GO Annotation
| Category | Term ID | Term description |
|---|---|---|
| Process | GO:0000122 | Negative regulation of transcription by RNA polymerase II |
| Process | GO:0001775 | Cell activation |
| Process | GO:0002376 | Immune system process |
| Process | GO:0006139 | Nucleobase-containing compound metabolic process |
| Process | GO:0006259 | DNA metabolic process |
| Process | GO:0006260 | DNA replication |
| Process | GO:0006261 | DNA-templated DNA replication |
| Process | GO:0006268 | DNA unwinding involved in DNA replication |
| Process | GO:0006355 | Regulation of transcription, DNA-templated |
| Process | GO:0006357 | Regulation of transcription by RNA polymerase II |
| Process | GO:0006417 | Regulation of translation |
| Process | GO:0006725 | Cellular aromatic compound metabolic process |
| Process | GO:0006807 | Nitrogen compound metabolic process |
| Process | GO:0006810 | Transport |
| Process | GO:0006996 | Organelle organization |
| Process | GO:0007017 | Microtubule-based process |
| Process | GO:0007018 | Microtubule-based movement |
| Process | GO:0007275 | Multicellular organism development |
| Process | GO:0007399 | Nervous system development |
| Process | GO:0008088 | Axo-dendritic transport |
| Process | GO:0008152 | Metabolic process |
| Process | GO:0008283 | Cell population proliferation |
| Process | GO:0008284 | Positive regulation of cell population proliferation |
| Process | GO:0009889 | Regulation of biosynthetic process |
| Process | GO:0009890 | Negative regulation of biosynthetic process |
| Process | GO:0009892 | Negative regulation of metabolic process |
| Process | GO:0009987 | Cellular process |
| Process | GO:0010468 | Regulation of gene expression |
| Process | GO:0010556 | Regulation of macromolecule biosynthetic process |
| Process | GO:0010558 | Negative regulation of macromolecule biosynthetic process |
| Process | GO:0010605 | Negative regulation of macromolecule metabolic process |
| Process | GO:0010608 | Post-transcriptional regulation of gene expression |
| Process | GO:0010629 | Negative regulation of gene expression |
| Process | GO:0010970 | Transport along microtubule |
| Process | GO:0016043 | Cellular component organization |
| Process | GO:0017148 | Negative regulation of translation |
| Process | GO:0019219 | Regulation of nucleobase-containing compound metabolic process |
| Process | GO:0019222 | Regulation of metabolic process |
| Process | GO:0030154 | Cell differentiation |
| Process | GO:0030705 | Cytoskeleton-dependent intracellular transport |
| Process | GO:0031323 | Regulation of cellular metabolic process |
| Process | GO:0031324 | Negative regulation of cellular metabolic process |
| Process | GO:0031326 | Regulation of cellular biosynthetic process |
| Process | GO:0031327 | Negative regulation of cellular biosynthetic process |
| Process | GO:0031503 | Protein-containing complex localization |
| Process | GO:0032392 | DNA geometric change |
| Process | GO:0032501 | Multicellular organismal process |
| Process | GO:0032502 | Developmental process |
| Process | GO:0032508 | DNA duplex unwinding |
| Process | GO:0032943 | Mononuclear cell proliferation |
| Process | GO:0034248 | Regulation of cellular amide metabolic process |
| Process | GO:0034249 | Negative regulation of cellular amide metabolic process |
| Process | GO:0034641 | Cellular nitrogen compound metabolic process |
| Process | GO:0042127 | Regulation of cell population proliferation |
| Process | GO:0043170 | Macromolecule metabolic process |
| Process | GO:0043433 | Negative regulation of DNA-binding transcription factor activity |
| Process | GO:0044092 | Negative regulation of molecular function |
| Process | GO:0044237 | Cellular metabolic process |
| Process | GO:0044238 | Primary metabolic process |
| Process | GO:0044260 | Cellular macromolecule metabolic process |
| Process | GO:0045321 | Leukocyte activation |
| Process | GO:0045892 | Negative regulation of transcription, DNA-templated |
| Process | GO:0045934 | Negative regulation of nucleobase-containing compound metabolic process |
| Process | GO:0046483 | Heterocycle metabolic process |
| Process | GO:0046649 | Lymphocyte activation |
| Process | GO:0046651 | Lymphocyte proliferation |
| Process | GO:0046907 | Intracellular transport |
| Process | GO:0048518 | Positive regulation of biological process |
| Process | GO:0048519 | Negative regulation of biological process |
| Process | GO:0048522 | Positive regulation of cellular process |
| Process | GO:0048523 | Negative regulation of cellular process |
| Process | GO:0048731 | System development |
| Process | GO:0048856 | Anatomical structure development |
| Process | GO:0048869 | Cellular developmental process |
| Process | GO:0050673 | Epithelial cell proliferation |
| Process | GO:0050789 | Regulation of biological process |
| Process | GO:0050794 | Regulation of cellular process |
| Process | GO:0051090 | Regulation of DNA-binding transcription factor activity |
| Process | GO:0051171 | Regulation of nitrogen compound metabolic process |
| Process | GO:0051172 | Negative regulation of nitrogen compound metabolic process |
| Process | GO:0051179 | Localization |
| Process | GO:0051234 | Establishment of localization |
| Process | GO:0051246 | Regulation of protein metabolic process |
| Process | GO:0051248 | Negative regulation of protein metabolic process |
| Process | GO:0051252 | Regulation of RNA metabolic process |
| Process | GO:0051253 | Negative regulation of RNA metabolic process |
| Process | GO:0051276 | Chromosome organization |
| Process | GO:0051641 | Cellular localization |
| Process | GO:0051649 | Establishment of localization in cell |
| Process | GO:0060255 | Regulation of macromolecule metabolic process |
| Process | GO:0065007 | Biological regulation |
| Process | GO:0065009 | Regulation of molecular function |
| Process | GO:0070661 | Leukocyte proliferation |
| Process | GO:0071103 | DNA conformation change |
| Process | GO:0071704 | Organic substance metabolic process |
| Process | GO:0071840 | Cellular component organization or biogenesis |
| Process | GO:0080090 | Regulation of primary metabolic process |
| Process | GO:0090304 | Nucleic acid metabolic process |
| Process | GO:0098935 | Dendritic transport |
| Process | GO:0098961 | Dendritic transport of ribonucleoprotein complex |
| Process | GO:0098963 | Dendritic transport of messenger ribonucleoprotein complex |
| Process | GO:0099111 | Microtubule-based transport |
| Process | GO:1901360 | Organic cyclic compound metabolic process |
| Process | GO:1902679 | Negative regulation of RNA biosynthetic process |
| Process | GO:1903506 | Regulation of nucleic acid-templated transcription |
| Process | GO:1903507 | Negative regulation of nucleic acid-templated transcription |
| Process | GO:2000112 | Regulation of cellular macromolecule biosynthetic process |
| Process | GO:2000113 | Negative regulation of cellular macromolecule biosynthetic process |
| Process | GO:2001141 | Regulation of RNA biosynthetic process |
| Component | GO:0005622 | Intracellular anatomical structure |
| Component | GO:0005634 | Nucleus |
| Component | GO:0005737 | Cytoplasm |
| Component | GO:0014069 | Postsynaptic density |
| Component | GO:0030054 | Cell junction |
| Component | GO:0030425 | Dendrite |
| Component | GO:0032279 | Asymmetric synapse |
| Component | GO:0032838 | Plasma membrane bounded cell projection cytoplasm |
| Component | GO:0032839 | Dendrite cytoplasm |
| Component | GO:0036477 | Somatodendritic compartment |
| Component | GO:0042995 | Cell projection |
| Component | GO:0043005 | Neuron projection |
| Component | GO:0043025 | Neuronal cell body |
| Component | GO:0043226 | Organelle |
| Component | GO:0043227 | Membrane-bounded organelle |
| Component | GO:0043229 | Intracellular organelle |
| Component | GO:0043231 | Intracellular membrane-bounded organelle |
| Component | GO:0044297 | Cell body |
| Component | GO:0045202 | Synapse |
| Component | GO:0097447 | Dendritic tree |
| Component | GO:0098794 | Postsynapse |
| Component | GO:0098978 | Glutamatergic synapse |
| Component | GO:0098984 | Neuron to neuron synapse |
| Component | GO:0099568 | Cytoplasmic region |
| Component | GO:0099572 | Postsynaptic specialization |
| Component | GO:0110165 | Cellular anatomical entity |
| Component | GO:0120025 | Plasma membrane bounded cell projection |
| Component | GO:0120111 | Neuron projection cytoplasm |
| Function | GO:0000900 | mRNA regulatory element binding translation repressor activity |
| Function | GO:0000976 | Transcription cis-regulatory region binding |
| Function | GO:0000977 | RNA polymerase II transcription regulatory region sequence-specific DNA binding |
| Function | GO:0000981 | DNA-binding transcription factor activity, RNA polymerase II-specific |
| Function | GO:0001067 | Transcription regulatory region nucleic acid binding |
| Function | GO:0003676 | Nucleic acid binding |
| Function | GO:0003677 | DNA binding |
| Function | GO:0003690 | Double-stranded DNA binding |
| Function | GO:0003691 | Double-stranded telomeric DNA binding |
| Function | GO:0003697 | Single-stranded DNA binding |
| Function | GO:0003700 | DNA-binding transcription factor activity |
| Function | GO:0003723 | RNA binding |
| Function | GO:0005488 | Binding |
| Function | GO:0005515 | Protein binding |
| Function | GO:0008134 | Transcription factor binding |
| Function | GO:0030371 | Translation repressor activity |
| Function | GO:0032422 | Purine-rich negative regulatory element binding |
| Function | GO:0042162 | Telomeric DNA binding |
| Function | GO:0043565 | Sequence-specific DNA binding |
| Function | GO:0045182 | Translation regulator activity |
| Function | GO:0046332 | SMAD binding |
| Function | GO:0090079 | Translation regulator activity, nucleic acid binding |
| Function | GO:0097159 | Organic cyclic compound binding |
| Function | GO:0098772 | Molecular function regulator activity |
| Function | GO:0140110 | Transcription regulator activity |
| Function | GO:0140297 | DNA-binding transcription factor binding |
| Function | GO:0140416 | Transcription regulator inhibitor activity |
| Function | GO:0140678 | Molecular function inhibitor activity |
| Function | GO:1901363 | Heterocyclic compound binding |
| Function | GO:1990837 | Sequence-specific double-stranded DNA binding |
Pathway annotation
| Category | Term ID | Term description |
|---|---|---|
| WikiPathways | WP5242 | Comprehensive IL-17A signaling |
| WikiPathways | WP544 | Exercise-induced circadian regulation |
Expression profile of Pura in omics data
| ID | GSE106799 |
| Title | Propofol alters the mRNA profile in the developing mouse hippocampus |
| Organism | Mus musculus |
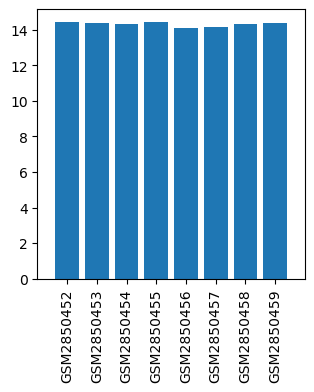
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2850452 | Control 1 | tissue: hippocampus age: 7 days-old | 14.442515 |
| GSM2850453 | Control 2 | tissue: hippocampus age: 7 days-old | 14.385873 |
| GSM2850454 | Control 3 | tissue: hippocampus age: 7 days-old | 14.339093 |
| GSM2850455 | Control 4 | tissue: hippocampus age: 7 days-old | 14.425244 |
| GSM2850456 | Propofol 1 | tissue: hippocampus age: 7 days-old | 14.101999 |
| GSM2850457 | Propofol 2 | tissue: hippocampus age: 7 days-old | 14.182906 |
| GSM2850458 | Propofol 3 | tissue: hippocampus age: 7 days-old | 14.315009 |
| GSM2850459 | Propofol 4 | tissue: hippocampus age: 7 days-old | 14.370253 |
| ID | GSE109165 |
| Title | Suppression of RGSz1 function optimizes the actions of opioid analgesics by mechanisms that involve the Wnt/β-catenin pathway |
| Organism | Mus musculus |
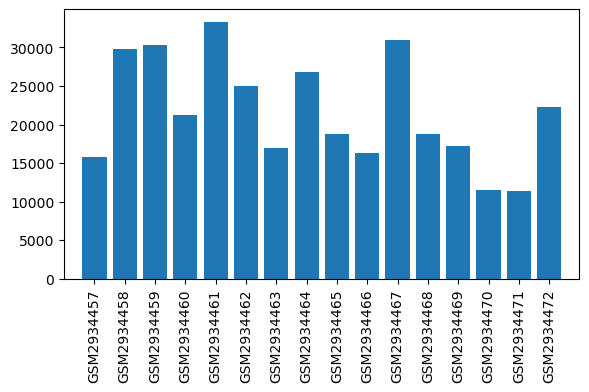
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM2934457 | WT_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 15744.0 |
| GSM2934458 | WT_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 29813.0 |
| GSM2934459 | WT_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 30301.0 |
| GSM2934460 | WT_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: naïve | 21201.0 |
| GSM2934461 | KO_Naive_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 33308.0 |
| GSM2934462 | KO_Naive_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 24962.0 |
| GSM2934463 | KO_Naive_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 16967.0 |
| GSM2934464 | KO_Naive_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: naïve | 26778.0 |
| GSM2934465 | WT_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 18709.0 |
| GSM2934466 | WT_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 16321.0 |
| GSM2934467 | WT_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 30997.0 |
| GSM2934468 | WT_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: wildtype treatment: morphine | 18752.0 |
| GSM2934469 | KO_Morphine_1 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 17242.0 |
| GSM2934470 | KO_Morphine_2 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 11441.0 |
| GSM2934471 | KO_Morphine_3 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 11434.0 |
| GSM2934472 | KO_Morphine_4 | tissue: brain brain region: Periaqueductal gray region Sex: Male genotype: RGSz1 KO treatment: morphine | 22309.0 |
| ID | GSE125076 |
| Title | A mouse model of acute post-surgical pain |
| Organism | Mus musculus |
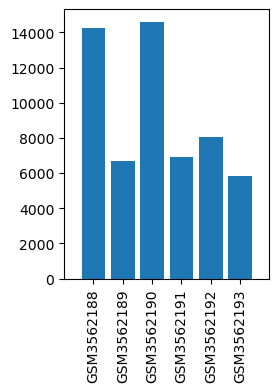
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM3562188 | DRG_WT_sham_1 | strain: C57BL/6 gender: male age: 8 weeks treatment: naïve | 14276.0 |
| GSM3562189 | DRG_WT_sham_2 | strain: C57BL/6 gender: male age: 8 weeks treatment: naïve | 6664.0 |
| GSM3562190 | DRG_WT_sham_3 | strain: C57BL/6 gender: male age: 8 weeks treatment: naïve | 14593.0 |
| GSM3562191 | DRG_WT_PSP_1 | strain: C57BL/6 gender: male age: 8 weeks treatment: 24h post surgery | 6918.0 |
| GSM3562192 | DRG_WT_PSP_2 | strain: C57BL/6 gender: male age: 8 weeks treatment: 24h post surgery | 8077.0 |
| GSM3562193 | DRG_WT_PSP_3 | strain: C57BL/6 gender: male age: 8 weeks treatment: 24h post surgery | 5825.0 |
| ID | GSE126662 |
| Title | Comparison between opioid (morphine) and vehicle treatments in mouse trigeminal ganglia and nucleus accumbens |
| Organism | Mus musculus |
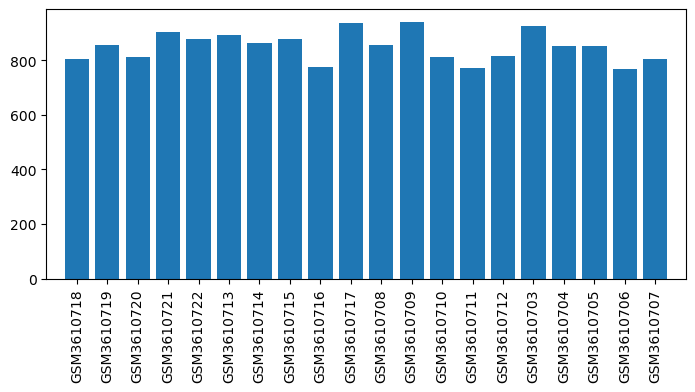
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM3610703 | TG_M1: OIH_M1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 926.584 |
| GSM3610704 | TG_M2: OIH_M2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 852.985 |
| GSM3610705 | TG_M3: OIH_M3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 851.3849999999999 |
| GSM3610706 | TG_M4: OIH_M4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 769.3836666666666 |
| GSM3610707 | TG_M5: OIH_M5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 806.2316666666666 |
| GSM3610708 | TG_V1: VehicleOIH_V1_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 856.3416666666666 |
| GSM3610709 | TG_V2: VehicleOIH_V2_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 940.3983333333332 |
| GSM3610710 | TG_V3: VehicleOIH_V3_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 810.777 |
| GSM3610711 | TG_V4: VehicleOIH_V4_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 770.5210000000001 |
| GSM3610712 | TG_V5: VehicleOIH_V5_Trigeminal_Ganglia | tissue: Trigeminal ganglia strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 816.5390000000001 |
| GSM3610713 | NAc_M1: OIH_M1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 892.5676666666667 |
| GSM3610714 | NAc_M2: OIH_M2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 861.7633333333333 |
| GSM3610715 | NAc_M3: OIH_M3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 879.3176666666667 |
| GSM3610716 | NAc_M4: OIH_M4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 776.3253333333332 |
| GSM3610717 | NAc_M5: OIH_M5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: OIH age: 9-12 weeks old Sex: male | 937.9780000000001 |
| GSM3610718 | NAc_V1: VehicleOIH_V1_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 806.2316666666667 |
| GSM3610719 | NAc_V2: VehicleOIH_V2_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 854.6790000000001 |
| GSM3610720 | NAc_V3: VehicleOIH_V3_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 813.0526666666666 |
| GSM3610721 | NAc_V4: VehicleOIH_V4_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 904.0423333333334 |
| GSM3610722 | NAc_V5: VehicleOIH_V5_Nucleus_Accumbens | tissue: Nucleus accumbens strain: C57BL/6J treatment: Vehicle age: 9-12 weeks old Sex: male | 879.5693333333334 |
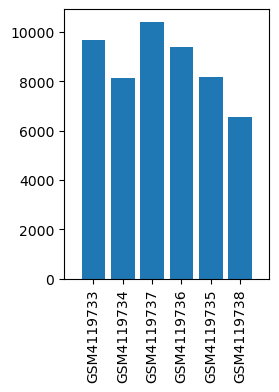
| ID | GSE138802 |
| Title | Mesocorticolimbic circuit mechanisms underlying the effects of ketamine on dopamine: a translational imaging study |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM4119733 | PLc_1_Sal | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Saline | 9684.43 |
| GSM4119734 | PLc_10_Sal | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Saline | 8138.34 |
| GSM4119735 | PLc_6_Sal | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Saline | 8178.28 |
| GSM4119736 | PLc_5_Ket | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Ketamine | 9384.06 |
| GSM4119737 | PLc_3_Ket | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Ketamine | 10405.32 |
| GSM4119738 | PLc_9_Ket | strain: C57Bl/6 age: 9 weeks tissue: Pre-limbic cortex treatment: Ketamine | 6569.85 |
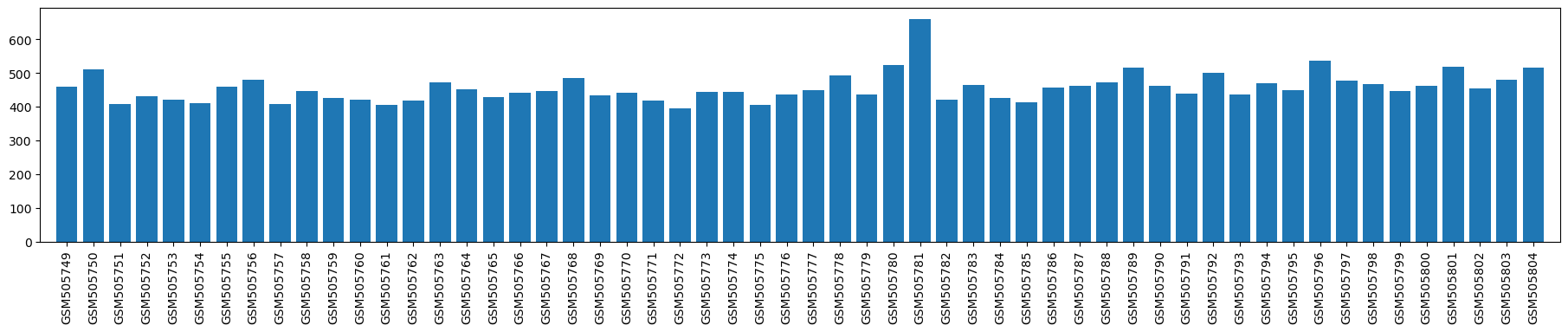
| ID | GSE20160 |
| Title | Gene Expression Data from Prefrontal Cortex and Nucleus Accumbens from Inbred Strains of Mice |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM505749 | 129S1/SvImJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: 129S1/SvImJ | 459.241 |
| GSM505750 | A/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: A/J | 511.5258333333333 |
| GSM505751 | AKR/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: AKR/J | 408.1285 |
| GSM505752 | BALB/cByJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: BALB/cByJ | 431.8443333333333 |
| GSM505753 | BTBR_T+_tf/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: BTBR_T+_tf/J | 420.1191666666667 |
| GSM505754 | BUB/BnJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: BUB/BnJ | 409.5461666666667 |
| GSM505755 | C3H/HeJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C3H/HeJ | 458.397 |
| GSM505756 | C57BL/6J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C57BL/6J | 480.15599999999995 |
| GSM505757 | C57BR/cdJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C57BR/cdJ | 407.4531666666667 |
| GSM505758 | C58/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: C58/J | 445.2293333333334 |
| GSM505759 | CBA/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: CBA/J | 426.44883333333337 |
| GSM505760 | CE/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: CE/J | 421.7148333333333 |
| GSM505761 | DBA/2J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: DBA/2J | 404.19066666666663 |
| GSM505762 | FVB/NJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: FVB/NJ | 417.253 |
| GSM505763 | I/LnJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: I/LnJ | 471.52433333333335 |
| GSM505764 | KK/HlJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: KK/HlJ | 452.032 |
| GSM505765 | MA/MyJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: MA/MyJ | 427.53133333333335 |
| GSM505766 | MRL_MpJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: MRL/MpJ | 441.1376666666667 |
| GSM505767 | NOD/LtJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NOD/LtJ | 445.022 |
| GSM505768 | NON/LtJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NON/LtJ | 485.8258333333333 |
| GSM505769 | NZO/HILtJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NZO/HILtJ | 433.15366666666665 |
| GSM505770 | NZW_LacJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: NZW_LacJ | 441.55216666666666 |
| GSM505771 | P/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: P/J | 418.5613333333333 |
| GSM505772 | PL/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: PL/J | 394.29033333333336 |
| GSM505773 | RIIIS/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: RIIIS/J | 444.4086666666667 |
| GSM505774 | SJL/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: SJL/J | 442.9026666666667 |
| GSM505775 | SM/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: SM/J | 404.647 |
| GSM505776 | SWR/J_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: SWR/J | 436.9738333333333 |
| GSM505777 | WSB/EiJ_Male_Frontal_Cortex | gender: male tissue: Frontal Cortex strain: WSB/EiJ | 448.05233333333337 |
| GSM505778 | 129S1/SvImJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: 129S1/SvImJ | 491.5363333333333 |
| GSM505779 | A/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: A/J | 435.54266666666666 |
| GSM505780 | AKR/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: AKR/J | 524.0063333333334 |
| GSM505781 | BALB/cByJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: BALB/cByJ | 660.0248333333333 |
| GSM505782 | BTBR_T+_tf/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: BTBR_T+_tf/J | 421.4148333333333 |
| GSM505783 | BUB/BnJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: BUB/BnJ | 462.9563333333333 |
| GSM505784 | C3H/HeJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C3H/HeJ | 426.57116666666667 |
| GSM505785 | C57BL/6J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C57BL/6J | 411.59133333333335 |
| GSM505786 | C57BR/cdJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C57BR/cdJ | 457.4336666666666 |
| GSM505787 | C58/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: C58/J | 462.21316666666667 |
| GSM505788 | CBA/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: CBA/J | 471.70533333333333 |
| GSM505789 | CE/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: CE/J | 516.4603333333333 |
| GSM505790 | DBA/2J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: DBA/2J | 461.31166666666667 |
| GSM505791 | FVB/NJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: FVB/NJ | 437.30299999999994 |
| GSM505792 | I/LnJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: I/LnJ | 498.9745 |
| GSM505793 | KK/HlJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: KK/HlJ | 435.46250000000003 |
| GSM505794 | MA/MyJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: MA/MyJ | 469.14433333333335 |
| GSM505795 | NOD/LtJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: NOD/LtJ | 450.03 |
| GSM505796 | NON/LtJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: NON/LtJ | 537.4658333333333 |
| GSM505797 | NZW/LacJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: NZW/LacJ | 476.907 |
| GSM505798 | P/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: P/J | 466.9166666666667 |
| GSM505799 | PL/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: PL/J | 445.2656666666667 |
| GSM505800 | RIIIS/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: RIIIS/J | 460.55749999999995 |
| GSM505801 | SJL/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: SJL/J | 519.2246666666666 |
| GSM505802 | SM/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: SM/J | 453.26516666666663 |
| GSM505803 | SWR/J_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: SWR/J | 479.9998333333333 |
| GSM505804 | WSB/EiJ_Male_Nuc_Acc | gender: male tissue: Nucleus Accumbens strain: WSB/EiJ | 516.7501666666667 |
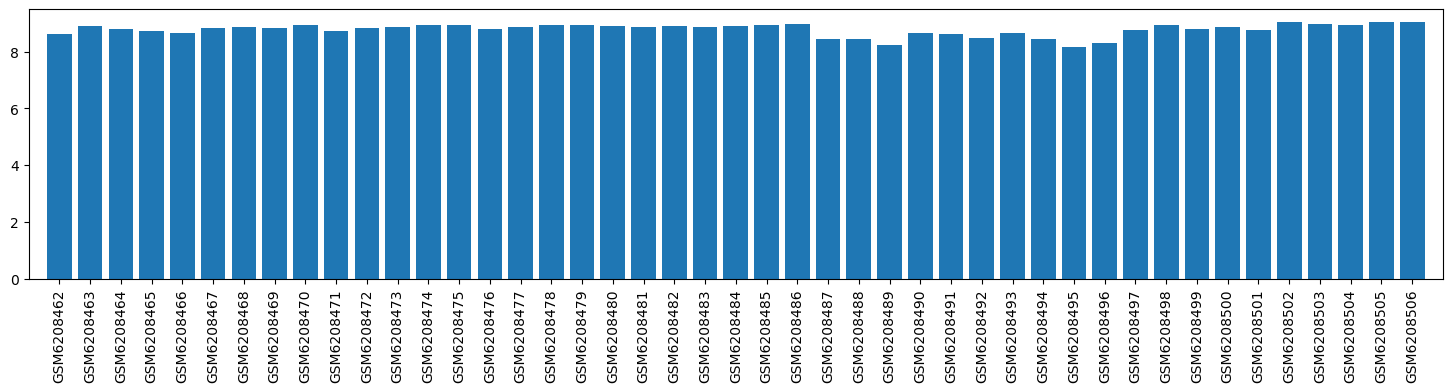
| ID | GSE205201 |
| Title | Hepatic responses of mice and rats to equivalent acetaminophen (APAP) insult [mouse] |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6208462 | Mouse_Baseline_0h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.606477 |
| GSM6208463 | Mouse_Baseline_0h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.883573 |
| GSM6208464 | Mouse_Baseline_0h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.780517 |
| GSM6208465 | Mouse_Baseline_0h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.707764 |
| GSM6208466 | Mouse_Baseline_0h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Untreated hours after treatment: 0 | 8.65406 |
| GSM6208467 | Mouse_Vehicle_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 8.833677 |
| GSM6208468 | Mouse_Vehicle_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 8.879439 |
| GSM6208469 | Mouse_Vehicle_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 8.837558 |
| GSM6208470 | Mouse_Vehicle_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 8.919347 |
| GSM6208471 | Mouse_Vehicle_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 3 | 8.715877 |
| GSM6208472 | Mouse_Vehicle_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 8.835131 |
| GSM6208473 | Mouse_Vehicle_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 8.865685 |
| GSM6208474 | Mouse_Vehicle_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 8.93512 |
| GSM6208475 | Mouse_Vehicle_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 8.940561 |
| GSM6208476 | Mouse_Vehicle_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 6 | 8.81211 |
| GSM6208477 | Mouse_Vehicle_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 8.855925 |
| GSM6208478 | Mouse_Vehicle_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 8.922967 |
| GSM6208479 | Mouse_Vehicle_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 8.929026 |
| GSM6208480 | Mouse_Vehicle_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 8.907361 |
| GSM6208481 | Mouse_Vehicle_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 9 | 8.860872 |
| GSM6208482 | Mouse_Vehicle_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 8.892549 |
| GSM6208483 | Mouse_Vehicle_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 8.870877 |
| GSM6208484 | Mouse_Vehicle_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 8.898988 |
| GSM6208485 | Mouse_Vehicle_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 8.932853 |
| GSM6208486 | Mouse_Vehicle_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: Vehicle (1% [w/v] hydroxyethylcellulose, o.g.) hours after treatment: 24 | 8.983988 |
| GSM6208487 | Mouse_APAP_03h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 8.441013 |
| GSM6208488 | Mouse_APAP_03h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 8.425114 |
| GSM6208489 | Mouse_APAP_03h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 8.244994 |
| GSM6208490 | Mouse_APAP_03h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 8.661547 |
| GSM6208491 | Mouse_APAP_03h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 3 | 8.628887 |
| GSM6208492 | Mouse_APAP_06h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 8.475861 |
| GSM6208493 | Mouse_APAP_06h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 8.645143 |
| GSM6208494 | Mouse_APAP_06h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 8.453845 |
| GSM6208495 | Mouse_APAP_06h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 8.17459 |
| GSM6208496 | Mouse_APAP_06h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 6 | 8.286887 |
| GSM6208497 | Mouse_APAP_09h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 8.742116 |
| GSM6208498 | Mouse_APAP_09h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 8.928916 |
| GSM6208499 | Mouse_APAP_09h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 8.811128 |
| GSM6208500 | Mouse_APAP_09h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 8.876576 |
| GSM6208501 | Mouse_APAP_09h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 9 | 8.754104 |
| GSM6208502 | Mouse_APAP_24h_1 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 9.027708 |
| GSM6208503 | Mouse_APAP_24h_2 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 8.973237 |
| GSM6208504 | Mouse_APAP_24h_3 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 8.926929 |
| GSM6208505 | Mouse_APAP_24h_4 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 9.04087 |
| GSM6208506 | Mouse_APAP_24h_5 | tissue: liver strain: C57Bl/6J Sex: male treatment: 300 mg/kg acetamnophen (o.g.) hours after treatment: 24 | 9.041917 |
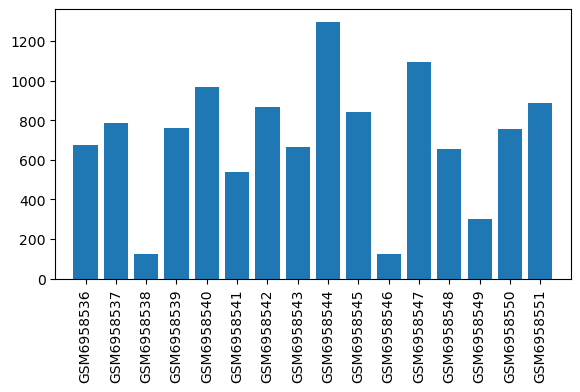
| ID | GSE223541 |
| Title | Oxycodone withdrawal induces HDAC1/2-dependent transcriptional maladaptations in the reward pathway in a murine model of peripheral nerve injury |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM6958536 | B1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sa | 675.0 |
| GSM6958537 | C1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 787.0 |
| GSM6958538 | F1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 127.0 |
| GSM6958539 | H1 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 759.0 |
| GSM6958540 | B2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 967.0 |
| GSM6958541 | C2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 540.0 |
| GSM6958542 | F2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 868.0 |
| GSM6958543 | H2 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 665.0 |
| GSM6958544 | B3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 1296.0 |
| GSM6958545 | C3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 844.0 |
| GSM6958546 | F3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 123.0 |
| GSM6958547 | H3 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 1095.0 |
| GSM6958548 | B4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: Sal | 653.0 |
| GSM6958549 | C4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: Sham condition: oxy | 303.0 |
| GSM6958550 | F4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: Sal | 757.0 |
| GSM6958551 | H4 | tissue: Brain strain: C57BL/6 Sex: male cell type: Nac genotype: SNI condition: oxy | 889.0 |
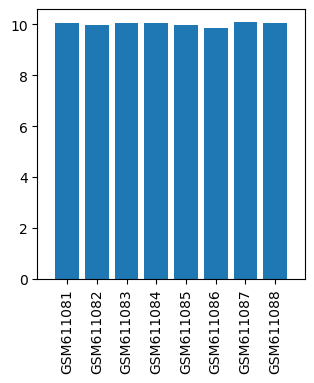
| ID | GSE24829 |
| Title | Gene expression data from striatal regions of MPTP-intoxicated mouse brain by acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM611081 | Control mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 10.04611 |
| GSM611082 | Control mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 9.97113 |
| GSM611083 | MPTP-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 10.05897 |
| GSM611084 | MPTP-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 10.04045 |
| GSM611085 | MPTP and acupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.99136 |
| GSM611086 | MPTP and acupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.8482 |
| GSM611087 | MPTP and nonacupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 10.09966 |
| GSM611088 | MPTP and nonacupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 10.03827 |
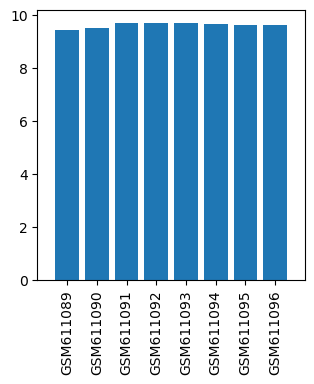
| ID | GSE24830 |
| Title | Gene expression data from cervical spinal cord regions of MPTP-intoxicated mouse by acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM611089 | Control mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 9.39743 |
| GSM611090 | Control mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 9.49079 |
| GSM611091 | MPTP-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 9.67542 |
| GSM611092 | MPTP-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 9.68217 |
| GSM611093 | MPTP and acupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.68491 |
| GSM611094 | MPTP and acupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.64677 |
| GSM611095 | MPTP and nonacupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 9.60647 |
| GSM611096 | MPTP and nonacupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 9.60152 |
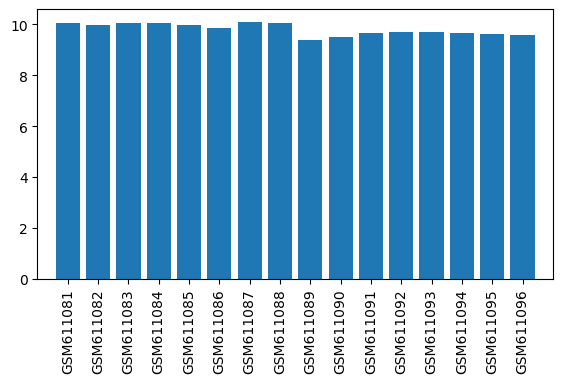
| ID | GSE24838 |
| Title | Gene expression data from brain striatal and spinal cord regions of MPTP-intoxicated mouse following acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM611081 | Control mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 10.04611 |
| GSM611082 | Control mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: Control | 9.97113 |
| GSM611083 | MPTP-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 10.05897 |
| GSM611084 | MPTP-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP-treated | 10.04045 |
| GSM611085 | MPTP and acupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.99136 |
| GSM611086 | MPTP and acupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.8482 |
| GSM611087 | MPTP and nonacupoints acupuncture-treated mouse 1 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 10.09966 |
| GSM611088 | MPTP and nonacupoints acupuncture-treated mouse 2 striatal regions | gender: Male strain: C57BL/6 tissue: Striatal tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 10.03827 |
| GSM611089 | Control mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 9.39743 |
| GSM611090 | Control mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: Control | 9.49079 |
| GSM611091 | MPTP-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 9.67542 |
| GSM611092 | MPTP-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP-treated | 9.68217 |
| GSM611093 | MPTP and acupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.68491 |
| GSM611094 | MPTP and acupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 9.64677 |
| GSM611095 | MPTP and nonacupoints acupuncture-treated mouse 1 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 9.60647 |
| GSM611096 | MPTP and nonacupoints acupuncture-treated mouse 2 cervical spinal cord regions | gender: Male strain: C57BL/6 tissue: Cervical spinal cord tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 9.60152 |
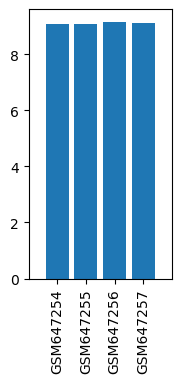
| ID | GSE26364 |
| Title | Gene expression changes in alpha GABA receptors in the mice brains after prolonged ketamine administration |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM647254 | Saline PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 9.07979 |
| GSM647255 | Saline PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 9.08178 |
| GSM647256 | Ketamine PFC 1 | strain: ICR tissue: prefrontal cortex Sex: female | 9.1613 |
| GSM647257 | Ketamine PFC 2 | strain: ICR tissue: prefrontal cortex Sex: female | 9.13544 |
| ID | GSE31827 |
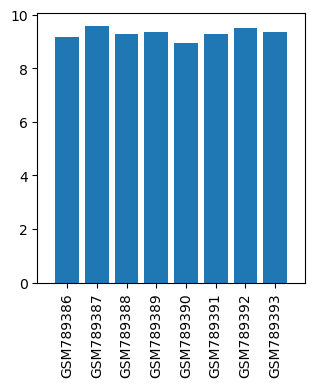
| Title | Gene expression profiling in murine mesenchymal stem cells treated with local anesthetics (ropivacaine) |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM789386 | MSC_control1 | tissue: bone marrow strain: C57B6/J agent: no treatment sample collection time: 24 hrs | 9.184 |
| GSM789387 | MSC_control2 | tissue: bone marrow strain: C57B6/J agent: no treatment sample collection time: 24 hrs | 9.584 |
| GSM789388 | MSC_control3 | tissue: bone marrow strain: C57B6/J agent: no treatment sample collection time: 24 hrs | 9.275 |
| GSM789389 | MSC_control4 | tissue: bone marrow strain: C57B6/J agent: no treatment sample collection time: 24 hrs | 9.371 |
| GSM789390 | MSC_ROPI1 | tissue: bone marrow strain: C57B6/J agent: ropivacaine 100 uM sample collection time: 24 hrs | 8.956 |
| GSM789391 | MSC_ROPI2 | tissue: bone marrow strain: C57B6/J agent: ropivacaine 100 uM sample collection time: 24 hrs | 9.284 |
| GSM789392 | MSC_ROPI3 | tissue: bone marrow strain: C57B6/J agent: ropivacaine 100 uM sample collection time: 24 hrs | 9.497 |
| GSM789393 | MSC_ROPI4 | tissue: bone marrow strain: C57B6/J agent: ropivacaine 100 uM sample collection time: 24 hrs | 9.346 |
| ID | GSE35138 |
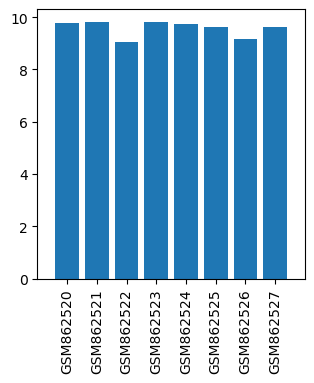
| Title | Gene expression data from thalamic regions of MPTP-intoxicated mouse brain by acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM862520 | Control mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.79442 |
| GSM862521 | Control mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.82147 |
| GSM862522 | MPTP-treated mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.0531 |
| GSM862523 | MPTP-treated mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.80789 |
| GSM862524 | MPTP and acupoints acupuncture-treated mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.74253 |
| GSM862525 | MPTP and acupoints acupuncture-treated mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.60856 |
| GSM862526 | MPTP and nonacupoints acupuncture-treated mouse 1 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.15423 |
| GSM862527 | MPTP and nonacupoints acupuncture-treated mouse 2 thalamic regions | gender: Male strain: C57BL/6 tissue: thalamic tissues developmental stage: Adult | 9.62821 |
| ID | GSE48293 |
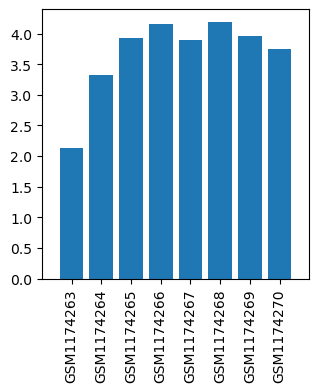
| Title | Gene expression data from substantia nigral regions of MPTP-intoxicated mouse brain by acupuncture |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1174263 | Control mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: Control | 2.13821 |
| GSM1174264 | Control mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: Control | 3.33046 |
| GSM1174265 | MPTP-treated mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP-treated | 3.92955 |
| GSM1174266 | MPTP-treated mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP-treated | 4.15633 |
| GSM1174267 | MPTP and acupoints acupuncture-treated mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 3.89853 |
| GSM1174268 | MPTP and acupoints acupuncture-treated mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and acupoints acupuncture-treated | 4.18866 |
| GSM1174269 | MPTP and nonacupoints acupuncture-treated mouse 1 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 3.95851 |
| GSM1174270 | MPTP and nonacupoints acupuncture-treated mouse 2 substantia nigral regions | gender: Male strain: C57BL/6 tissue: substantia nigral tissues developmental stage: Adult treatment: MPTP and nonacupoints acupuncture-treated | 3.74807 |
| ID | GSE57771 |
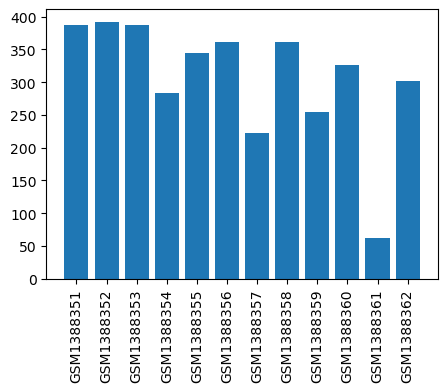
| Title | Electroacupuncture Improves Trinitrobenzene Sulphonic Acid-Induced Colitis, Evaluated by Transcriptomic Study |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM1388351 | Mock_Spleen_1 | strain: FVB tissue: Spleen | 387.1667 |
| GSM1388352 | Mock_Spleen_2 | strain: FVB tissue: Spleen | 391.9722 |
| GSM1388353 | TNBS_Spleen_1 | strain: FVB tissue: Spleen | 387.6111 |
| GSM1388354 | TNBS_Spleen_2 | strain: FVB tissue: Spleen | 283.4444 |
| GSM1388355 | EA_Spleen_1 | strain: FVB tissue: Spleen | 345.0278 |
| GSM1388356 | EA_Spleen_2 | strain: FVB tissue: Spleen | 361.7222 |
| GSM1388357 | Mock_Colon_1 | strain: FVB tissue: Colon | 222.0 |
| GSM1388358 | Mock_Colon_2 | strain: FVB tissue: Colon | 361.7222 |
| GSM1388359 | TNBS_Colon_1 | strain: FVB tissue: Colon | 254.3889 |
| GSM1388360 | TNBS_Colon_2 | strain: FVB tissue: Colon | 326.4444 |
| GSM1388361 | EA_Colon_1 | strain: FVB tissue: Colon | 62.27778 |
| GSM1388362 | EA_Colon_2 | strain: FVB tissue: Colon | 301.9722 |
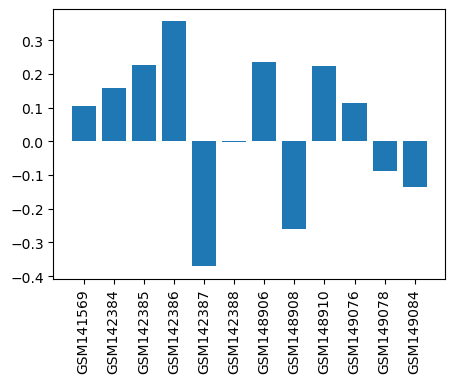
| ID | GSE6479 |
| Title | Analgesia of naloxone and morphine in wild-type and NY1DD sickle cell mice using BMAP cDNA microarrays |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM141569 | NY1DD Sickle Cell Mouse brains, NLX treated, 10mins. SCNLX-I 817 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.10557565 |
| GSM142384 | NY1DD Sickle Cell Mouse brains, NLX treated, 10mins. 819 SCNLX2 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.15797159 |
| GSM142385 | NY1DD Sickle Cell Mouse brains, NLX treated, 10mins. 825 SCNLX3 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.22549807 |
| GSM142386 | NY1DD Sickle Cell Mouse brains, MOR treated, 10mins. 819 SCMOR1 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Morphine for 10 mins.; Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.35582054 |
| GSM142387 | NY1DD Sickle Cell Mouse brains, MOR treated, 10mins. 823 SCMOR2 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Morphine for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.37100935 |
| GSM142388 | NY1DD Sickle Cell Mouse brains, MOR treated, 10mins. 825 SCMOR3 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Morphine for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.0009253729 |
| GSM148906 | C57BL/6J wild-type mouse brains, MOR treated, 10mins. 916 WTMOR 1 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Morphine for 10 mins.; Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.23432954 |
| GSM148908 | C57BL/6J wild-type mouse brains, MOR treated, 10mins. 916 WTMOR 2 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Morphine for 10 mins.; Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.2605937 |
| GSM148910 | C57BL/6J wild-type mouse brains, MOR treated, 10mins. 925 WTMOR3 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Morphine for 10 mins.; Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.2240316 |
| GSM149076 | C57BL/6J wild-type mouse brains, NLX treated, 10mins. 915 WTNLX 1 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.11331368 |
| GSM149078 | C57BL/6J wild-type mouse brains, NLX treated, 10mins. 915 WTNLX 2 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.08923228 |
| GSM149084 | C57BL/6J wild-type mouse brains, NLX treated, 10mins. 921 WTNLX 3 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.13728905 |
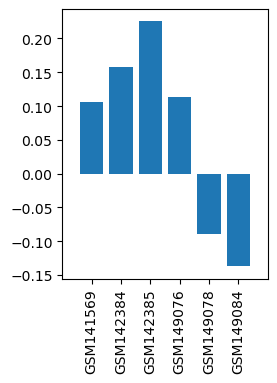
| ID | GSE6578 |
| Title | Analgesia of naloxone in wild-type and NY1DD transgenic mouse model of sickle cell anemia using BMAP cDNA microarrays |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM141569 | NY1DD Sickle Cell Mouse brains, NLX treated, 10mins. SCNLX-I 817 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.; Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.10557565 |
| GSM142384 | NY1DD Sickle Cell Mouse brains, NLX treated, 10mins. 819 SCNLX2 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.15797159 |
| GSM142385 | NY1DD Sickle Cell Mouse brains, NLX treated, 10mins. 825 SCNLX3 | Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:NY1DD Sickle Cell Mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.22549807 |
| GSM149076 | C57BL/6J wild-type mouse brains, NLX treated, 10mins. 915 WTNLX 1 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.;Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | 0.11331368 |
| GSM149078 | C57BL/6J wild-type mouse brains, NLX treated, 10mins. 915 WTNLX 2 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.; Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.08923228 |
| GSM149084 | C57BL/6J wild-type mouse brains, NLX treated, 10mins. 921 WTNLX 3 | Strain:C57BL/6J Mice Gender: Male Age: 9 wks Drug: Naloxone for 10 mins.; Strain:C57BL/6J wild-type mice Gender: Male Age: 9 wks Drug: Saline for 10 mins. | -0.13728905 |
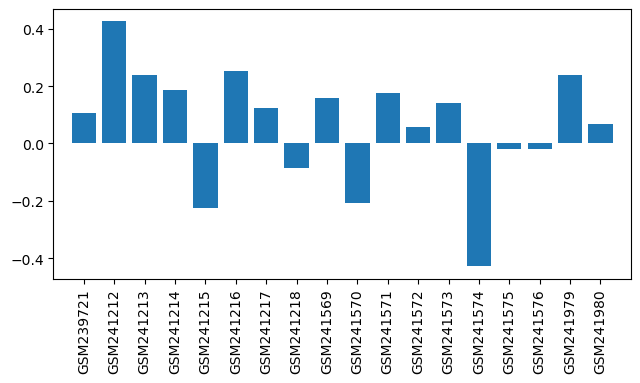
| ID | GSE9525 |
| Title | Effect of short- and long-term morphine treatment on gene expression in the hypothalamus and pituitary |
| Organism | Mus musculus |
| GSM | Sample info | Characteristics | Value |
|---|---|---|---|
| GSM239721 | Hypothalamus 4day(H1/L1) | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.107 |
| GSM241212 | Hypothalamus 4day(H1s/L1s);dye swap | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.426 |
| GSM241213 | Hypothalamus 4day(H2/L2);biological replicate | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.237 |
| GSM241214 | Hypothalamus 4day(H2s/L2s);biological replicate_dye swap | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.188 |
| GSM241215 | Pituitary 4day(P1) | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | -0.224 |
| GSM241216 | Pituitary 4day(P1s);dye swap | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | 0.253 |
| GSM241217 | Pituitary 4day(P2/L1);biological replicate | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | 0.124 |
| GSM241218 | Pituitary 4day(P2s/L1s);biological replicate_dye swap | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | -0.084 |
| GSM241569 | Hypothalamus 6hour(H1/S1) | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.159 |
| GSM241570 | Hypothalamus 6hour(H1s/S1s);dye swap | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | -0.207 |
| GSM241571 | Hypothalamus 6hour (H2/S2);biological replicate | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.176 |
| GSM241572 | Hypothalamus 6hour (H2s/S2s);biological replicate_dye swap | Strain: C57BL/6, Gender: male, Tissue: hypothalamus;Strain: C57BL/6, Gender: male, Tissue: hypothalamus | 0.059 |
| GSM241573 | Pituitary 6hour (P1/S1) | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | 0.14 |
| GSM241574 | Pituitary 6hour(P1s/S1s);dye swap | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | -0.428 |
| GSM241575 | Pituitary 6hour(P2/S2);biological replicate | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | -0.019 |
| GSM241576 | Pituitary 6hour (P2s/S2s);biological replicate_dye swap | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | -0.019 |
| GSM241979 | Pituitary 4day(L2);biological replicate | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | 0.237 |
| GSM241980 | Pituitary 4day(L2s);biological replicate_dye swap | Strain: C57BL/6, Gender: male, Tissue: pituitary;Strain: C57BL/6, Gender: male, Tissue: pituitary | 0.069 |

