Gene Information
|
Gene Name
|
CTLA4 |
|
Gene ID
|
1493
|
|
Gene Full Name
|
cytotoxic T-lymphocyte associated protein 4 |
|
Gene Alias
|
ALPS5|CD|CD152|CELIAC3|CTLA-4|GRD4|GSE|IDDM12 |
|
Transcripts
|
ENSG00000163599
|
|
Virus
|
HPV |
|
Gene Type
|
protein-coding |
|
HPA Location Info
|

|
|
Membrane Info
|
CD markers, Disease related genes, FDA approved drug targets, Human disease related genes, Predicted intracellular proteins, Predicted membrane proteins |
|
Uniport_ID
|
P16410
|
|
HGNC ID
|
HGNC:2505
|
|
OMIM ID
|
123890 |
|
Summary
|
This gene is a member of the immunoglobulin superfamily and encodes a protein which transmits an inhibitory signal to T cells. The protein contains a V domain, a transmembrane domain, and a cytoplasmic tail. Alternate transcriptional splice variants, encoding different isoforms, have been characterized. The membrane-bound isoform functions as a homodimer interconnected by a disulfide bond, while the soluble isoform functions as a monomer. Mutations in this gene have been associated with insulin-dependent diabetes mellitus, Graves disease, Hashimoto thyroiditis, celiac disease, systemic lupus erythematosus, thyroid-associated orbitopathy, and other autoimmune diseases. [provided by RefSeq, Jul 2008] |
Target gene [CTLA4] related to VISs 
Integration Table: if previous studies reported that target gene was altered by virus integration events, the overlap between VISs in this literature and Cistrome factors was listed in this section
| DVID |
Chromosome |
HM |
TFBS |
CA |
Sum of Overlapped Records |
Detail |
| 5011305 |
chr2 |
89 |
221 |
2 |
312 |
View |
Target gene [CTLA4] related to Omics data
| Data ID |
Experiment type |
Sample number |
Platform |
|
C GSE183048
|
Chip-seq |
24 |
Illumina HiSeq 4000 (Homo sapiens) |
|
S GSE226620
|
scRNA-seq |
30 |
HiSeq X Ten (Homo sapiens) |
|
GSE40774
|
Expression array |
134 |
Agilent-026652 Whole Human Genome Microarray 4x44K v2 (Probe Name version) |
|
S GSE139324
|
scRNA-seq |
63 |
Illumina NextSeq 500 (Homo sapiens) |
|
GSE169622
|
Methylation profiling (Array) |
9 |
Infinium MethylationEPIC |
|
GSE65858
|
Expression array |
270 |
Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE55550
|
Expression array |
155 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
GSE181805
|
Expression array |
25 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE196215
|
RNA-seq |
8 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
S GSE164690
|
scRNA-seq |
51 |
Illumina NextSeq 500 (Homo sapiens) |
|
S GSE197461
|
scRNA-seq |
16 |
Illumina NovaSeq 6000 (Homo sapiens) |
|
GSE140662
|
Expression array |
8 |
[HTA-2_0] Affymetrix Human Transcriptome Array 2.0 [transcript (gene) version] |
|
GSE55542
|
Expression array |
36 |
Agilent-039494 SurePrint G3 Human GE v2 8x60K Microarray 039381 (Probe Name version) |
|
C GSE143026
|
ATAC-seq;Chip-seq;RNA-seq |
30 |
Illumina HiSeq 2500 (Homo sapiens) |
|
TCGA_CESC
|
DNA methylation sequencing;RNA-seq |
288 |
TCGA |
|
GSE51993
|
Expression array |
48 |
Illumina Human v2 MicroRNA expression beadchip;Illumina HumanHT-12 V4.0 expression beadchip |
|
GSE199029
|
Variation;RNA-seq |
44 |
NanoString Human nCounter PanCancer IO360 Panel |
|
GSE165883
|
RNA-seq |
20 |
Illumina NextSeq 500 (Homo sapiens) |
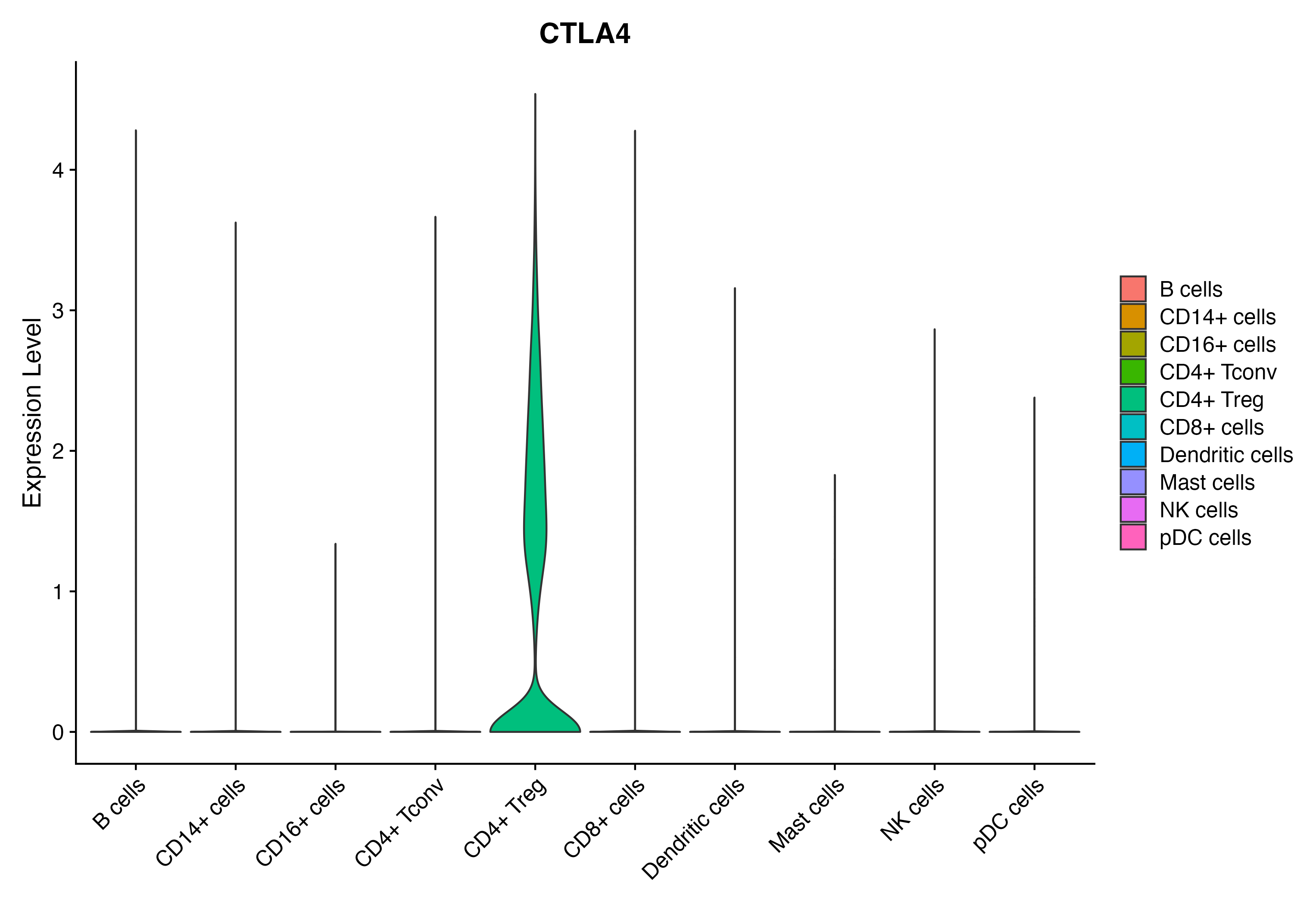
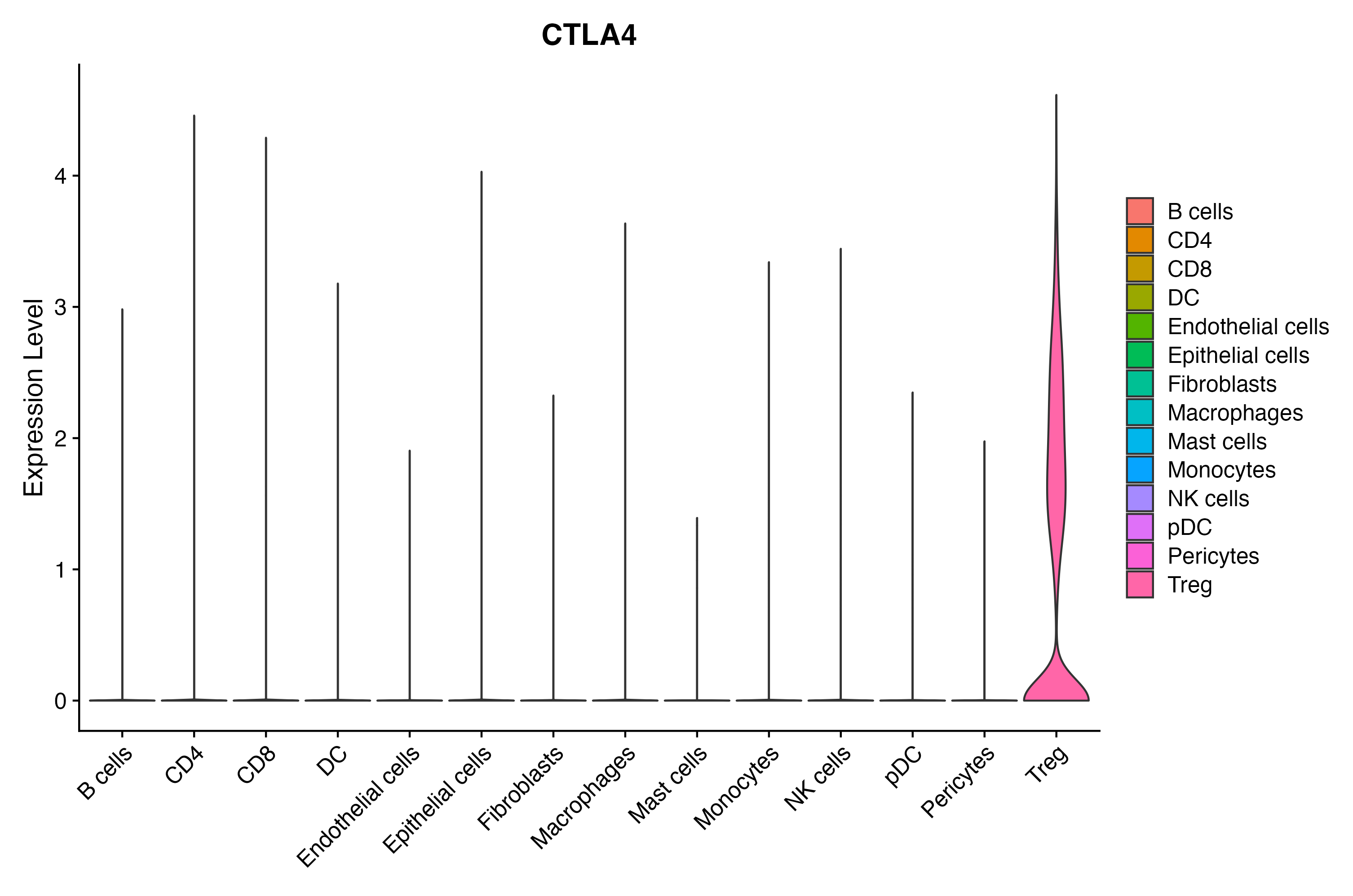
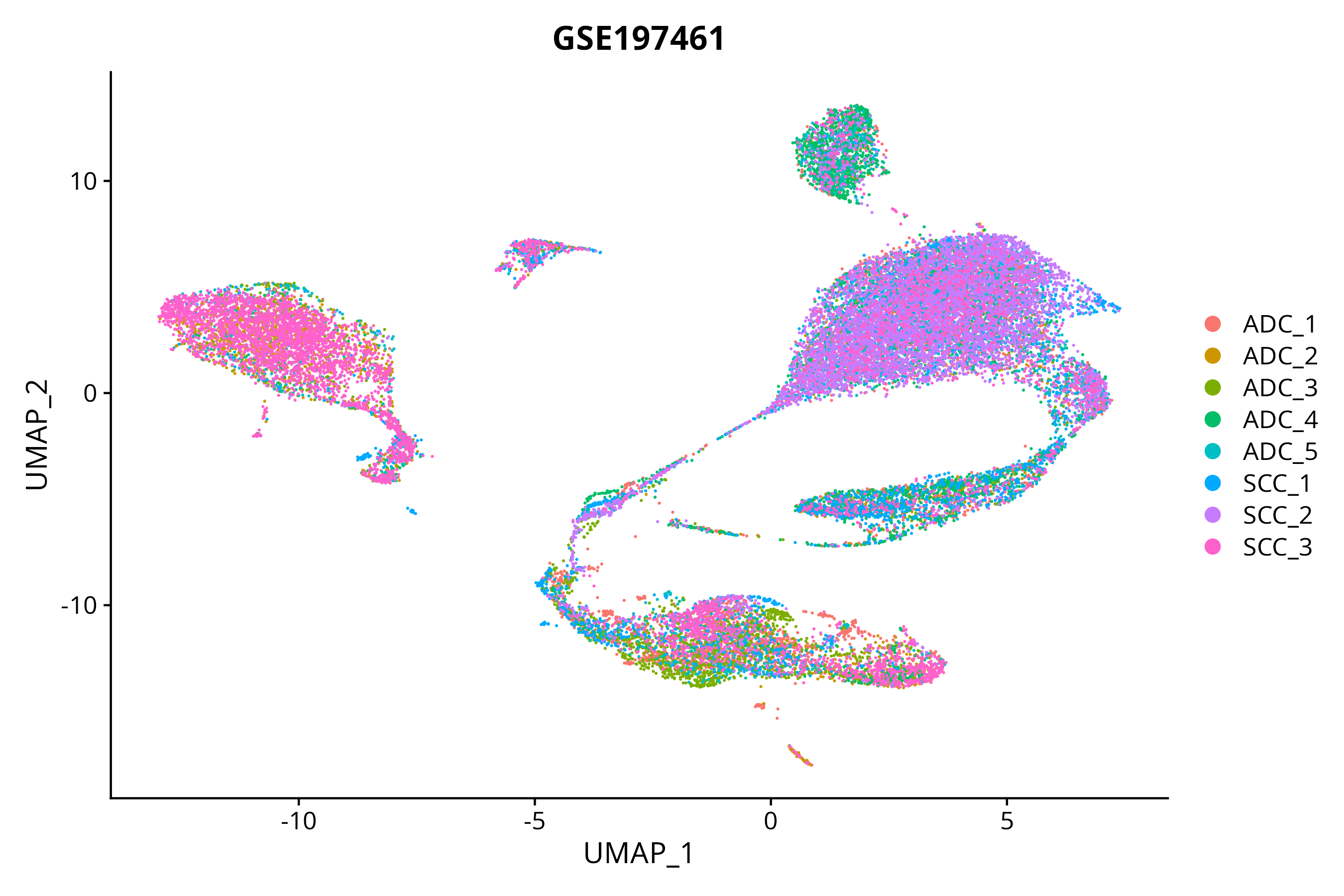
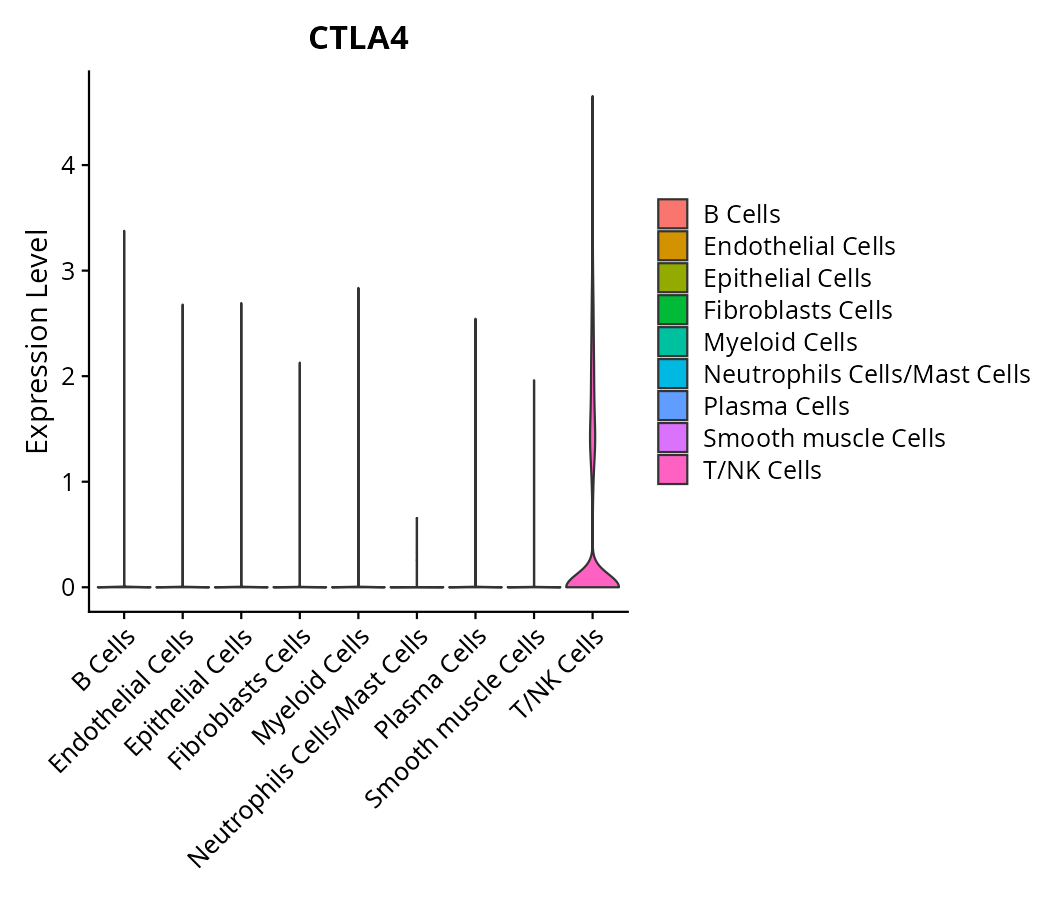
When the query gene is differentially changed in the dataset, a feature/violin plot will be displayed.
> Dataset: GSE226620 - Gene expression in cell subsets
|
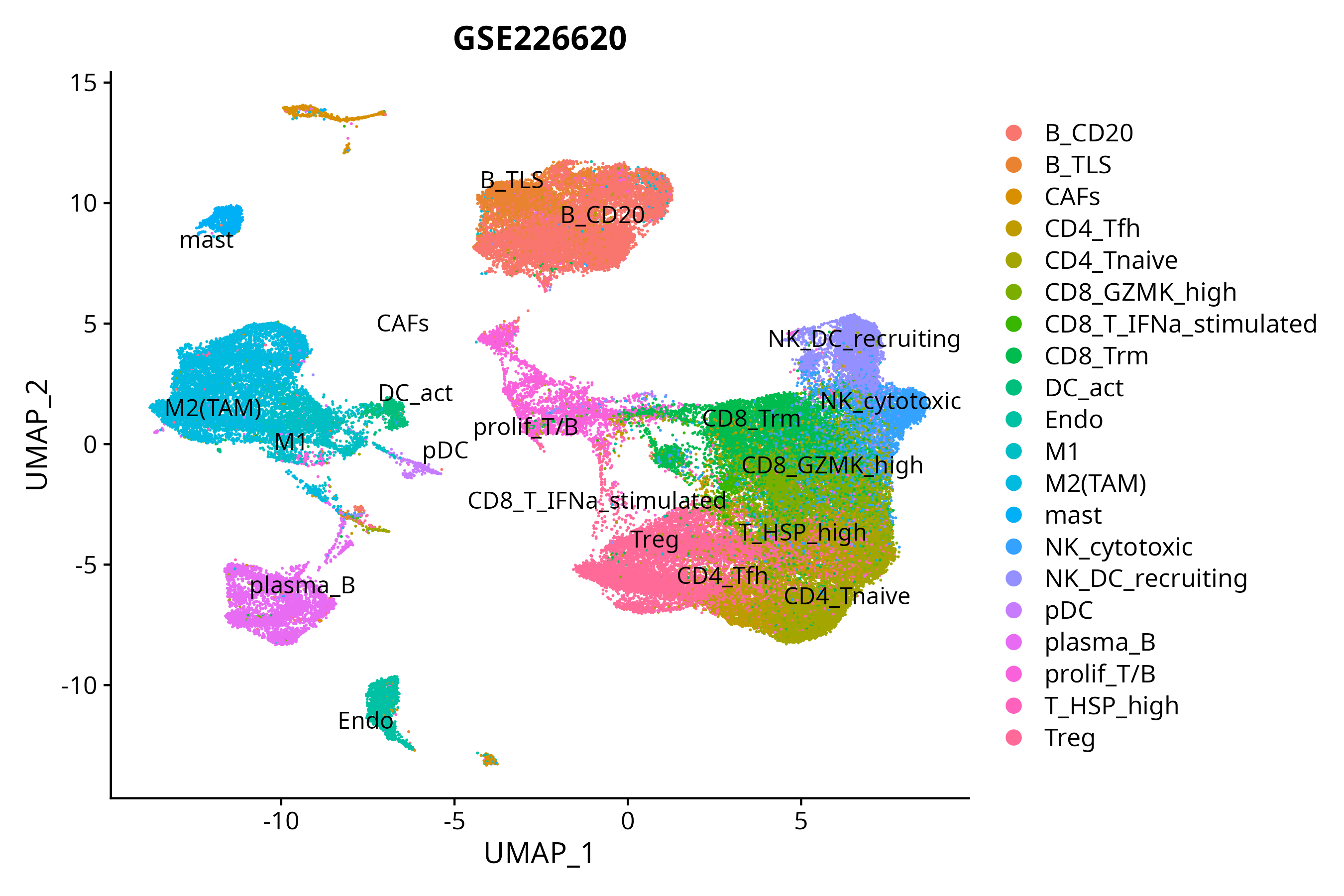
|
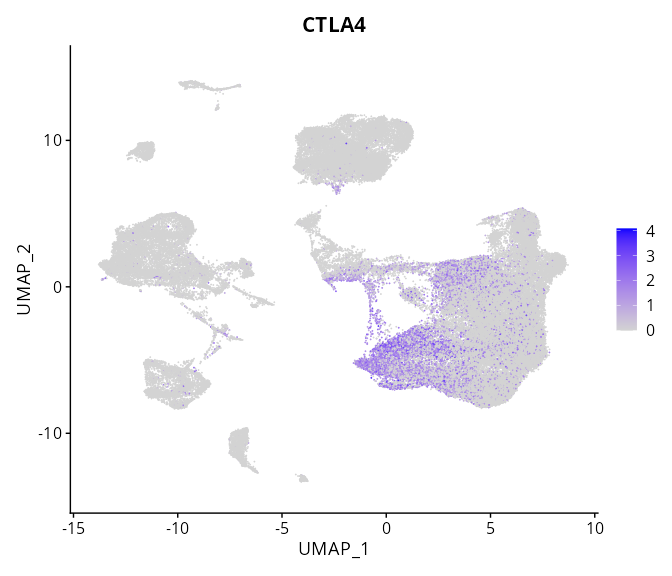
|

|
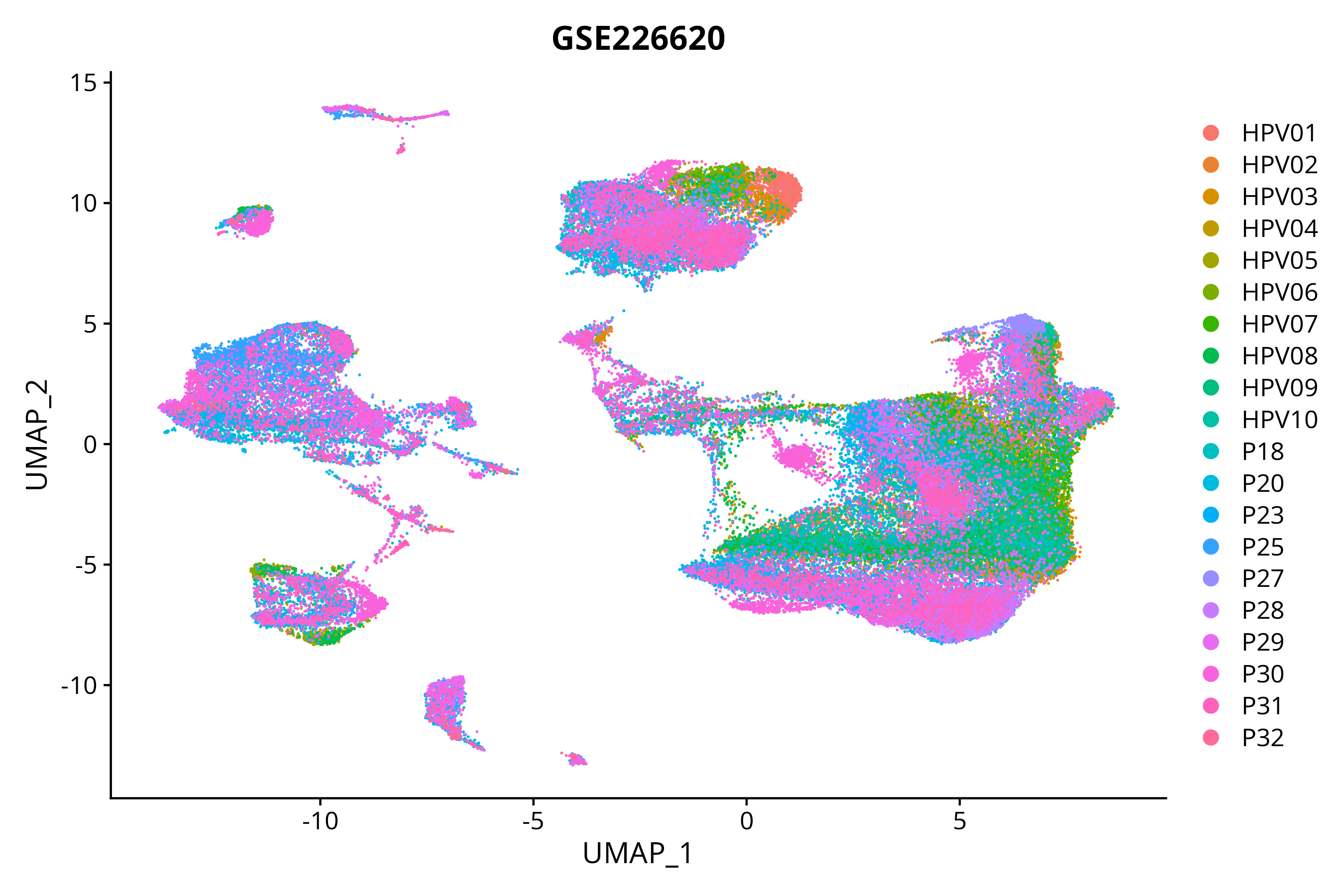
|
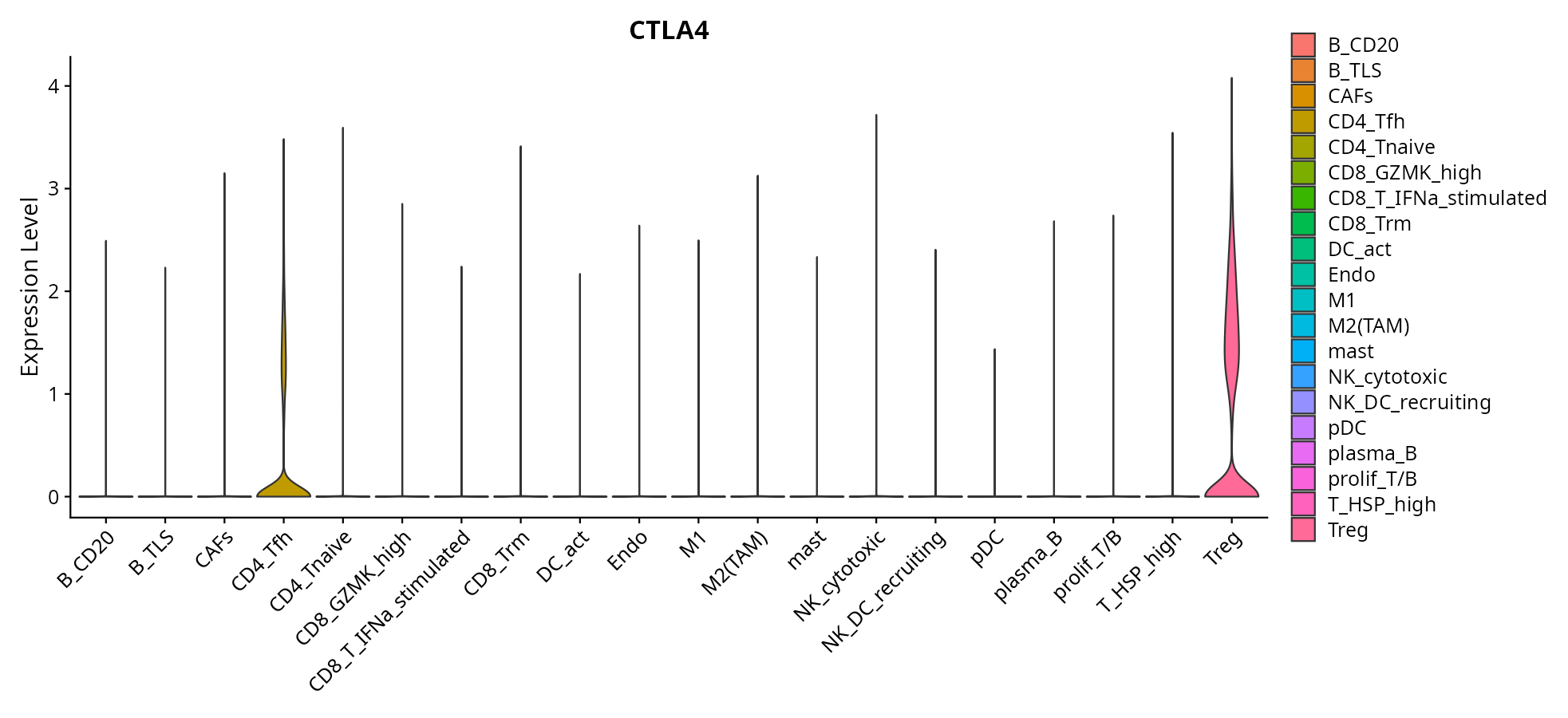
|
> Dataset: GSE139324 - Gene expression in cell subsets
|
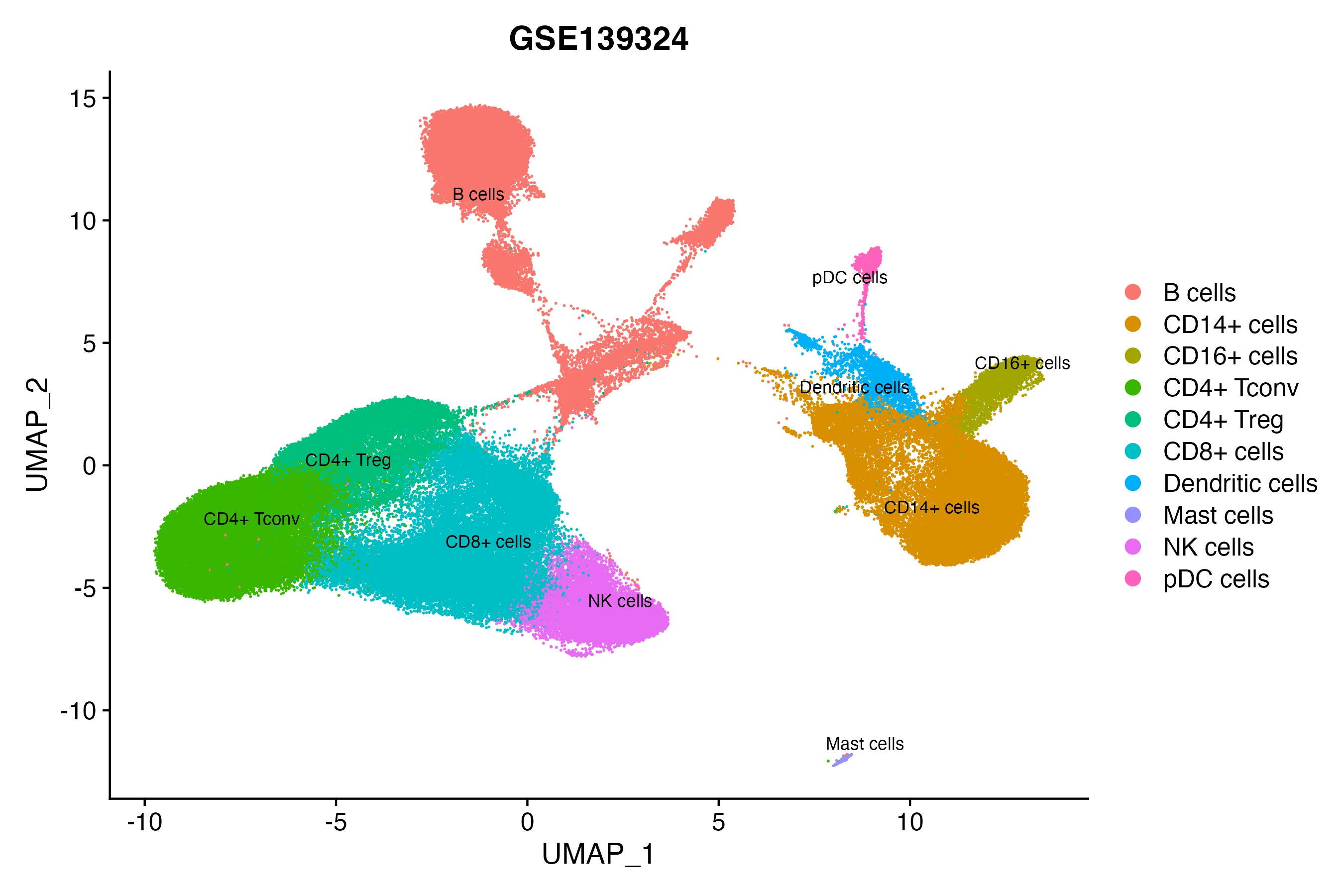
|
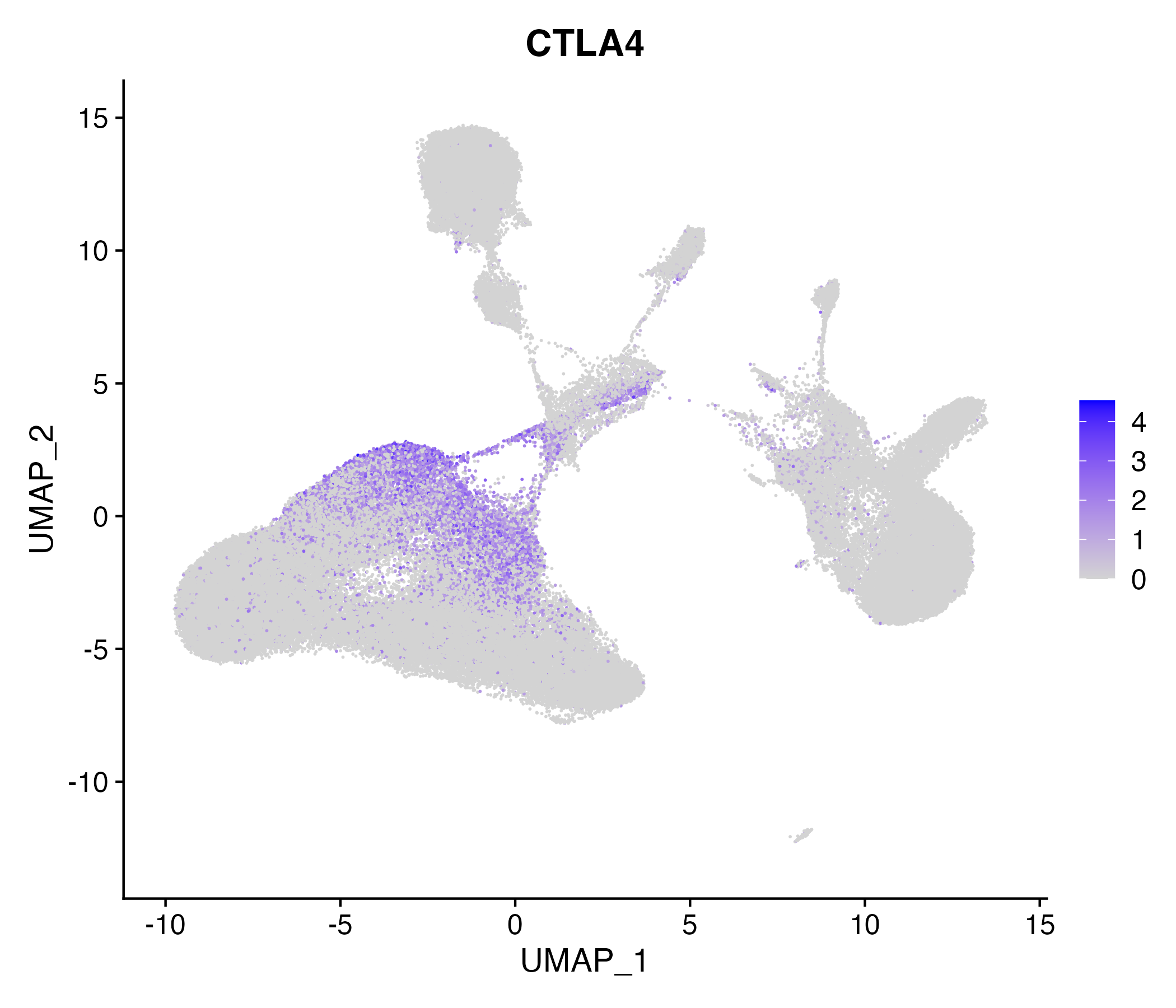
|
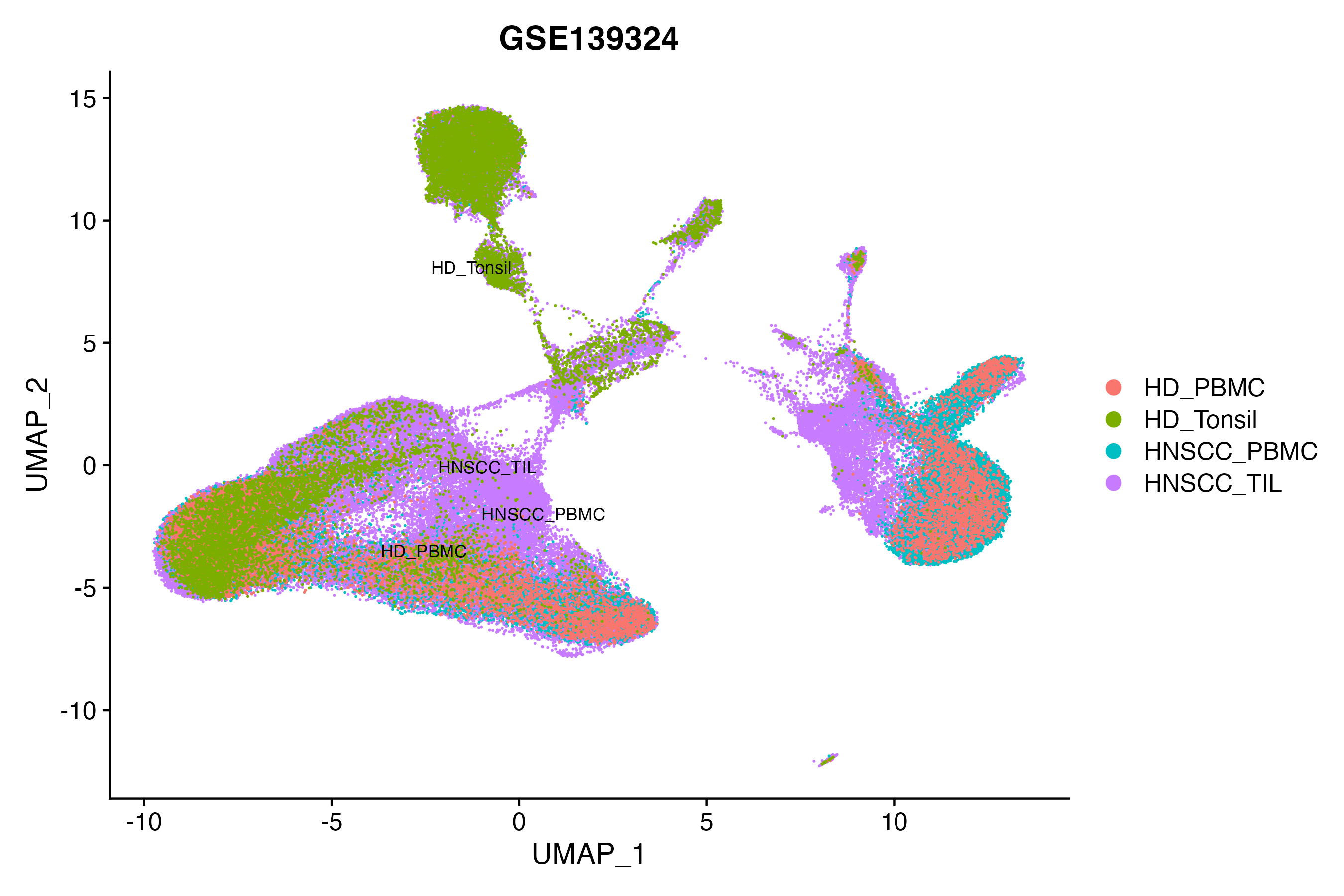
|
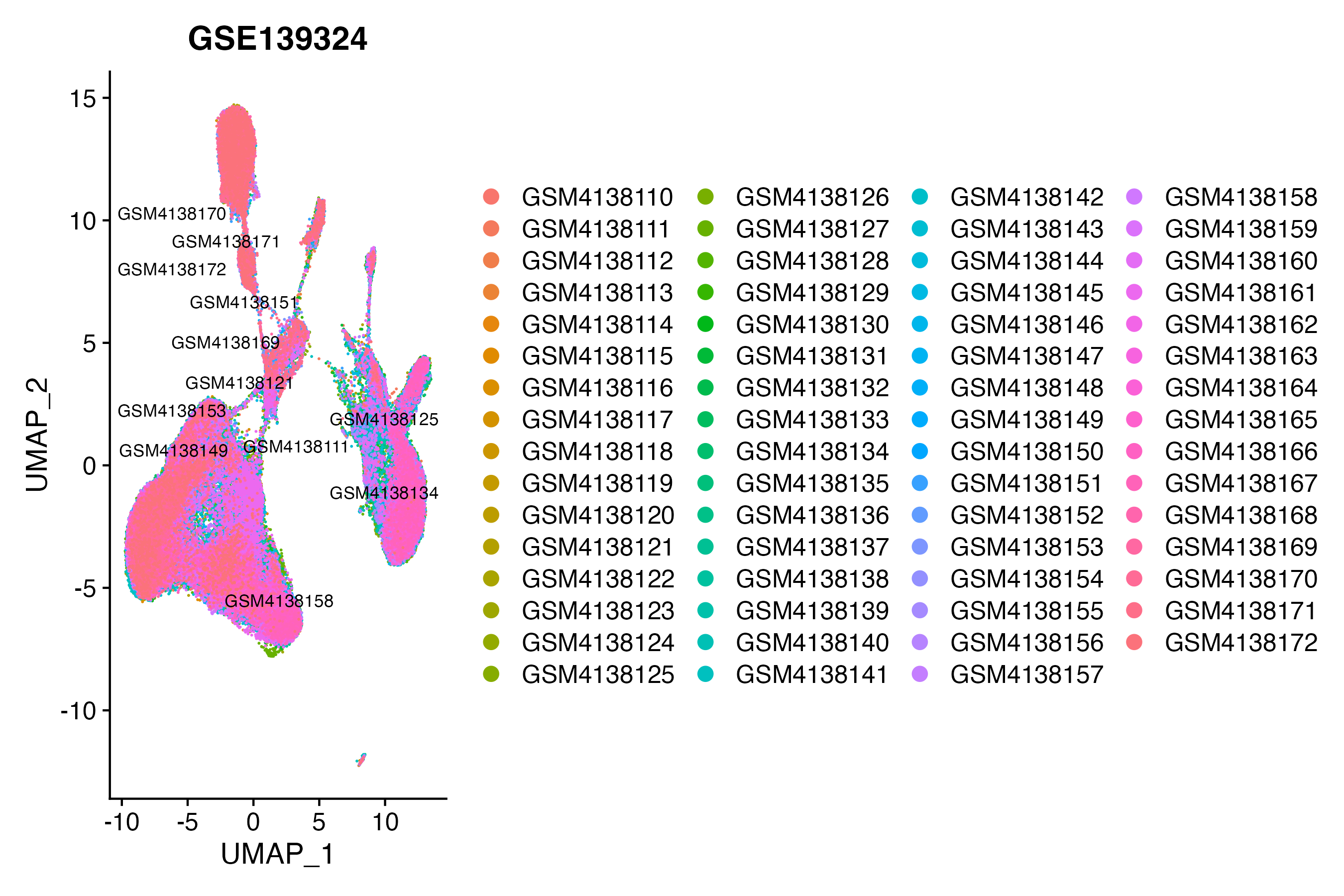
|
|
> Dataset: GSE164690 - Gene expression in cell subsets
|
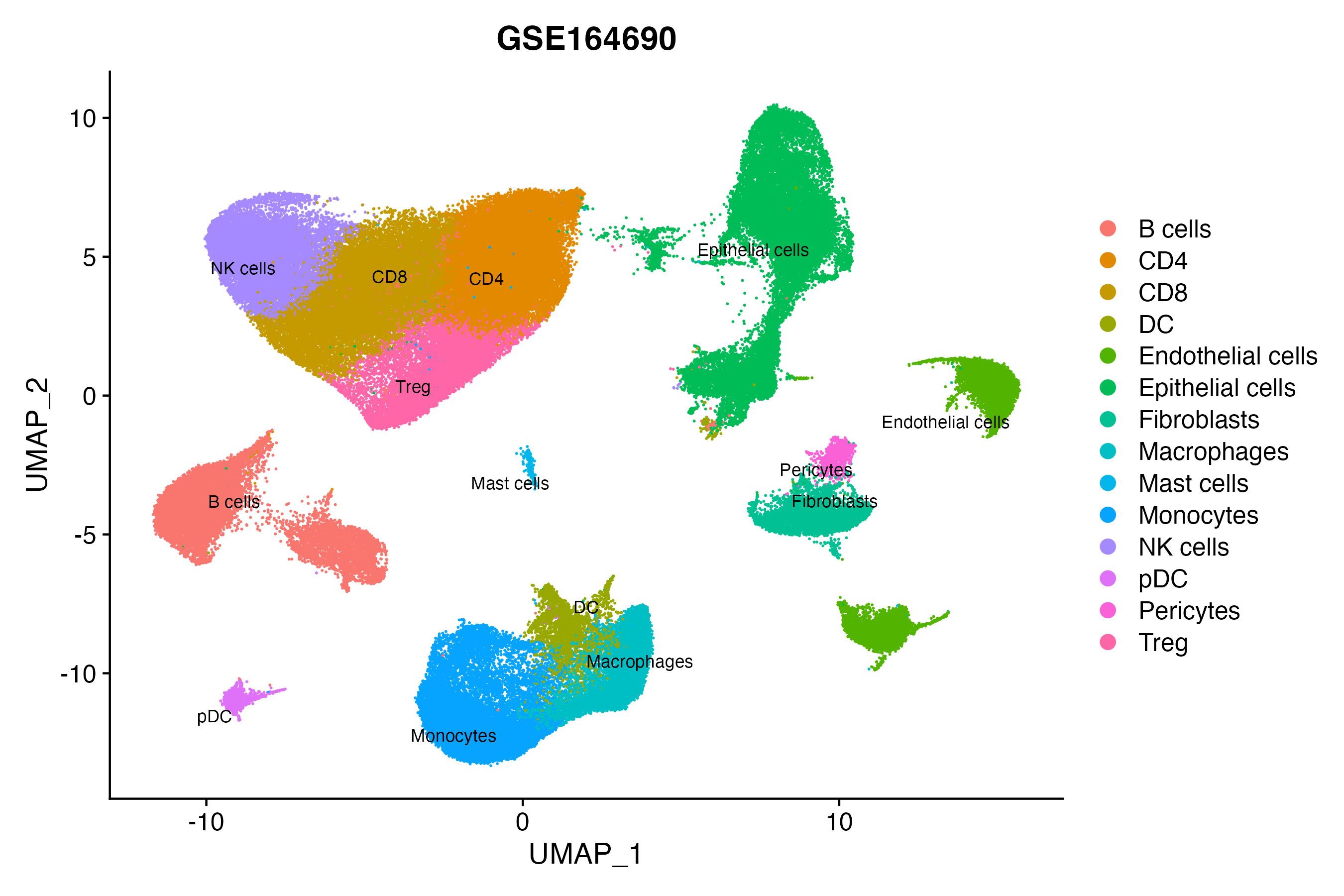
|
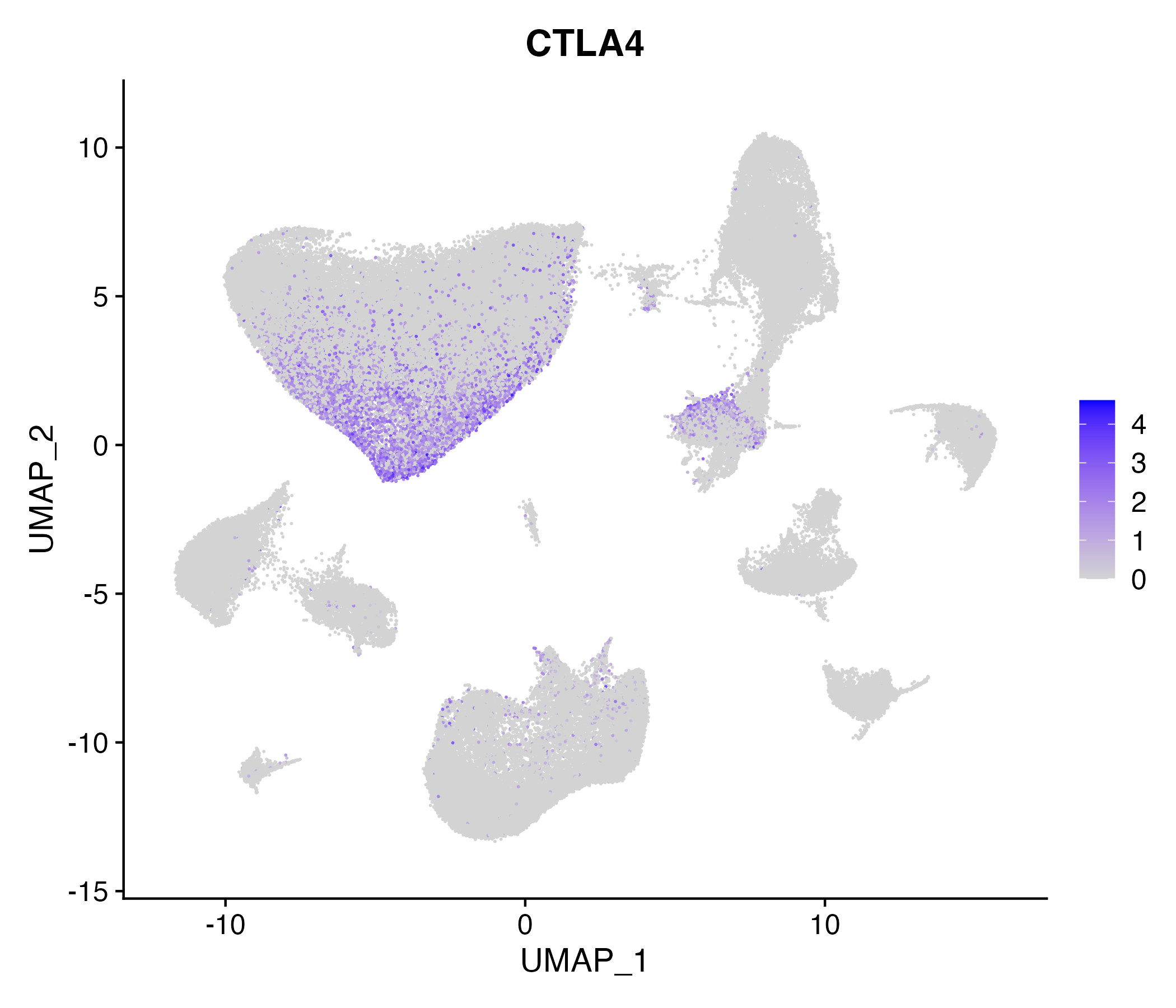
|

|
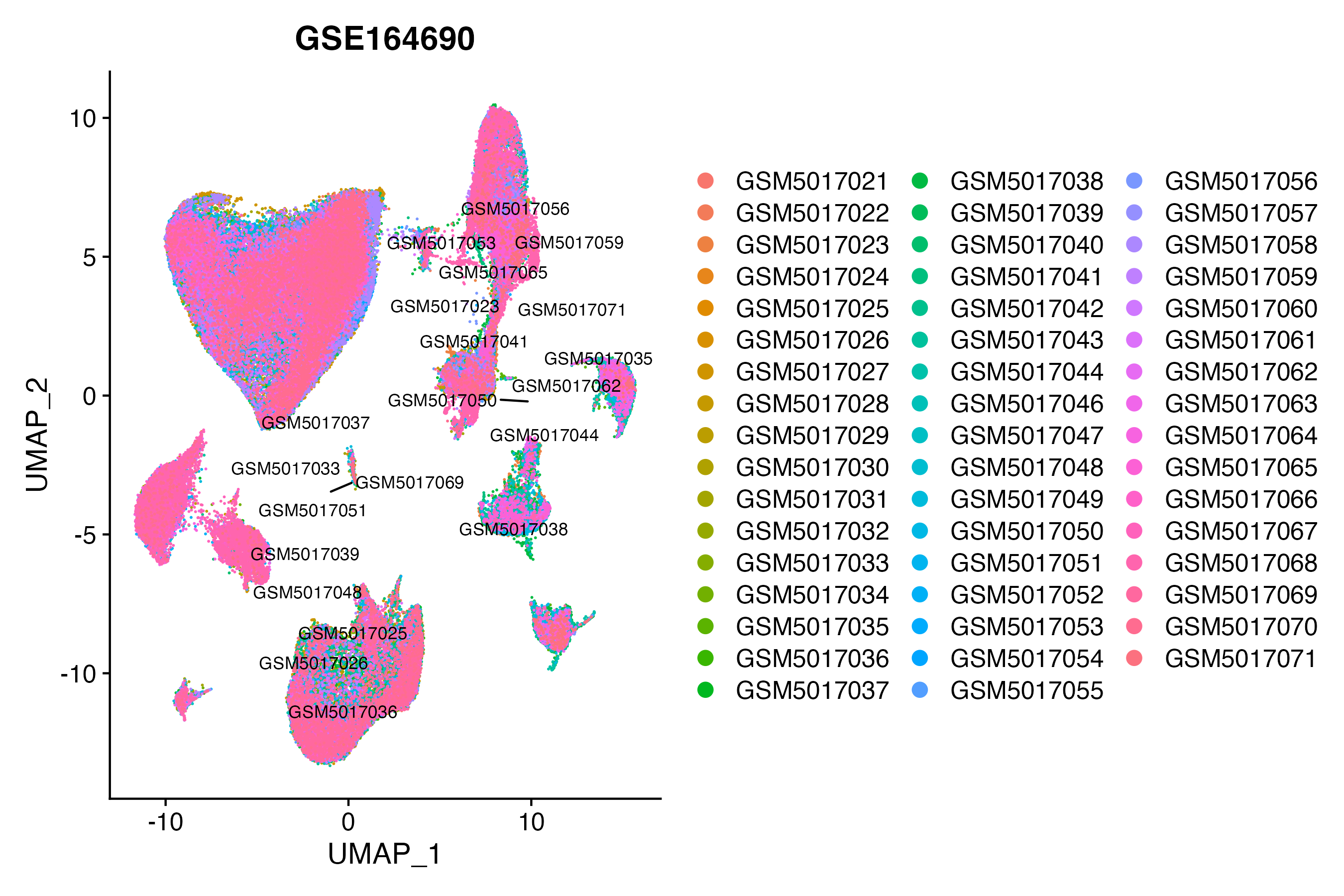
|
|
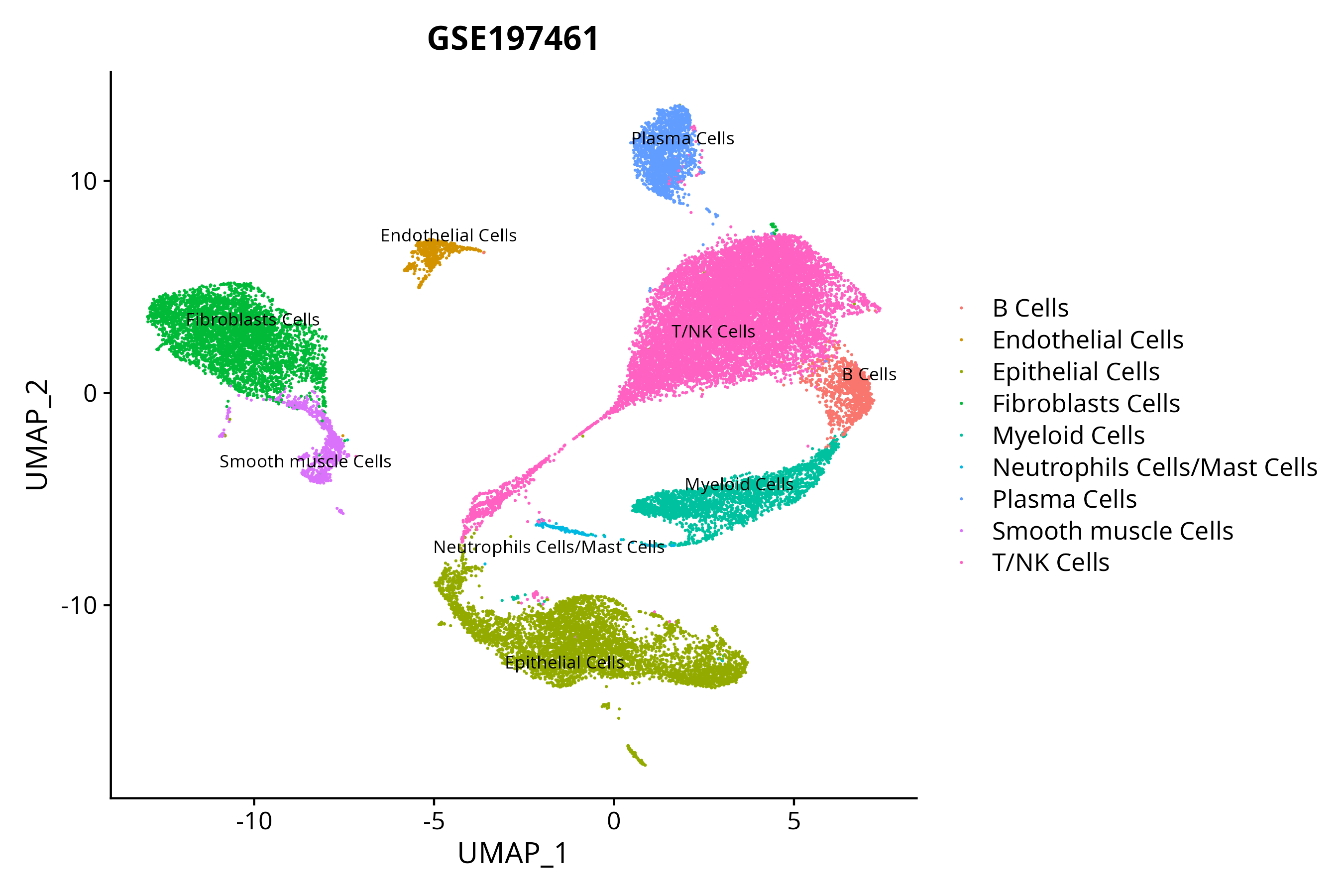
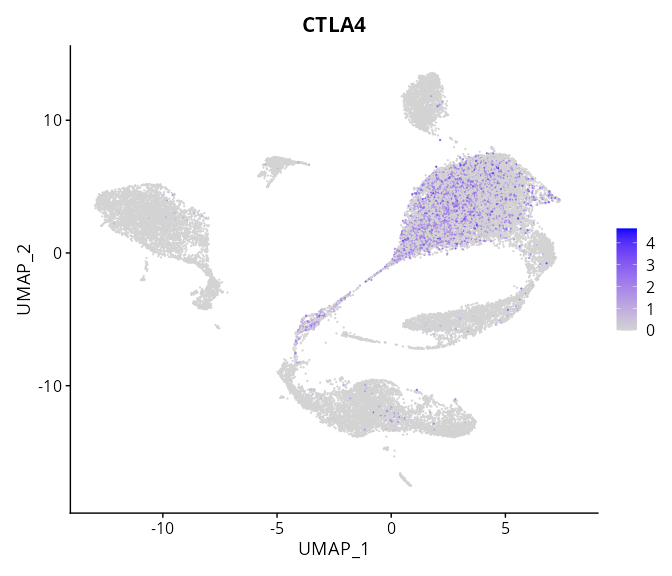
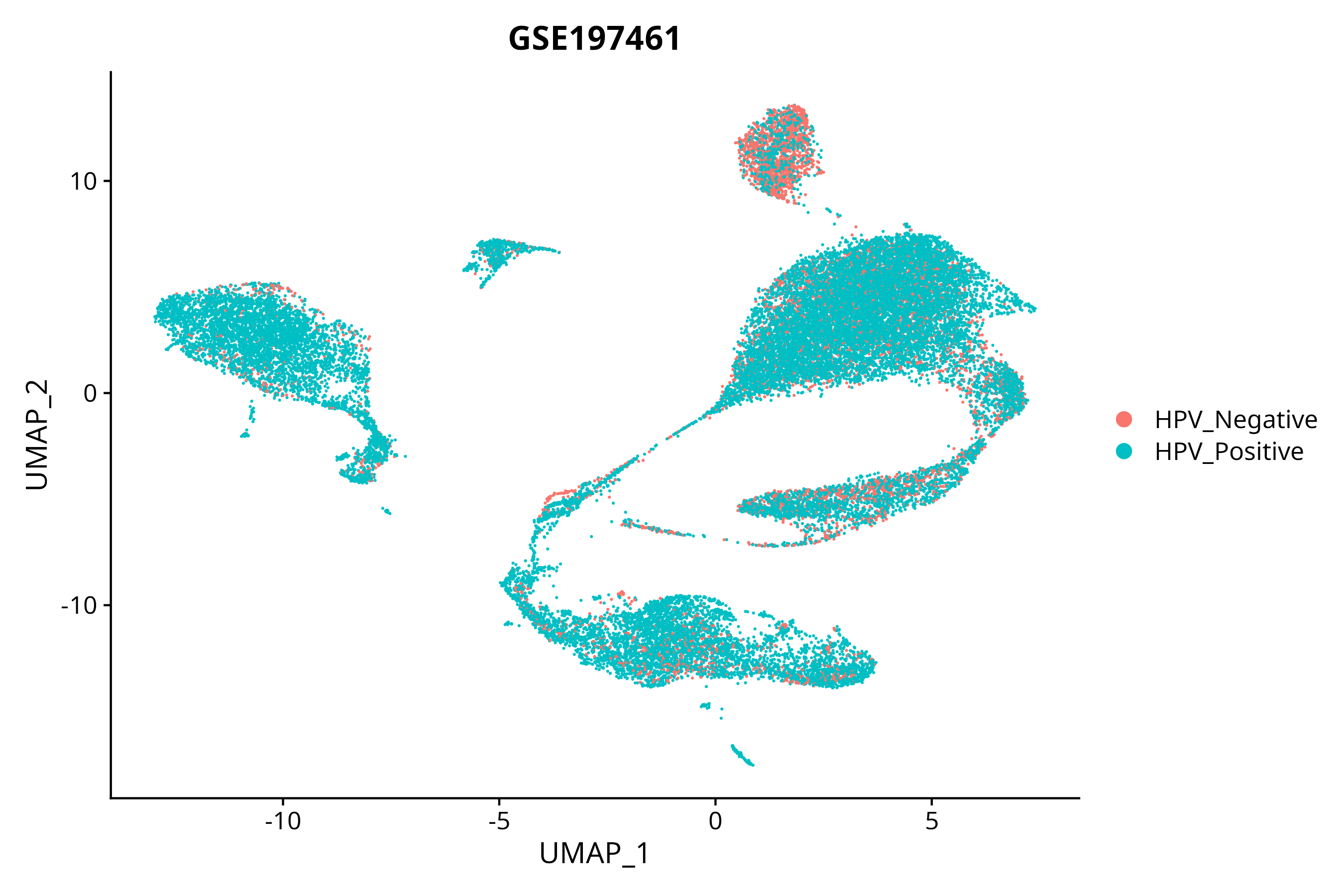
> Dataset: GSE197461 - Gene expression in cell subsets
|
|
|
|
|
|
|
|
 HPV Target gene Detail Information
HPV Target gene Detail Information