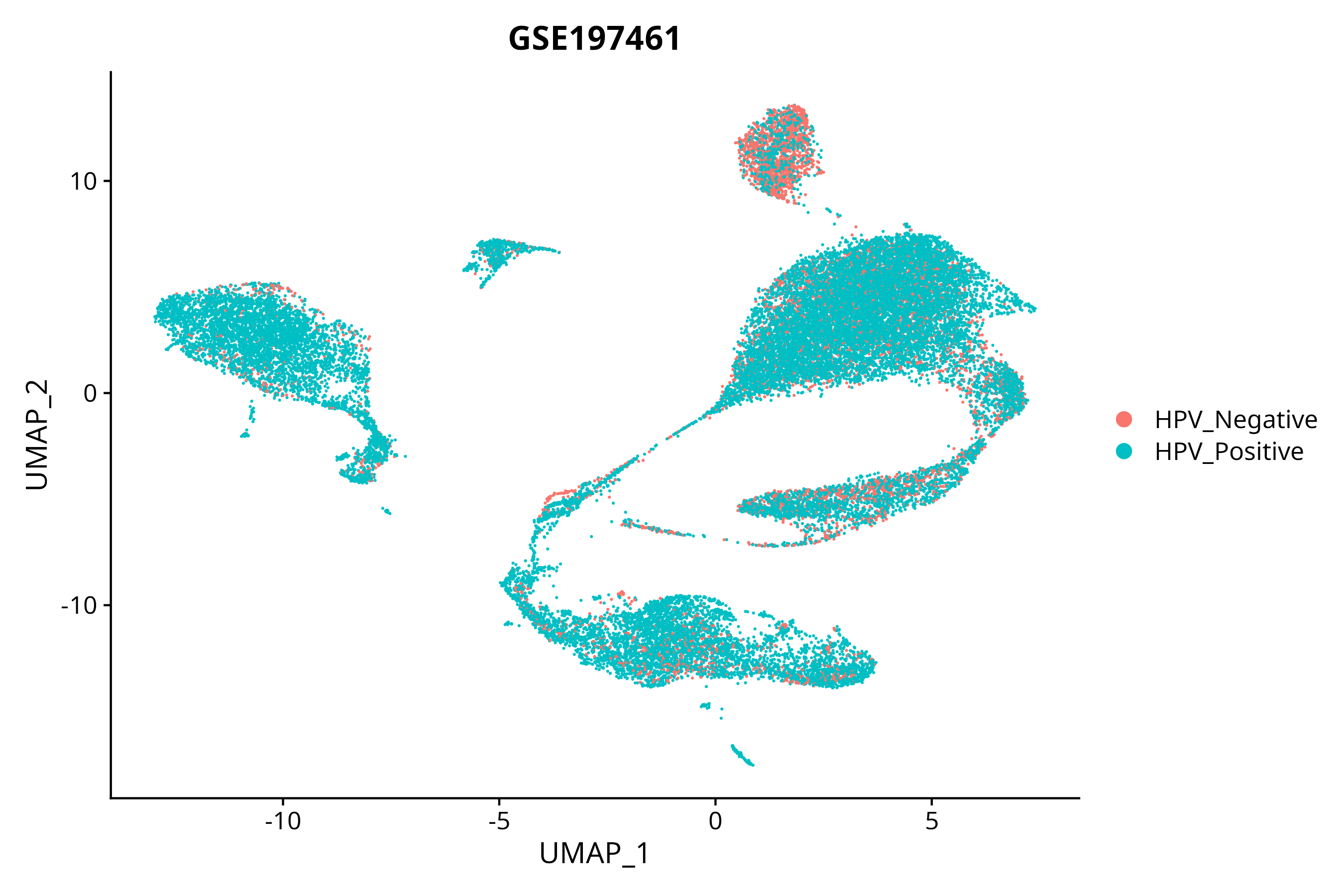
|
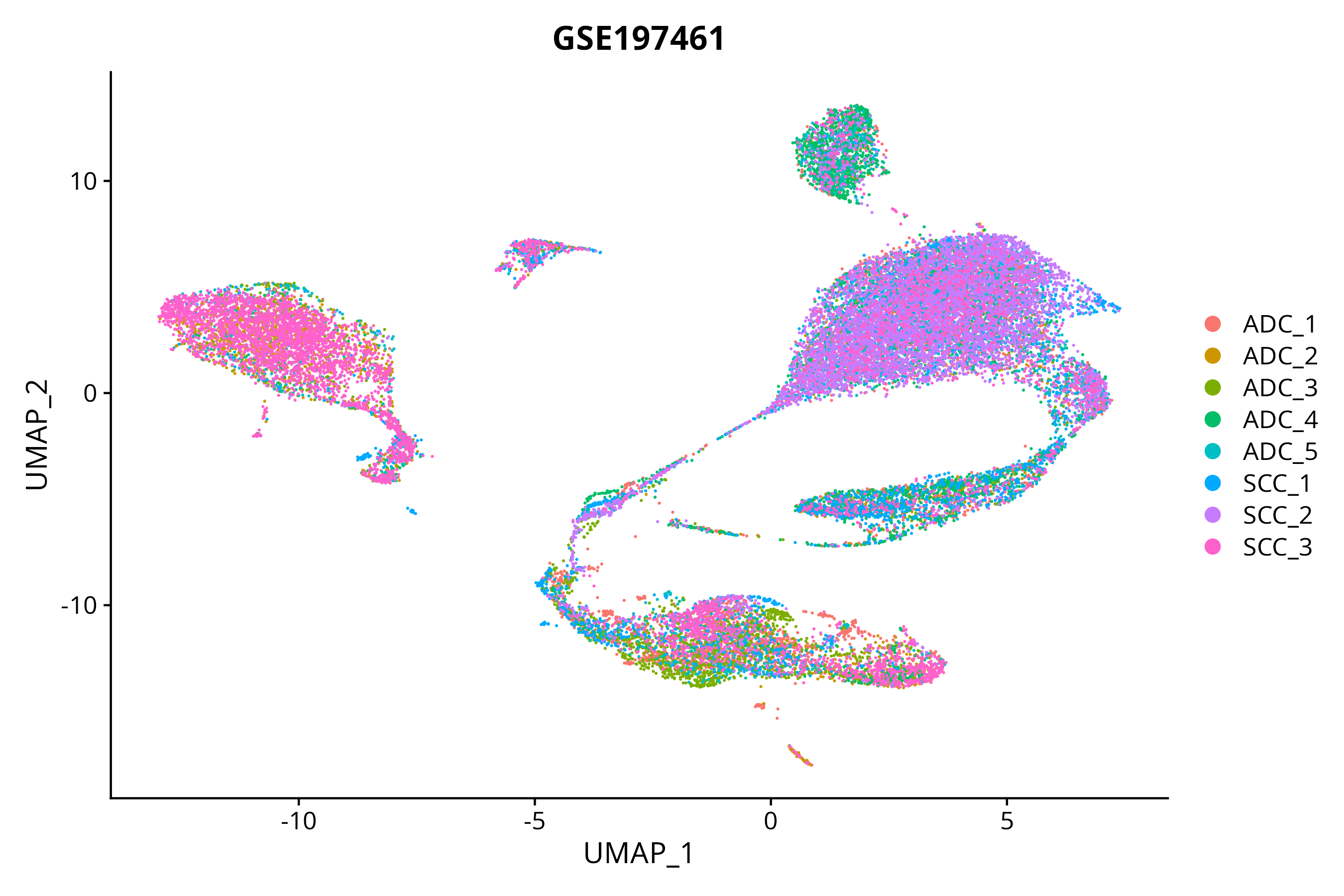
|
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information