Gene Information
|
Gene Name
|
PTPRK |
|
Gene ID
|
5796
|
|
Gene Full Name
|
protein tyrosine phosphatase receptor type K |
|
Gene Alias
|
R-PTP-kappa |
|
Transcripts
|
ENSG00000152894
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|
Cell Junctions;Vesicles, Plasma membrane, Connecting piece, Flagellar centriole; 
|
|
Membrane Info
|
Cancer-related genes, Enzymes, Plasma proteins, Predicted intracellular proteins, Predicted membrane proteins |
|
Uniport_ID
|
Q15262
|
|
HGNC ID
|
HGNC:9674
|
|
OMIM ID
|
602545 |
|
Summary
|
The protein encoded by this gene is a member of the protein tyrosine phosphatase (PTP) family. PTPs are known to be signaling molecules that regulate a variety of cellular processes including cell growth, differentiation, mitotic cycle, and oncogenic transformation. This PTP possesses an extracellular region, a single transmembrane region, and two tandem catalytic domains, and thus represents a receptor-type PTP. The extracellular region contains a meprin-A5 antigen-PTP mu (MAM) domain, an Ig-like domain and four fibronectin type III-like repeats. This PTP was shown to mediate homophilic intercellular interaction, possibly through the interaction with beta- and gamma-catenin at adherens junctions. Expression of this gene was found to be stimulated by TGF-beta 1, which may be important for the inhibition of keratinocyte proliferation. [provided by RefSeq, Jul 2008] |
Target gene [PTPRK] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5002085 |
chr6 |
7 |
9 |
0 |
16 |
View |
| 5004876 |
chr6 |
19 |
1 |
1 |
21 |
View |
| 5011235 |
chr6 |
3 |
1 |
2 |
6 |
View |
| 5011386 |
chr6 |
0 |
0 |
0 |
0 |
View |
| 5016164 |
chr6 |
19 |
12 |
1 |
32 |
View |
Target gene [PTPRK] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
GSE226620
|
scRNA-seq |
30 |
HiSeq X Ten (Homo sapiens) |
|
GSE40774
|
Expression array |
134 |
Agilent-026652 Whole Human Genome Microarray 4x44K v2 (Probe Name version) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE65858
|
Expression array |
270 |
Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE164690
|
scRNA-seq |
51 |
Illumina NextSeq 500 (Homo sapiens) |
|
GSE138080
|
Expression array |
35 |
Agilent-014850 Whole Human Genome Microarray 4x44K G4112F (Feature Number version) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
S GSE197461
|
scRNA-seq |
16 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
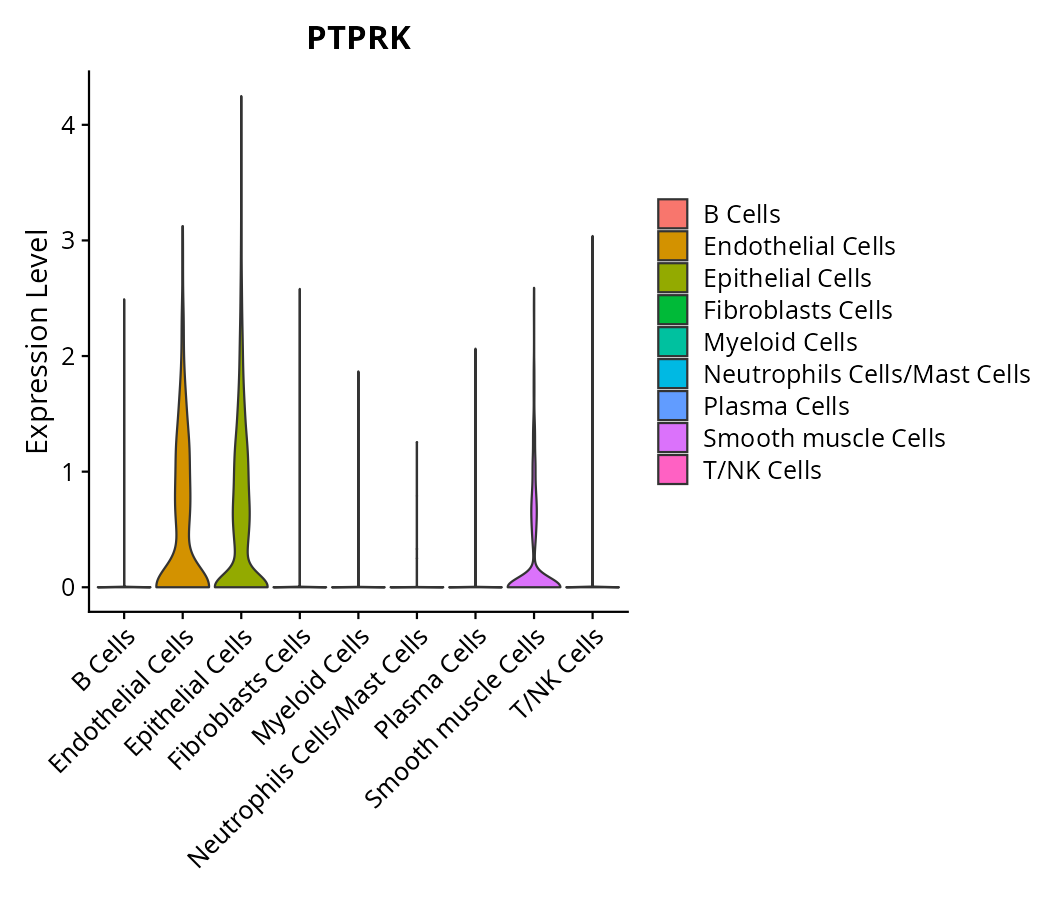
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE197461 - Gene expression in cell subsets
|
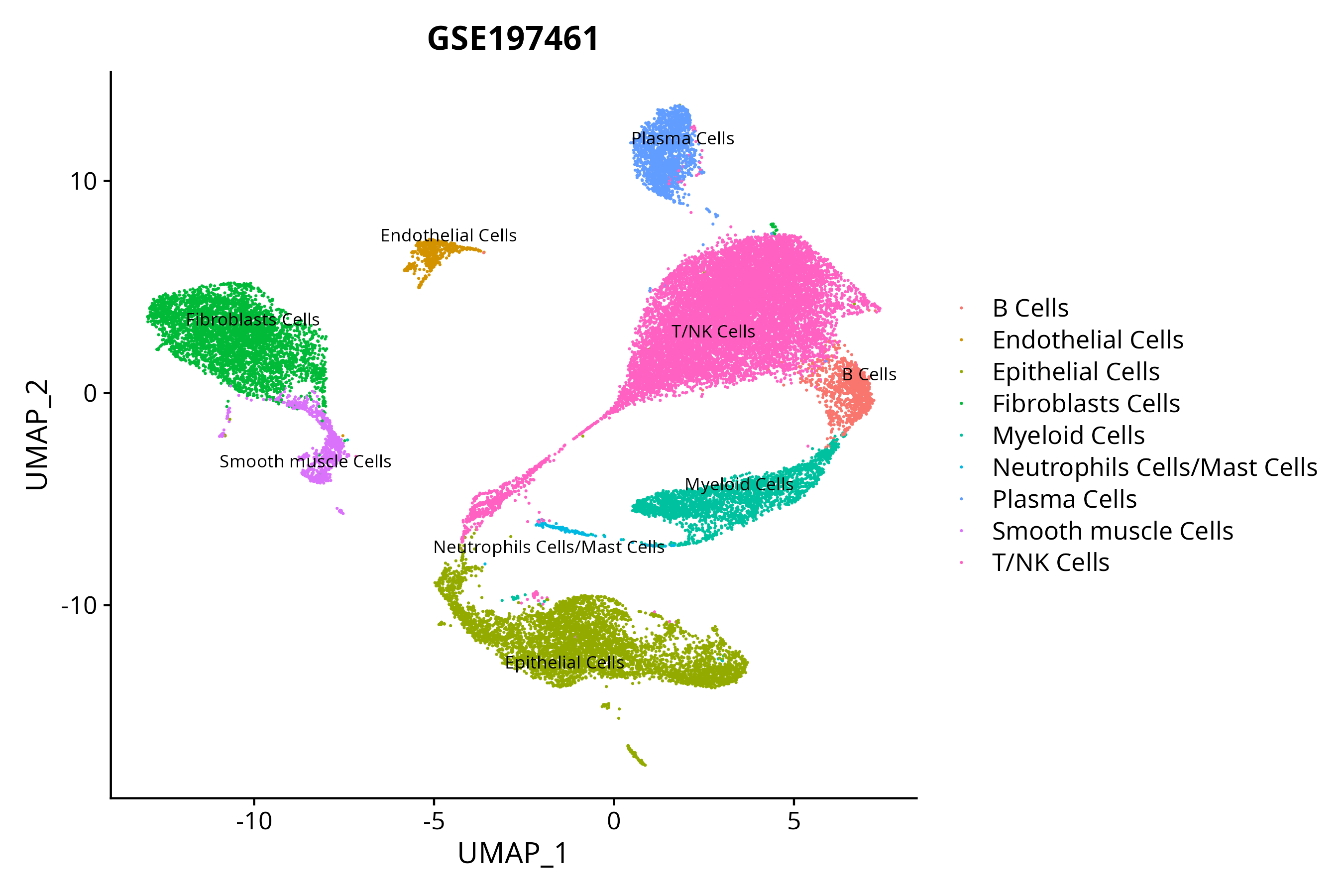
|
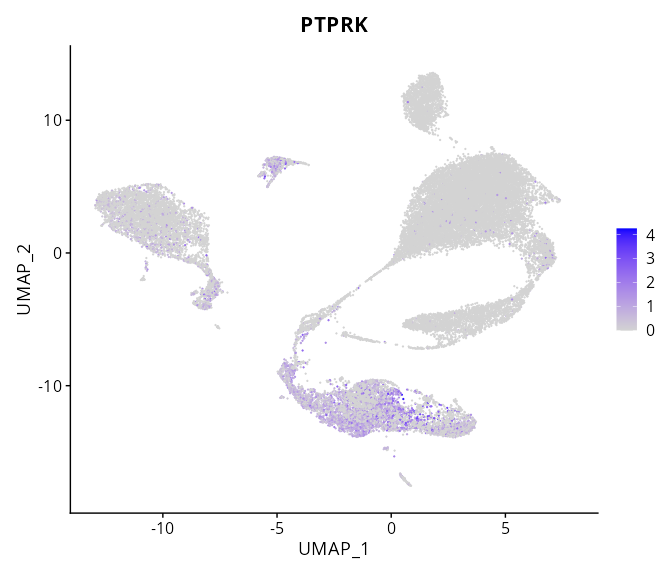
|
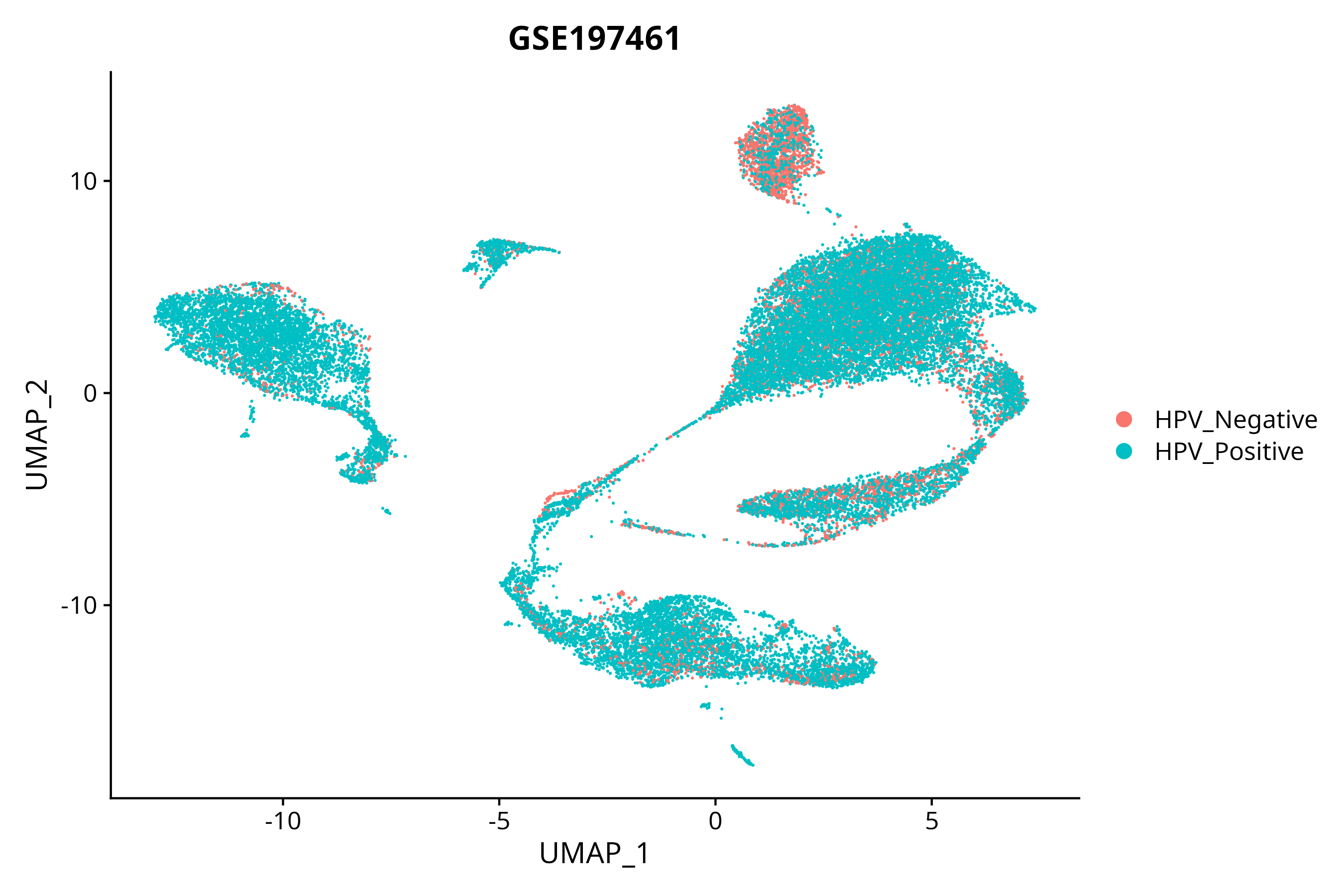
|
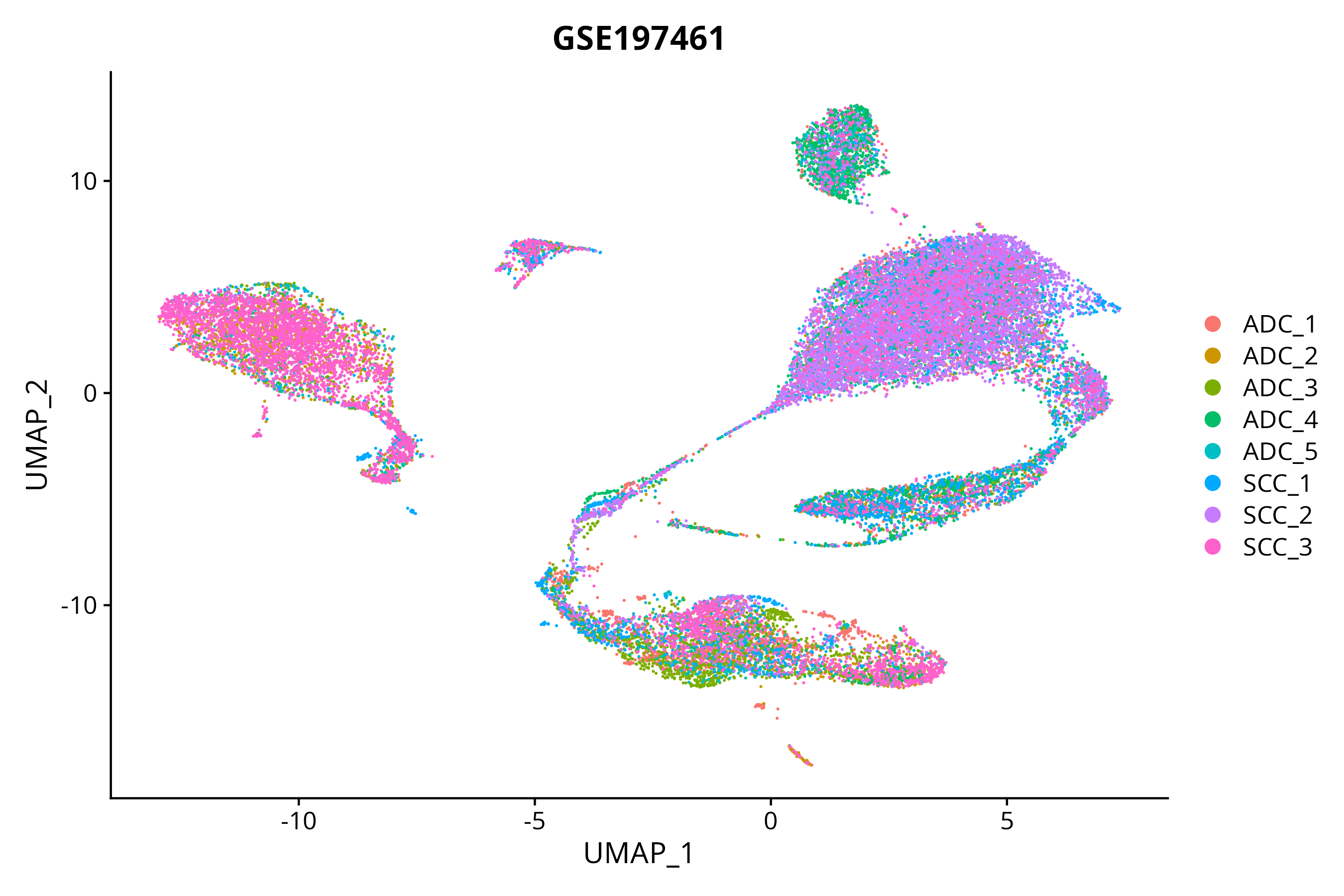
|
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information